Abstract
Systemic insulin sensitivity shows a diurnal rhythm with a peak upon waking1,2. The molecular mechanism that underlies this temporal pattern is unclear. Here we show that the nuclear receptors REV-ERB-α and REV-ERB-β (referred to here as ‘REV-ERB’) in the GABAergic (γ-aminobutyric acid-producing) neurons in the suprachiasmatic nucleus (SCN) (SCNGABA neurons) control the diurnal rhythm of insulin-mediated suppression of hepatic glucose production in mice, without affecting diurnal eating or locomotor behaviours during regular light–dark cycles. REV-ERB regulates the rhythmic expression of genes that are involved in neurotransmission in the SCN, and modulates the oscillatory firing activity of SCNGABA neurons. Chemogenetic stimulation of SCNGABA neurons at waking leads to glucose intolerance, whereas restoration of the temporal pattern of either SCNGABA neuron firing or REV-ERB expression rescues the time-dependent glucose metabolic phenotype caused by REV-ERB depletion. In individuals with diabetes, an increased level of blood glucose after waking is a defining feature of the ‘extended dawn phenomenon’3,4. Patients with type 2 diabetes with the extended dawn phenomenon exhibit a differential temporal pattern of expression of REV-ERB genes compared to patients with type 2 diabetes who do not have the extended dawn phenomenon. These findings provide mechanistic insights into how the central circadian clock regulates the diurnal rhythm of hepatic insulin sensitivity, with implications for our understanding of the extended dawn phenomenon in type 2 diabetes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are freely available from the corresponding authors upon request. RNA-seq data are available at the Gene Expression Omnibus (GEO) with the accession code GSE150840. Source data are provided with this paper.
Change history
15 June 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-03654-5
References
Shi, S. Q., Ansari, T. S., McGuinness, O. P., Wasserman, D. H. & Johnson, C. H. Circadian disruption leads to insulin resistance and obesity. Curr. Biol. 23, 372–381 (2013).
Coomans, C. P. et al. Detrimental effects of constant light exposure and high-fat diet on circadian energy metabolism and insulin sensitivity. FASEB J. 27, 1721–1732 (2013).
O’Neal, T. B. & Luther, E. E. Dawn phenomenon. https://www.statpearls.com/articlelibrary/viewarticle/20266/ (StatPearls Publishing, 2020).
Monnier, L., Colette, C., Dejager, S. & Owens, D. Magnitude of the dawn phenomenon and its impact on the overall glucose exposure in type 2 diabetes: is this of concern? Diabetes Care 36, 4057–4062 (2013).
Hastings, M. H., Maywood, E. S. & Brancaccio, M. Generation of circadian rhythms in the suprachiasmatic nucleus. Nat. Rev. Neurosci. 19, 453–469 (2018).
Zhang, R., Lahens, N. F., Ballance, H. I., Hughes, M. E. & Hogenesch, J. B. A circadian gene expression atlas in mammals: implications for biology and medicine. Proc. Natl Acad. Sci. USA 111, 16219–16224 (2014).
Cho, H. et al. Regulation of circadian behaviour and metabolism by REV-ERB-α and REV-ERB-β. Nature 485, 123–127 (2012).
Zhang, Y. et al. Discrete functions of nuclear receptor Rev-erbα couple metabolism to the clock. Science 348, 1488–1492 (2015).
Doi, M. et al. Circadian regulation of intracellular G-protein signalling mediates intercellular synchrony and rhythmicity in the suprachiasmatic nucleus. Nat. Commun. 2, 327 (2011).
Tu, S. et al. Takusan: a large gene family that regulates synaptic activity. Neuron 55, 69–85 (2007).
Panda, S. et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109, 307–320 (2002).
Adelmant, G., Bègue, A., Stéhelin, D. & Laudet, V. A functional Rev-erbα responsive element located in the human Rev-erbα promoter mediates a repressing activity. Proc. Natl Acad. Sci. USA 93, 3553–3558 (1996).
Carroll, M. F. & Schade, D. S. The dawn phenomenon revisited: implications for diabetes therapy. Endocr. Pract. 11, 55–64 (2005).
Porcellati, F., Lucidi, P., Bolli, G. B. & Fanelli, C. G. Thirty years of research on the dawn phenomenon: lessons to optimize blood glucose control in diabetes. Diabetes Care 36, 3860–3862 (2013).
Cuesta, M., Boudreau, P., Cermakian, N. & Boivin, D. B. Rapid resetting of human peripheral clocks by phototherapy during simulated night shift work. Sci. Rep. 7, 16310 (2017).
Akashi, M. et al. Noninvasive method for assessing the human circadian clock using hair follicle cells. Proc. Natl Acad. Sci. USA 107, 15643–15648 (2010).
la Fleur, S. E., Kalsbeek, A., Wortel, J., Fekkes, M. L. & Buijs, R. M. A daily rhythm in glucose tolerance: a role for the suprachiasmatic nucleus. Diabetes 50, 1237–1243 (2001).
Coomans, C. P. et al. The suprachiasmatic nucleus controls circadian energy metabolism and hepatic insulin sensitivity. Diabetes 62, 1102–1108 (2013).
Foppen, E., Tan, A. A., Ackermans, M. T., Fliers, E. & Kalsbeek, A. Suprachiasmatic nucleus neuropeptides and their control of endogenous glucose production. J. Neuroendocrinol. 28, https://doi.org/10.1111/jne.12365 (2016).
Kalsbeek, A., Yi, C.-X., La Fleur, S. E. & Fliers, E. The hypothalamic clock and its control of glucose homeostasis. Trends Endocrinol. Metab. 21, 402–410 (2010).
Bolli, G. B. et al. Demonstration of a dawn phenomenon in normal human volunteers. Diabetes 33, 1150–1153 (1984).
Van Cauter, E., Polonsky, K. S. & Scheen, A. J. Roles of circadian rhythmicity and sleep in human glucose regulation. Endocr. Rev. 18, 716–738 (1997).
Boden, G., Chen, X. & Urbain, J. L. Evidence for a circadian rhythm of insulin sensitivity in patients with NIDDM caused by cyclic changes in hepatic glucose production. Diabetes 45, 1044–1050 (1996).
Radziuk, J. & Pye, S. Diurnal rhythm in endogenous glucose production is a major contributor to fasting hyperglycaemia in type 2 diabetes. Suprachiasmatic deficit or limit cycle behaviour? Diabetologia 49, 1619–1628 (2006).
Albus, H., Vansteensel, M. J., Michel, S., Block, G. D. & Meijer, J. H. A GABAergic mechanism is necessary for coupling dissociable ventral and dorsal regional oscillators within the circadian clock. Curr. Biol. 15, 886–893 (2005).
Choi, H. J. et al. Excitatory actions of GABA in the suprachiasmatic nucleus. J. Neurosci. 28, 5450–5459 (2008).
Freeman, G. M., Jr, Krock, R. M., Aton, S. J., Thaben, P. & Herzog, E. D. GABA networks destabilize genetic oscillations in the circadian pacemaker. Neuron 78, 799–806 (2013).
Vong, L. et al. Leptin action on GABAergic neurons prevents obesity and reduces inhibitory tone to POMC neurons. Neuron 71, 142–154 (2011).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Siepka, S. M. & Takahashi, J. S. Methods to record circadian rhythm wheel running activity in mice. Methods Enzymol. 393, 230–239 (2005).
Atasoy, D., Aponte, Y., Su, H. H. & Sternson, S. M. A FLEX switch targets channelrhodopsin-2 to multiple cell types for imaging and long-range circuit mapping. J. Neurosci. 28, 7025–7030 (2008).
Sprengel, R. & Hasan, M. T. Tetracycline-controlled genetic switches. Handb. Exp. Pharmacol. 178, 49–72 (2007).
Ochoa, C. D., Alexeyev, M., Pastukh, V., Balczon, R. & Stevens, T. Pseudomonas aeruginosa exotoxin Y is a promiscuous cyclase that increases endothelial tau phosphorylation and permeability. J. Biol. Chem. 287, 25407–25418 (2012).
Hockemeyer, D. et al. A drug-inducible system for direct reprogramming of human somatic cells to pluripotency. Cell Stem Cell 3, 346–353 (2008).
Roth, B. L. DREADDs for neuroscientists. Neuron 89, 683–694 (2016).
Krashes, M. J. et al. Rapid, reversible activation of AgRP neurons drives feeding behavior in mice. J. Clin. Invest. 121, 1424–1428 (2011).
Armbruster, B. N., Li, X., Pausch, M. H., Herlitze, S. & Roth, B. L. Evolving the lock to fit the key to create a family of G protein-coupled receptors potently activated by an inert ligand. Proc. Natl Acad. Sci. USA 104, 5163–5168 (2007).
Ren, H. et al. FoxO1 target Gpr17 activates AgRP neurons to regulate food intake. Cell 149, 1314–1326 (2012).
Liu, T. et al. Fasting activation of AgRP neurons requires NMDA receptors and involves spinogenesis and increased excitatory tone. Neuron 73, 511–522 (2012).
Fenselau, H. et al. A rapidly acting glutamatergic ARC→PVH satiety circuit postsynaptically regulated by α-MSH. Nat. Neurosci. 20, 42–51 (2017).
Itri, J., Michel, S., Waschek, J. A. & Colwell, C. S. Circadian rhythm in inhibitory synaptic transmission in the mouse suprachiasmatic nucleus. J. Neurophysiol. 92, 311–319 (2004).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Danne, T. et al. International consensus on use of continuous glucose monitoring. Diabetes Care 40, 1631–1640 (2017).
Faraco, G. et al. Dietary salt promotes neurovascular and cognitive dysfunction through a gut-initiated TH17 response. Nat. Neurosci. 21, 240–249 (2018).
Peixoto, R. T., Wang, W., Croney, D. M., Kozorovitskiy, Y. & Sabatini, B. L. Early hyperactivity and precocious maturation of corticostriatal circuits in Shank3B−/− mice. Nat. Neurosci. 19, 716–724 (2016).
Witton, J. et al. Hippocampal circuit dysfunction in the Tc1 mouse model of Down syndrome. Nat. Neurosci. 18, 1291–1298 (2015).
Xu, P. et al. Estrogen receptor-α in medial amygdala neurons regulates body weight. J. Clin. Invest. 125, 2861–2876 (2015).
Perusini, J. N. et al. Optogenetic stimulation of dentate gyrus engrams restores memory in Alzheimer’s disease mice. Hippocampus 27, 1110–1122 (2017).
Wang, W. et al. Chemogenetic activation of prefrontal cortex rescues synaptic and behavioral deficits in a mouse model of 16p11.2 deletion syndrome. J. Neurosci. 38, 5939–5948 (2018).
Acknowledgements
We thank M. Lazar for the Nr1d1loxP and Nr1d2loxP mice; Q. Tong and Y. Xu for critical reading of the manuscript and technical guidance; H. Liu for technical consultation; X. Wang for statistics consultation; C. Yu and S. S. B. Lee for technical assistance; K. Oka and B. Arenkiel for viral vector production; and C. Ljungberg (U54HD083092 and 1S10OD016167) for some of the histology studies. The Mouse Metabolism and Phenotyping Core at Baylor College of Medicine was supported by R01DK114356 and UM1HG006348. The authors were supported by grants from the National Natural Science Foundation of China (81971458 and 31671222 to G.D.); the American Diabetes Association (ADA1-17-PDF-138 to Y. He, ADA1-19-PDF-012 to W.Z. and NIH P20 GM135002 to Y. He); the US Department of Agriculture (Cris51000-064-01S to Y.X.); the National Key R&D Program of China (2016YFC0901204 and 2018YFC1311801 to L.C. and 2017YFC1001300 to G.D.); and the NIH (R01DK111436, R01HL153320, RF1AG069966 and R01ES027544 to Z.S.). We also thank the John S. Dunn Foundation, the Mrs. Clifford Elder White Graham Endowed Research Fund, the Cardiovascular Research Institute at Baylor College of Medicine, the Dan L Duncan Comprehensive Cancer Center (P30CA125123), the Texas Medical Center Digestive Diseases Center (P30DK056338), the SPORE program in lymphoma at Baylor College of Medicine (P50 CA126752) and the Gulf Coast Center for Precision Environmental Health (P30ES030285).
Author information
Authors and Affiliations
Contributions
Z.S. conceived the study. G.D. identified the mouse phenotype and oversaw the human study. X.L. performed gene expression analysis, chemogenetic studies, and stereotaxic injections in mice. X.H. recruited study participants and supervised the human study. W.Z. performed histological studies and ChIP. Y.G. coordinated the insulin clamp. W.L. made DNA constructs. S.Q. performed some of the mouse metabolic tests. J.S. performed gene expression analyses in human samples. J.W., F.L., J.W. and C.C. collected human blood samples, made clinical measurements and coordinated clinical studies. Y. He performed patch clamp recording. P.B. performed the initial mouse crossbreeding. G.D., X.L. and Y.G. maintained the mouse lines. P.S. performed CLAMS and insulin clamp analyses. G.D., X.L., W.Z., Y.G., J.S., J.W., Y. He, P.B., T.Y. and Z.S. analysed the data. K.Z., Y. Han and C.I.A. performed statistical analyses. Y.X., X.H. and Z.S. interpreted the data. L.C. and Z.S. obtained funding. G.D. and Z.S. wrote the manuscript with input from other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no financial or non-financial conflict of interest. No patent was involved in the study.
Additional information
Peer review information Nature thanks Erik D. Herzog, Satchidananda Panda and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Behavioural characterization of knockout mice.
a, RNAscope analysis of Nr1d1 (REV-ERB-α) gene expression at ZT6–9 and ZT18–21 in three-month-old wild-type mice. Scale bars, 200 μm b, Representative wheel-running actogram for five-month-old mice in light–dark conditions (LD) or constant darkness (DD). c, Phase angle of the light entrainment in the last day of the light–dark conditions (n = 7 mice). d–i, Representative chi-square periodograms (d, e, h, i) and period length (f, g) for five-month-old mice in light–dark conditions or constant darkness (n = 7 mice). j, Average wheel-running activity in constant darkness after normalization to the intrinsic period (tau) (n = 7 mice). Data are mean ± s.e.m. *P <0.05 by two-sided t-test. Statistical details are in Supplementary Table 1.
Extended Data Fig. 2 Metabolic characterization of knockout mice on a normal chow diet.
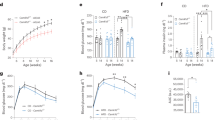
a, Daily food intake in three-month-old mice in home cages (n = 4 cages across 20 days). b, c, Food intake measured by the comprehensive laboratory animal monitoring system (CLAMS) in the 6 h (b) or 12 h (c) before GTT analyses at four months old (n = 5 mice). Box plot centre lines, box limits and whiskers represent the median, quartiles and minimum and maximum values, respectively. d, Blood glucose levels in four-month-old mice (n = 14 wild-type mice; n = 10 knockout mice). e, Serum insulin levels (n = 10 mice per group). f, Blood glucagon levels (n = 12 mice per group). g, Blood corticosterone levels (n = 11 mice per group). h, Blood GLP-1 levels (n = 12 wild-type mice; n = 10 knockout mice). i, Blood growth hormone (GH) levels (n = 11 wild-type mice; n = 12 knockout mice). j, k, GTTs in five-month-old mice at ZT6–8 (j) or ZT12–14 (k), with vgat-cre mice serving as the wild-type control (n = 7 mice). l, Body weight for clamp analyses at five months old (n = 4 mice). m, n, Blood glucose levels (m) and glucose infusion rate (n) during clamp analyses (n = 4 mice). o, Hyperinsulinaemia-mediated suppression of hepatic glucose production in the clamp analyses (n = 4 mice). Data are mean ± s.e.m. *P < 0.05 by two-way ANOVA or two-sided t-test. Statistical details are in Supplementary Table 1.
Extended Data Fig. 3 Metabolic characterization of knockout mice on a high-fat diet.
a, Body weight on HFD. HFD started at 10 months (n = 12 mice). b, GTTs at ZT6–8 after two weeks on HFD (n = 12 mice). c, GTTs at ZT12–14 after three weeks on HFD (n = 12 mice). d, ITTs at ZT6–8 after four weeks on HFD (n = 12 mice). e, ITTs at ZT12–14 after five weeks on HFD (n = 12 mice). f, Injection of streptozotocin (STZ) six weeks after HFD. g, h, Body weight (g) and blood glucose levels (h) at ZT10 after STZ injection (n = 12 mice). i, Blood glucose levels at the indicated ZTs two weeks after STZ injection, n = 12 mice. Data are mean ± s.e.m. *P < 0.05 by two-way ANOVA or two-sided t-test. Statistical details are in Supplementary Table 1.
Extended Data Fig. 4 Gene expression analysis of different brain regions.
a–e, RT–qPCR analysis of the indicated brain-region-specific marker genes (Vip (a), Pmch (b), Crf (c), Rfrp (also known as Npvf) (d) and Pomc1 (also known as Pomc) (e)) for brain regions isolated from both wild-type and knockout mice at ZT6 at the age of three months (n = 12 mice). Box plot centre lines, box limits and whiskers represent the median, quartiles and minimum and maximum values, respectively. f–i, RT–qPCR analysis comparing mRNA expression of Nr1d1 (f), Nr1d2 (g), Bmal1 (h) and Npas2 (i) in wild-type and knockout mice at ZT6 at the age of three months (n = 6 mice). Data are mean ± s.e.m. *P < 0.05 by two-way ANOVA or two-sided t-test. Statistical details are in Supplementary Table 1.
Extended Data Fig. 5 Electrophysiological and molecular characterization of knockout mice.
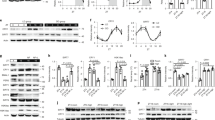
a–c, Brain slice patch-clamp representative traces for spontaneous firing (a), mEPSCs (b) and mIPSCs (c) at ZT12–14. d–g, Temporal pattern of expression of Rgs16 (d), α7-Takusan (Gm10406) (e), Nr1d1 (f) and Nr1d2 (g) in the hypothalamus in light–dark conditions from CircaDB (http://circadb.hogeneschlab.org). h–k, RT–qPCR analysis of the mRNA levels of Nr1d1 (h), Nr1d2 (i), Bmal1 (j) and Npas2 (k) in the SCN of three-month-old mice (n = 6 mice). Primers for Nr1d1 and Nr1d2 did not span the floxed exons. l, RNAscope of Rgs16 at the SCN in wild-type and knockout mice at the indicated ZTs. Scale bars, 100 μm. m, Quantification of Rgs16 staining (n = 5 wild-type mice at ZT4; n = 3 knockout mice at ZT4; n = 4 wild-type or knockout mice at ZT16). n, In situ hybridization analysis of Takusan Gm3500 staining. Scale bars, 25 μm. o, Quantification of in situ hybridization analysis of Takusan Gm3500 (n = 4 wild-type mice at ZT4; n = 6 wild-type mice at ZT16; n = 3 knockout mice at ZT4 or ZT16). p, Genome browser views of transcription start sites (green arrows) and nearby AGGTCA elements (red arrows) for the indicated genes on GRCm38. q, ChIP–qPCR analysis of Nr1d1 in the hypothalamus of three-month-old wild-type mice at ZT9 and ZT21—the peak and the trough of REV-ERB-α expression, respectively (n = 4 samples). Hypothalami from five mice were pooled as one sample. The negative control primers target a gene desert region on chromosome 17. The primer sequences of ChIP–qPCR assays are in Supplementary Table 6. Data are mean ± s.e.m. *P < 0.05 by two-way ANOVA or two-sided t-test. Statistical details are in Supplementary Table 1.
Extended Data Fig. 6 Metabolic characterization of mice overexpressing RGS16 or α7-Takusan in SCNGABA neurons.
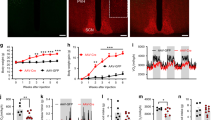
a, Validation of the injection with GFP fluorescence signals. Scale bar, 200 μm. b–e, GTTs (b, c) and ITTs (d, e) at the indicated ZTs in four-month-old vgat-cre mice injected with AAV-FLEX vectors encoding GFP, RGS16 or α7-Takusan (n = 7 mice). f, Body weight of three-month-old vgat-cre mice at three weeks after AAV injection (n = 14 mice). Data are mean ± s.e.m. *P < 0.05 for RGS16 or α7-Takusan versus the GFP control by two-way ANOVA followed by Holm–Sidak’s test. Statistical details are in Supplementary Table 1.
Extended Data Fig. 7 Rhythmicity of SCNGABA neuron firing in glucose metabolism.
a, Experimental design for chemogenetic activation of the SCNGABA neurons in wild-type mice with hM3Dq. b, Body weight of vgat-cre mice injected with AAV expressing hM3Dq or control mCherry (n = 11 mice). Mice were injected at the age of two months. c, Experimental design for chemogenetic repression of the SCNGABA neurons in wild-type and knockout mice with hM4Di. d, Body weight of wild-type and knockout mice injected with AAV expressing hM4Di (n = 12 wild-type mice; n = 14 knockout mice). Mice were injected at the age of two months. e, f, GTTs in wild-type or knockout mice injected with AAV expressing hM4Di at the indicated ZTs in the presence of saline (e) or CNO (f) (n = 12 wild-type mice; n = 14 knockout mice). Data are mean ± s.e.m. *P < 0.05 by two-sided t-test.
Extended Data Fig. 8 Rhythmicity of REV-ERB expression in SCNGABA neurons in glucose metabolism.
a, Experimental design for inducible re-expression of REV-ERB-α in the SCNGABA neurons of knockout mice. Mice were injected virus at the age of 2.5 months. b, Body weight at the time of euthanasia (n = 9 mice). c, d, GTTs in 4–4.5-month-old mice at ZT6–8 after injection of doxycycline at ZT0 (c) or ZT18 (d) (n = 9 mice). e, RT–qPCR analysis of the SCN from knockout mice with inducible re-expression of REV-ERB-α. Doxycycline was injected at ZT0 and the brain was collected at ZT12–14 (n = 4 mice). Data are mean ± s.e.m. *P < 0.05 by two-sided t-test. Statistical details are in Supplementary Table 1.
Extended Data Fig. 9 Assessment of CGM performance.
a, A representative comparison between fingertip glucometer reading and CGM reading for an individual at different times of the day. b, Pearson correlation coefficient between CGM and fingertip readings (n = 16 DP−; n = 11 DP+). c, MARD, the average of the absolute error between all CGM values and matched reference values (n = 16 DP−; n = 11 DP+). Data are mean ± s.e.m.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-6. Supplementary Table 1. Statistical tests. Statistical details were shown for results with significant differences, including the names of the statistical methods, p values, and sample sizes for those not included in the main figure legends due to limited space. Supplementary Table 2. Primer sequences for RT-qPCR and ChIP-qPCR. Nucleic acid sequences (from the 5’ end to the 3’ end) were shown for RT-qPCR primers, ChIP-qPCR primers, and the ISH probe. Supplementary Table 3. Inclusion and exclusion criteria for patient recruitment. Supplementary Table 4. Characteristics of human subjects. Supplementary Table 5. Cardiopulmonary Coupling-Polysomnography (CPC-PSG) of human subjects. Supplementary Table 6. Medication usage in human subjects.
Source data
Rights and permissions
About this article
Cite this article
Ding, G., Li, X., Hou, X. et al. REV-ERB in GABAergic neurons controls diurnal hepatic insulin sensitivity. Nature 592, 763–767 (2021). https://doi.org/10.1038/s41586-021-03358-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03358-w
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.