Abstract
Human pluripotent and trophoblast stem cells have been essential alternatives to blastocysts for understanding early human development1,2,3,4. However, these simple culture systems lack the complexity to adequately model the spatiotemporal cellular and molecular dynamics that occur during early embryonic development. Here we describe the reprogramming of fibroblasts into in vitro three-dimensional models of the human blastocyst, termed iBlastoids. Characterization of iBlastoids shows that they model the overall architecture of blastocysts, presenting an inner cell mass-like structure, with epiblast- and primitive endoderm-like cells, a blastocoel-like cavity and a trophectoderm-like outer layer of cells. Single-cell transcriptomics further confirmed the presence of epiblast-, primitive endoderm-, and trophectoderm-like cells. Moreover, iBlastoids can give rise to pluripotent and trophoblast stem cells and are capable of modelling, in vitro, several aspects of the early stage of implantation. In summary, we have developed a scalable and tractable system to model human blastocyst biology; we envision that this will facilitate the study of early human development and the effects of gene mutations and toxins during early embryogenesis, as well as aiding in the development of new therapies associated with in vitro fertilization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All processed datasets are available at http://blastoid.ddnetbio.com/. Raw and processed next-generation sequencing datasets from iBlastoids were deposited at the Gene Expression Omnibus (GEO) repository under accession number GSE156596. Published human embryo datasets used in this study were obtained from Blakeley et al.12 and Petropoulos et al.13 under accession numbers GSE66507 and E-MTAB-3929 (ArrayExpress, https://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-3929/), respectively. For Extended Data Fig. 6h, datasets were obtained from Table S1 of Okae et al.25 and Linneberg-Agerholm et al.53 GSE138012 for the RNA-seq integration. There are no restrictions on data availability. Source data are provided with this paper.
Code availability
All data were analysed with standard programs and packages as detailed above. Scripts can be found at http://github.com/jibsch/iBlastoids_scripts.
References
Rossant, J. & Tam, P. P. L. New insights into early human development: lessons for stem cell derivation and differentiation. Cell Stem Cell 20, 18–28 (2017).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Fu, J., Warmflash, A. & Lutolf, M. P. Stem-cell-based embryo models for fundamental research and translation. Nat. Mater. 20, 132–144 (2021).
Harrison, S. E., Sozen, B., Christodoulou, N., Kyprianou, C. & Zernicka-Goetz, M. Assembly of embryonic and extraembryonic stem cells to mimic embryogenesis in vitro. Science 356, eaal1810 (2017).
Sozen, B. et al. Self-assembly of embryonic and two extra-embryonic stem cell types into gastrulating embryo-like structures. Nat. Cell Biol. 20, 979–989 (2018).
Rivron, N. C. et al. Blastocyst-like structures generated solely from stem cells. Nature 557, 106–111 (2018).
Zhang, S. et al. Implantation initiation of self-assembled embryo-like structures generated using three types of mouse blastocyst-derived stem cells. Nat. Commun. 10, 496 (2019).
Sozen, B. et al. Self-organization of mouse stem cells into an extended potential blastoid. Dev. Cell 51, 698–712 (2019).
Li, R. et al. Generation of blastocyst-like structures from mouse embryonic and adult cell cultures. Cell 179, 687–702 (2019).
Liu, X. et al. Reprogramming roadmap reveals route to human induced trophoblast stem cells. Nature 586, 101–107 (2020).
Blakeley, P. et al. Defining the three cell lineages of the human blastocyst by single-cell RNA-seq. Development 142, 3613 (2015).
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Shahbazi, M. N. et al. Self-organization of the human embryo in the absence of maternal tissues. Nat. Cell Biol. 18, 700–708 (2016).
Xiang, L. et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos. Nature 577, 537–542 (2020).
Qin, H. et al. YAP induces human naive pluripotency. Cell Rep. 14, 2301–2312 (2016).
Liu, L. et al. An integrated chromatin accessibility and transcriptome landscape of human pre-implantation embryos. Nat. Commun. 10, 364 (2019).
Durruthy-Durruthy, J. et al. Spatiotemporal reconstruction of the human blastocyst by single-cell gene-expression analysis informs induction of naive pluripotency. Dev. Cell 38, 100–115 (2016).
Fogarty, N. M. E. et al. Genome editing reveals a role for OCT4 in human embryogenesis. Nature 550, 67–73 (2017).
Niakan, K. K. & Eggan, K. Analysis of human embryos from zygote to blastocyst reveals distinct gene expression patterns relative to the mouse. Dev. Biol. 375, 54–64 (2013).
Deglincerti, A. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016).
Roode, M. et al. Human hypoblast formation is not dependent on FGF signalling. Dev. Biol. 361, 358–363 (2012).
Kovacic, B., Vlaisavljevic, V., Reljic, M. & Cizek-Sajko, M. Developmental capacity of different morphological types of day 5 human morulae and blastocysts. Reprod. Biomed. Online 8, 687–694 (2004).
Posfai, E. et al. Evaluating totipotency using criteria of increasing stringency. Nat. Cell Biol. 23, 49–60 (2021).
Okae, H. et al. Derivation of human trophoblast stem cells. Cell Stem Cell 22, 50–63 (2018).
Tyser, R. C. V., Mahammadov, E., Nakanoh, S. & Vallier, L. A spatially resolved single cell atlas of human gastrulation. Preprint at https://doi.org/10.1101/2020.07.21.213512 (2020).
Chen, D. et al. Human primordial germ cells are specified from lineage-primed progenitors. Cell Rep. 29, 4568–4582 (2019).
West, R. C. et al. Dynamics of trophoblast differentiation in peri-implantation-stage human embryos. Proc. Natl Acad. Sci. USA 116, 22635–22644 (2019).
Alpha Scientists in Reproductive Medicine and ESHRE Special Interest Group of Embryology. The Istanbul consensus workshop on embryo assessment: proceedings of an expert meeting. Hum. Reprod. 26, 1270–1283 (2011).
Hyun, I., Munsie, M., Pera, M. F., Rivron, N. C. & Rossant, J. Toward guidelines for research on human embryo models formed from stem cells. Stem Cell Rep. 14, 169–174 (2020).
Warnock—report of the committee of inquiry into human fertilisation and embryology. Ir. Nurs. News 5, 7–8 (1985).
Guo, G. et al. Naive pluripotent stem cells derived directly from isolated cells of the human inner cell mass. Stem Cell Rep. 6, 437–446 (2016).
Tan, J. P., Liu, X. & Polo, J. M. Generation of human blastocyst-like structures by somatic cell reprogramming. Protoc. Exch. https://doi.org/10.21203/rs.3.pex-1347/v1 (2021).
Liu, X. et al. Comprehensive characterization of distinct states of human naive pluripotency generated by reprogramming. Nat. Methods 14, 1055–1062 (2017).
Zhou, F. et al. Reconstituting the transcriptome and DNA methylome landscapes of human implantation. Nature 572, 660–664 (2019).
Paynter, J. M., Chen, J., Liu, X. & Nefzger, C. M. Propagation and maintenance of mouse embryonic stem cells. Methods Mol. Biol. 1940, 33–45 (2019).
Rostovskaya, M., Stirparo, G. G. & Smith, A. Capacitation of human naïve pluripotent stem cells for multi-lineage differentiation. Development 146, dev172916 (2019).
Boroviak, T. et al. Single cell transcriptome analysis of human, marmoset and mouse embryos reveals common and divergent features of preimplantation development. Development 145, dev167833 (2018).
Lim, H. Y. G. et al. Keratins are asymmetrically inherited fate determinants in the mammalian embryo. Nature 585, 404–409 (2020).
Gerri, C. et al. Initiation of a conserved trophectoderm program in human, cow and mouse embryos. Nature 587, 443–447 (2020).
Aberkane, A. et al. Expression of adhesion and extracellular matrix genes in human blastocysts upon attachment in a 2D co-culture system. Mol. Hum. Reprod. 24, 375–387 (2018).
Bankhead, P. et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Lam, A. Q. et al. Rapid and efficient differentiation of human pluripotent stem cells into intermediate mesoderm that forms tubules expressing kidney proximal tubular markers. J. Am. Soc. Nephrol. 25, 1211–1225 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Huang, Y., McCarthy, D. J. & Stegle, O. Vireo: Bayesian demultiplexing of pooled single-cell RNA-seq data without genotype reference. Genome Biol. 20, 273 (2019).
Wickham, H. et al. dplyr: a grammar of data manipulation. R version 1.0.5 https://CRAN.R-project.org/package=dplyr (2021).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Kolde, R. & Vilo, J. GOsummaries: an R package for visual functional annotation of experimental data. F1000Res. 4, 574 (2015).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Scialdone, A. et al. Computational assignment of cell-cycle stage from single-cell transcriptome data. Methods 85, 54–61 (2015).
Linneberg-Agerholm, M. et al. Naïve human pluripotent stem cells respond to Wnt, Nodal and LIF signalling to produce expandable naïve extra-embryonic endoderm. Development 146, dev180620 (2019).
Acknowledgements
We thank the staff at Monash Micro Imaging for providing high-quality technical input and the use of imaging facilities, and acknowledge the use of the services and facilities of Micromon and Monash Flowcore Facility at Monash University. The schematics were created with BioRender.com. This work was supported by the National Health and Medical Research Council (NHMRC) project grants APP1104560 and APP2004627 to J.M.P. and APP1069830 to R.L. X.L. was supported by a Monash bridging postdoctoral fellowship and the Carmela and Carmelo Ridolfo Prize in Stem Cell Research. J.P.T. was supported by a Monash International Tuition Scholarship and Research Training Program Scholarship. J.M.P. and R.L. were supported by Silvia and Charles Viertel Senior Medical Research Fellowships. J.M.P. was supported by an ARC Future Fellowship. R.L. was supported by a Howard Hughes Medical Institute International Research Scholarship. O.J.L.R. and J.F.O. were supported by a Singapore National Research Foundation Competitive Research Programme (NRF-CRP20-2017-0002). The Australian Regenerative Medicine Institute is supported by grants from the State Government of Victoria and the Australian Government.
Author information
Authors and Affiliations
Contributions
J.M.P. and X.L. conceptualized the study and designed the experiments. X.L. and J.P.T. performed reprogramming experiments and generation of human iBlastoids, and collection and isolation of single cells for scRNA-seq and functional characterization of iBlastoids with support from Y.B.Y.S., J.C., G.S., M.M. and Y.Z. under the supervision of J.M.P. and with advice from A.T.C. D.P. helped with sequencing of iBlastoid scRNA-seq libraries under the supervision of R.L. X.L. and J.P.T. performed the immunostaining of iBlastoids with help from A.A. and J.Z. with image acquisition and analysis. J.S. performed the scRNA-seq analyses for the iBlastoids as well as the integration across the human blastocyst datasets with support from X.L., J.F.O., J.M.P. and O.J.L.R. J.F.O. and O.J.L.R. developed the interface for the interactive online tool. S.M.L. assisted with the application for ethical approval. X.L., J.P.T. and J.M.P. wrote the manuscript with input from R.L., A.T.C., O.J.L.R. and J.Z. All authors approved of and contributed to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
Although it is not directly related to this manuscript, O.J.L.R. and J.M.P. are co-inventors on a patent (WO/2017/106932) and are co-founders and shareholders of Mogrify Ltd., a cell therapy company. X.L., J.P.T. and J.M.P. are co-inventors on two provisional patent applications (application numbers: 2020904338 and 2020904340) filed by Monash University in relation to the generation of iBlastoids and derivation of stem cell lines from iBlastoids. The other authors declare no competing interests.
Additional information
Peer review information Nature thanks Jianping Fu, Josephine Johnston, Margherita Turco and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of day 21 reprogrammed cells and morphological assessment of iBlastoids.
a, scRNA-seq analysis of day 21 reprogrammed cells (6,816 cells) showing presence of EPI-like, TE-like and PE-like populations. b–d, Immunostaining of day 21 reprogrammed cells for OCT4, GATA6 and CDX2 (b) with expanded view of the region enclosed by dashed line (c) showing OCT4+GATA6+CDX2− cells (yellow arrows), OCT4+GATA6−CDX2+ cells (white arrows), OCT4−GATA6+CDX2+ cells (cyan arrows) and OCT4+GATA6+CDX2+ cells (orange arrows) (d; n = 3). Scale bars, 100 μm. e, Quantification of different existing cell populations in day 21 reprogrammed cells in b; n = 3. f, Measurement of x- and y-axis diameters of iBlastoids compared to blastocysts12,13,14,15,16,17,18,19; mean ± s.e.m., n = 18 for iBlastoids, 8 for blastocysts12,13,14,15,16,17,18,19. g, Representative iBlastoids with and without cavitation; n = 2. Scale bars, 20 μm. h, Quantification of iBlastoids with cavity formation, n = 628. i, Per-cell expression score for refractory cell signatures on UMAP of the scRNA-seq dataset for day 21 reprogrammed cells. j, Refractory cells adhering to microwell edges during iBlastoid formation, which propagated in fibroblast medium with classic fibroblast morphology; n = 2. Scale bars, 100 μm (arrows indicate step-wise extraction and culture of refractory cells). k, qRT–PCR analysis of refractory cells in j for pluripotency markers (left) and refractory cell markers (right); mean ± s.e.m., n = 3. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Extended Data Fig. 2 Immunostaining analysis of iBlastoids.
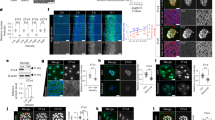
a, 3D and 2D representations of iBlastoids stained for CDX2 and NANOG; n = 5. b, iBlastoid stained for GATA2, NANOG, and SOX17, related to Fig. 1k; n = 2. c, Proportions of EPI-, PE- and TE-like cells out of total cells in iBlastoids; mean ± s.e.m., n = 5. d, iBlastoid stained for CDX2, OCT4, and GATA6, related to Fig. 1l; n = 2. e, iBlastoid stained for GATA3 and NANOG; n = 2. f, Assessment of proliferation marker Ki67 on iBlastoids stained for CDX2 and NANOG or CDX2 and GATA6; n = 2. Scale bars, 20 μm. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Extended Data Fig. 3 Time-course examination of iBlastoid formation.
a, Phase-contrast images of iBlastoid formation from day 1 to day 6; n = 5. Scale bar, 100 μm. b, Immunostaining of iBlastoids on days 1–5 for CDX2, OCT4 and GATA6; n = 2. Scale bars, 20 μm. c, Ratio of longest diameter (dmax) to shortest diameter (dmin) for TE-like and EPI-like cells in iBlastoids; mean ± s.e.m., n = 20. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Extended Data Fig. 4 scRNA-seq pipeline and quality controls.
a, Phase-contrast images of the iBlastoids selected for scRNA-seq; n = 200. Scale bar, 100 μm. b, scRNA-seq analysis strategy (see Methods). c, UMAP representation of donor cell distribution for iBlastoids. d, UMAP representation of the cell cycle stages for iBlastoid scRNA-seq library. e, Expression of Sendai-KLF4, Sendai-KOS and Sendai-MYC in iBlastoids. f, Expression of GATA3 and KRT8 on UMAP of iBlastoid dataset. g, Expression of EPI markers (POU5F1 and NANOG), TE markers (CDX2 and GATA2), and PE markers (SOX17 and GATA6) for Petropoulos scRNA-seq library13. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Extended Data Fig. 5 Scoring of the defined E5–7 EPI, TE, and PE signatures for blastocysts and iBlastoids.
a, Defined EPI, TE, and PE signatures for E5, E6, and E7 on Petropoulos dataset13. b, Defined EPI, TE, and PE signatures for E5, E6, and E7 on iBlastoid scRNA-seq dataset. c, d, Expression of refractory cell signature (c) and IFI27 expression (d) on UMAP of iBlastoid dataset. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Extended Data Fig. 6 scRNA-seq analysis of iBlastoid and blastocyst datasets.
a, Proportion of cells from each donor across all assigned clusters in the iBlastoid scRNA-seq dataset. b, Heat map showing gene expression profiles of each assigned cluster (top 10 genes each) in the iBlastoid scRNA-seq dataset. c, Unsupervised clustering of day 21 reprogrammed cells in the scRNA-seq dataset. d, Pearson correlation analysis for clusters from day 21 reprogrammed cells with annotated IM clusters from the iBlastoid scRNA-seq dataset. e, UMAP projection showing cell distribution of iBlastoids and blastocysts12,13 in the integrated dataset of iBlastoids and E5–7 blastocysts12,13. f, Expression of EPI markers (POU5F1 and NANOG), TE markers (CDX2 and GATA2), and PE markers (SOX17 and GATA6) for the integrated dataset. g, Per-cell expression score of EPI, TE, and PE signatures in the integrated dataset. h, PCA plot of pseudo-bulk RNA-seq data from scRNA-seq dataset of iBlastoids and blastocysts12,13 integrated with RNA-seq dataset of PS, TS and nEnd cell lines25,53. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Extended Data Fig. 7 Assessment of mural and polar TE gene expression in iBlastoid scRNA-seq dataset.
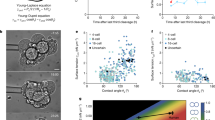
a, Unsupervised clustering of the integrated dataset with assigned cluster names. b, Proportion of cells for iBlastoid and human blastocyst datasets12,13 within each integrated cluster with respective original cell ID before integration analysis. c, Pearson correlation analysis of iBlastoid EPI, TE, and PE clusters with annotated EPI, TE, and PE clusters from blastocysts12,13. d, UMAP representation of E5–E7 TE cells of Petropoulos dataset13. e, f, Defined mural (e) and polar (f) TE signatures for the Petropoulos dataset13. g, Binned subtype scores for mural and polar TE signatures along UMAP component 1. h, Immunostaining of iBlastoid for CCR7; n = 4. Scale bar, 20 μm. i, CCR7 fluorescence intensity of polar and mural TE cells on iBlastoids; n = 4. Line, median; whiskers, maxima and minima; box limits, 25th to 75th percentiles. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Extended Data Fig. 8 Characterization of stem cells derived from iBlastoids.
a, Schematic of the experimental design for characterization of naive bPSCs, primed bPSCs and bTSCs. b, Immunostaining of naive bPSCs for KLF17 and NANOG, primed bPSCs for TRA160 and NANOG and bTSCs for KRT7 and GATA2, related to Fig. 3b–g; n = 2. c, qRT–PCR analysis of naive bPSCs for naive pluripotency markers, primed bPSCs for primed pluripotency markers and bTSCs for TSC markers; mean ± s.e.m., n = 4. d, e, EBs derived from #primed bPSCs (d) and primed bPSCs (e); n = 2. # indicates primed bPSCs generated from naive bPSCs. f, ScoreCard Assay analysis of EBs from #primed bPSCs and primed bPSCs; n = 2. g, h, Differentiation of #primed bPSCs (g) and primed bPSCs (h) into ectoderm, endoderm and mesoderm lineages; n = 2. i, Immunostaining of bTSC-differentiated EVTs for HLA-G; n = 2. j, Immunostaining of bTSC-differentiated STs for SDC1; n = 2. k, qRT–PCR analysis of bTSC-differentiated EVTs for EVT markers (ITGA1, ITGA5, FN1); mean ± s.e.m., n = 4. l, qRT–PCR analysis of bTSC-differentiated STs for ST markers (CSH1, SDC1, HSD3B1); mean ± s.e.m., n = 4. m, Fusion index of bTSC-differentiated STs; mean ± s.e.m., n = 12, two-tailed unpaired Student’s t-test. n, hCG protein level detected by hCG ELISA using conditioned medium collected from bTSC-differentiated STs; mean ± s.e.m., n = 2–3. Scale bars, 100 μm. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Extended Data Fig. 9 Assessment of epiblast development in attached iBlastoids.
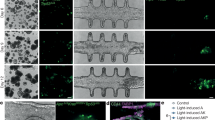
a, Schematic of the experimental design for iBlastoid in vitro attachment assay. b, Time-course qRT–PCR analysis of primitive streak markers (TBXT, EOMES, and MIXL1) in iBlastoids for up to day 5 in the attachment assay; mean ± s.e.m., n = 6. Positive control for TBXT, EOMES, and MIXL1 was generated using a previously published mesoderm differentiation protocol43. c, Attached iBlastoids stained for CDX2 and NANOG; n = 5. Scale bars, 100 μm. d, Attached iBlastoids stained for GATA2, OCT4, and SOX2; n = 2. Scale bars, 100 μm. e, Expanded view of attached iBlastoids for NANOG and SOX17 staining, related to Fig. 4b; n = 2. f, Expanded view of attached iBlastoids for OCT4 and GATA6 staining, related to Fig. 4c; n = 2. Scale bars for e, f, 50 μm. g, Z-section series of attached iBlastoids stained with F-actin, OCT4 and aPKC; bottom, expanded view of pro-amniotic cavity. n = 2. Scale bar, 10 μm. Arrows mark appearance of the pro-amniotic-like cavity. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Extended Data Fig. 10 Characterization of attached iBlastoids.
a, Immunostaining of iBlastoids, day 1 attached iBlastoids and day 3 attached iBlastoids for F-actin, OCT4 and aPKC; n = 2. Scale bars, 20 μm. Arrows mark appearance of the pro-amniotic-like cavity. b, Immunostaining of iBlastoids and attached iBlastoids for KRT7 and NANOG, related to Fig. 4e; n = 4. c, Attached iBlastoids stained for F-actin and NANOG, related to Fig. 4f; n = 2. d, Immunostaining of iBlastoids and attached iBlastoids for MMP2 and hCG, related to Fig. 4g; n = 2. e, Attached iBlastoids stained for HLA-G and NANOG with induced trophoblast stem cell-differentiated EVTs as positive control for HLA-G immunostaining (left). Scale bars for b–e, 50 μm. f, qRT–PCR analysis of ST markers (SDC1, HSD3B1; top) and EVT markers (ITGA5, FN1; bottom) in attached iBlastoids; mean ± s.e.m., n = 5. g, Assessment criteria29 used for scoring of iBlastoids. h, Phase-contrast images of iBlastoids included for scoring assessment; n = 24. Scale bar, 100 μm. i, Average grades of ICM and TE for iBlastoids in h according to g; n = 24. For more details on sample numbers, see ‘Statistics and Reproducibility’.
Supplementary information
Supplementary Figure 1
Flow cytometry gating strategy of dissociated iBlastoid cells for scRNA-seq. Cell debris were first excluded using the SSC vs FSC-H gate; Aggregates were then excluded via the FSC-A vs FSC-H; Dead cells were defined as PI high/positive populations and were excluded; Live cell (PI-negative/low) populations were sorted for scRNA-seq.
Supplementary Table 1
scRNA-seq dataset of iBlastoids. Information regarding each single cell for the iBlastoid scRNA-seq dataset used in this study.
Supplementary Table 2
scRNA-seq dataset of Petropoulos et al. Information regarding each single cell for the Petropoulos et al13 scRNA-seq dataset used in this study.
Supplementary Table 3
Top marker genes among each cluster in iBlastoid scRNA-seq dataset. Lists of DEGs among each cluster in iBlastoid scRNA-seq dataset, related to Extended Data Fig. 6b.
Supplementary Table 4
Integrated scRNA-seq dataset. Information regarding each single cell for the integrated iBlastoid, Petropoulos and Blakeley scRNA-seq dataset used in this study.
Supplementary Table 5
Polar and Mural TE gene signatures. Top 100 polar and mural DEGs at E6 obtained from Petropoulos et al13 for the generation of polar and mural TE gene signatures used in this study.
Supplementary Table 6
Pro-amniotic-like cavity formation in iBlastoids. Proportion of attached iBlastoids developing a pro-amniotic-like cavity.
Supplementary Table 7
Scoring of iBlastoids. Assessment of iBlastoids in accordance to IVF embryo grading criteria with score graded for each ICM-like and TE-like compartment, related to Extended Data Fig. 10g-i.
Supplementary Table 8
qRT-PCR primers. Information regarding the list of qRT-PCR primers used in this study.
Supplementary Table 9
Immunostaining antibodies. Information regarding the list of antibodies used for immunostaining in this study.
Video 1
: Z-series scanning of the iBlastoids Different confocal z-sections of the iBlastoids stained for CDX2 (cyan) and NANOG (orange) overlaid with Differential Interference Contrast (DIC). Scale bar, 10 μm.
Video 2
: 3D representation of the iBlastoids Representative iBlastoids stained for CDX2 (cyan), NANOG (orange) and DAPI (grey) overlaid with Differential Interference Contrast (DIC). ICM-like cellular compartment is indicated by an arrow in the video. Scale bar, 10 μm.
Video 3
: Z-series scanning of the pro-amniotic-like cavity in the attached iBlastoids. Different confocal z-sections of the iBlastoids stained for OCT4 (orange), aPKC (magenta) and F-actin (light blue) with the pro-amniotic-like cavity indicated by an arrow in the video. The pro-amniotic-like cavity surrounded by OCT4-expressing cells is highlighted by 3D segmentation of the cavity (grey). Scale bar, 10 μm.
Video 4
: 3D representation and z-series scanning of the attached iBlastoids Different confocal z-sections of the iBlastoids stained for F-actin (light blue), NANOG (orange) and DAPI (grey). Scale bar, 30 μm.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, X., Tan, J.P., Schröder, J. et al. Modelling human blastocysts by reprogramming fibroblasts into iBlastoids. Nature 591, 627–632 (2021). https://doi.org/10.1038/s41586-021-03372-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03372-y
This article is cited by
-
Hypoblast from human pluripotent stem cells regulates epiblast development
Nature (2024)
-
Self-renewing human naïve pluripotent stem cells dedifferentiate in 3D culture and form blastoids spontaneously
Nature Communications (2024)
-
VGLL1 cooperates with TEAD4 to control human trophectoderm lineage specification
Nature Communications (2024)
-
Reprogramming fibroblast into human iBlastoids
Nature Protocols (2024)
-
Modelling post-implantation human development to yolk sac blood emergence
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.