Abstract
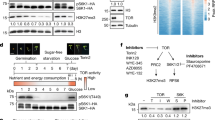
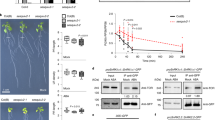
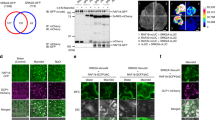
The evolutionarily conserved target of rapamycin (TOR) kinase acts as a master regulator that coordinates cell proliferation and growth by integrating nutrient, energy, hormone and stress signals in all eukaryotes1,2. Research has focused mainly on TOR-regulated translation, but how TOR orchestrates the global transcriptional network remains unclear. Here we identify ethylene-insensitive protein 2 (EIN2), a central integrator3,4,5 that shuttles between the cytoplasm and the nucleus, as a direct substrate of TOR in Arabidopsis thaliana. Glucose-activated TOR kinase directly phosphorylates EIN2 to prevent its nuclear localization. Notably, the rapid global transcriptional reprogramming that is directed by glucose–TOR signalling is largely compromised in the ein2-5 mutant, and EIN2 negatively regulates the expression of a wide range of target genes of glucose-activated TOR that are involved in DNA replication, cell wall and lipid synthesis and various secondary metabolic pathways. Chemical, cellular and genetic analyses reveal that cell elongation and proliferation processes that are controlled by the glucose–TOR–EIN2 axis are decoupled from canonical ethylene–CTR1–EIN2 signalling, and mediated by different phosphorylation sites. Our findings reveal a molecular mechanism by which a central signalling hub is shared but differentially modulated by diverse signalling pathways using distinct phosphorylation codes that can be specified by upstream protein kinases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The RNA-seq raw data for the glucose–EIN2-regulated transcriptome have been deposited in the NCBI Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) under the accession number: GSE114505. The RNA-seq raw data for the ethylene-regulated transcriptome can be downloaded from the NCBI SRA (https://www.ncbi.nlm.nih.gov/sra/) under the accession number: SRP017925. Source gel data for all immunoblots and radiograms (Figs. 1c–g, 2b, c, Extended Data Figs. 1f, 2c, 3b, k, 4b, 6e) are provided in Supplementary Fig. 1. Source data are provided with this paper.
References
Brunkard, J. O. Exaptive evolution of target of rapamycin signaling in multicellular eukaryotes. Dev. Cell 54, 142–155 (2020).
Liu, G. Y. & Sabatini, D. M. mTOR at the nexus of nutrition, growth, ageing and disease. Nat. Rev. Mol. Cell Biol. 21, 183–203 (2020).
Ju, C. et al. CTR1 phosphorylates the central regulator EIN2 to control ethylene hormone signaling from the ER membrane to the nucleus in Arabidopsis. Proc. Natl Acad. Sci. USA 109, 19486–19491 (2012).
Qiao, H. et al. Processing and subcellular trafficking of ER-tethered EIN2 control response to ethylene gas. Science 338, 390–393 (2012).
Wen, X. et al. Activation of ethylene signaling is mediated by nuclear translocation of the cleaved EIN2 carboxyl terminus. Cell Res. 22, 1613–1616 (2012).
Dobrenel, T. et al. TOR signaling and nutrient sensing. Annu. Rev. Plant Biol. 67, 261–285 (2016).
Fu, L., Wang, P. & Xiong, Y. Target of rapamycin signaling in plant stress responses. Plant Physiol. 182, 1613–1623 (2020).
Shi, L., Wu, Y. & Sheen, J. TOR signaling in plants: conservation and innovation. Development 145, dev160887 (2018).
Zhang, Z. et al. TOR signaling promotes accumulation of BZR1 to balance growth with carbon availability in Arabidopsis. Curr. Biol. 26, 1854–1860 (2016).
Li, X. et al. Differential TOR activation and cell proliferation in Arabidopsis root and shoot apexes. Proc. Natl Acad. Sci. USA 114, 2765–2770 (2017).
Xiong, Y. et al. Glucose-TOR signalling reprograms the transcriptome and activates meristems. Nature 496, 181–186 (2013).
Xiong, Y. & Sheen, J. Rapamycin and glucose-target of rapamycin (TOR) protein signaling in plants. J. Biol. Chem. 287, 2836–2842 (2012).
Ang, L. H. & Deng, X. W. Regulatory hierarchy of photomorphogenic loci: allele-specific and light-dependent interaction between the HY5 and COP1 loci. Plant Cell 6, 613–628 (1994).
Ma, D. et al. Cryptochrome 1 interacts with PIF4 to regulate high temperature-mediated hypocotyl elongation in response to blue light. Proc. Natl Acad. Sci. USA 113, 224–229 (2016).
Oh, E., Zhu, J. Y. & Wang, Z. Y. Interaction between BZR1 and PIF4 integrates brassinosteroid and environmental responses. Nat. Cell Biol. 14, 802–809 (2012).
Zhong, S. et al. Ethylene-orchestrated circuitry coordinates a seedling’s response to soil cover and etiolated growth. Proc. Natl Acad. Sci. USA 111, 3913–3920 (2014).
Alonso, J. M., Hirayama, T., Roman, G., Nourizadeh, S. & Ecker, J. R. EIN2, a bifunctional transducer of ethylene and stress responses in Arabidopsis. Science 284, 2148–2152 (1999).
Binder, B. M., Mortimore, L. A., Stepanova, A. N., Ecker, J. R. & Bleecker, A. B. Short-term growth responses to ethylene in Arabidopsis seedlings are EIN3/EIL1 independent. Plant Physiol. 136, 2921–2927 (2004).
An, F. et al. Ethylene-induced stabilization of ETHYLENE INSENSITIVE3 and EIN3-LIKE1 is mediated by proteasomal degradation of EIN3 binding F-box 1 and 2 that requires EIN2 in Arabidopsis. Plant Cell 22, 2384–2401 (2010).
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Nakano, T., Suzuki, K., Fujimura, T. & Shinshi, H. Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 140, 411–432 (2006).
Nelson, B. K., Cai, X. & Nebenführ, A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 51, 1126–1136 (2007).
Street, I. H. et al. Ethylene inhibits cell proliferation of the Arabidopsis root meristem. Plant Physiol. 169, 338–350 (2015).
Chang, K. N. et al. Temporal transcriptional response to ethylene gas drives growth hormone cross-regulation in Arabidopsis. eLife 2, e00675 (2013).
Zhang, J., Chen, Y., Lu, J., Zhang, Y. & Wen, C.-K. Uncertainty of EIN2Ser645/Ser924 inactivation by CTR1-mediated phosphorylation reveals the complexity of ethylene signaling. Plant Communications 1, 100046 (2020).
Gazzarrini, S. & McCourt, P. Cross-talk in plant hormone signalling: what Arabidopsis mutants are telling us. Ann. Bot. 91, 605–612 (2003).
Lorenzo, O., Piqueras, R., Sánchez-Serrano, J. J. & Solano, R. ETHYLENE RESPONSE FACTOR1 integrates signals from ethylene and jasmonate pathways in plant defense. Plant Cell 15, 165–178 (2003).
Peng, J. et al. Salt-induced stabilization of EIN3/EIL1 confers salinity tolerance by deterring ROS accumulation in Arabidopsis. PLoS Genet. 10, e1004664 (2014).
Zhuo, F., Xiong, F., Deng, K., Li, Z. & Ren, M. Target of rapamycin (TOR) negatively regulates ethylene signals in Arabidopsis. Int. J. Mol. Sci. 21, 2680 (2020).
Bleecker, A. B., Estelle, M. A., Somerville, C. & Kende, H. Insensitivity to ethylene conferred by a dominant mutation in Arabidopsis thaliana. Science 241, 1086–1089 (1988).
Mao, J. L. et al. Arabidopsis ERF1 mediates cross-talk between ethylene and auxin biosynthesis during primary root elongation by regulating asa1 expression. PLoS Genet. 12, e1005760 (2016).
Nakagawa, T. et al. Development of series of gateway binary vectors, pGWBs, for realizing efficient construction of fusion genes for plant transformation. J. Biosci. Bioeng. 104, 34–41 (2007).
Brand, L. et al. A versatile and reliable two-component system for tissue-specific gene induction in Arabidopsis. Plant Physiol. 141, 1194–1204 (2006).
Wang, J. & Chen, H. A novel CRISPR/Cas9 system for efficiently generating Cas9-free multiplex mutants in Arabidopsis. aBIOTECH 1, 6–14 (2020).
Kovtun, Y., Chiu, W. L., Zeng, W. & Sheen, J. Suppression of auxin signal transduction by a MAPK cascade in higher plants. Nature 395, 716–720 (1998).
Wiśniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Tsai, C. F. et al. Sequential phosphoproteomic enrichment through complementary metal-directed immobilized metal ion affinity chromatography. Anal. Chem. 86, 685–693 (2014).
Wang, P. et al. Reciprocal regulation of the TOR kinase and ABA receptor balances plant growth and stress response. Mol. Cell 69, 100–112 (2018).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Cho, Y. H., Yoo, S. D. & Sheen, J. Regulatory functions of nuclear hexokinase1 complex in glucose signaling. Cell 127, 579–589 (2006).
Kotogány, E., Dudits, D., Horváth, G. V. & Ayaydin, F. A rapid and robust assay for detection of S-phase cell cycle progression in plant cells and tissues by using ethynyl deoxyuridine. Plant Methods 6, 5 (2010).
Yoo, S. D., Cho, Y. H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protocols 2, 1565–1572 (2007).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Cox, M. P., Peterson, D. A. & Biggs, P. J. SolexaQA: at-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinformatics 11, 485 (2010).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Homer, N., Merriman, B. & Nelson, S. F. BFAST: an alignment tool for large scale genome resequencing. PLoS ONE 4, e7767 (2009).
Acknowledgements
We thank J. Sheen, T. D. Xu and X. Y. Zhu for critical reading of the manuscript, all members of the Xiong laboratory for discussion, C. B. Xiang for ERF1-OX seeds, W. F. Wang for the BZR1 antibody, R. X. Li for the CFP-HDEL plasmid and ABRC for the T-DNA insertional and point mutation mutants. This research was supported by funding from the National Natural Science Foundation of China (grant 31870269 to Y.X. and 31800199 to Yanlin Liu; the Chinese Academy of Sciences; and the Basic Forestry and Proteomics Research Centre, Haixia Institute of Science and Technology, Fujian Agriculture and Forestry University.
Author information
Authors and Affiliations
Contributions
Y.X., L.F., Yanlin Liu, S.-D.Y. and Y.-H.C. designed the experiments, and L.F. and Yanlin Liu performed most of the experiments and prepared the figures. G.Q., Yaxing Li, P.W., C.P., Z.Z. and C.C.L.W. carried out proteomics analyses. H.Z., Z.X. and R.L. conducted RNA-seq data analyses. X.Z., Y.W., S.Y. and L.F. generated the EIN2 transgenic lines. S.-D.Y. and Y.-H.C. provided ethylene-related mutants. Y.X., L.F. and Yanlin Liu wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Jose Alonso, Ronald Pierik and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Non-canonical role of EIN2, EIN3 and EIL1 in glucose–TOR signalling.
a, Quantification data of Fig. 1a. C2H4, ethylene. b, c, Glucose-promoted hypocotyl elongation depends on TOR kinase. Wild-type and tor-es lines were germinated for 4 days in the dark without or with glucose (Glc, 0.5% w/v). Representative images (b) and hypocotyl length (c). Scale bar, 1 mm. d, e, Glucose–TOR-regulated hypocotyl elongation is independent of ethylene sensing and biosynthesis. The tor-es seedlings were germinated in the dark without or with oestradiol (Est), 1-MCP (100 nl l−1) or AVG (1 μM) for 4 days. Representative images (d) and hypocotyl length (e). Scale bar, 1 mm. f, Oestradiol induces TOR silencing, as revealed by western blot. Experiments were repeated three times with similar results. g, h, Representative images (g) and hypocotyl length (h) of wild-type (Ler) and gin2 seedlings germinated in the dark with or without various chemical treatments for 4 days. Scale bar, 1 mm. i, Quantification data of Fig. 1b. j, Quantification data of Fig. 1c. n = 3 biological repeats. k, l, Representative images (k) and hypocotyl length (l) of wild-type (Col-0) and mutant seedlings germinated in the dark with or without various chemical treatments for 4 days. Scale bars, 1 mm. Data were analysed from 45 (a, c, i), 35 (e), 30 (h) or 52 (l) seedlings for each genotype and treatment from 3 experiments, and expressed as mean ± s.d. Statistical significance was determined by one-way ANOVA with Tukey’s post hoc test (a, e, h, i, j, l) or by two-sided Student’s t-test (c); n.s. not significant; P values are indicated. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Different dosage response curves between wild-type, etr1-1, ein2-5 and ein3 eil1 in response to rapamycin, Torin2 and AZD8055.
a, b, Representative images (a) and hypocotyl length (b) of wild type (Col-0) and mutant seedlings germinated in the dark with different concentration of rapamycin, Torin2 or AZD8055 for 4 days. Scale bars, 1 mm. Data were analysed from 30 (rapamycin and Torin2 treatment) or 90 (AZD8055 treatment) seedlings for each genotype and treatment from 3 experiments, and are expressed as mean ± s.d. Statistical significance was determined by two-sided Student’s t-test; P values are indicated. c, Rapamycin, Torin2 or AZD8055 inhibits TOR kinase activity in a concentration-dependent manner. p35S::S6K1-HA transgenic seedlings were germinated with different concentrations of rapamycin, Torin2 or AZD8055 in the dark for 4 days. TOR kinase activities were analysed based on phosphorylation of the conserved TOR substrate (pThr449 of S6K1) by western blot. Experiments were repeated three times with similar results. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 3 Glucose–TOR–EIN2 regulates EIN3 stability and the expression of ERFs to control hypocotyl elongation.
a, TOR interacts with EIN2-C, as revealed by yeast two-hybrid analyses. b, Inactivation of TOR triggers EIN3 accumulation depending on EIN2. p35S::EIN3-GFP/ein3 eil1 and p35S::EIN3-GFP/ein2-5 seedlings were treated with rapamycin (10 μM) or ACC (10 μM) for the indicated times. Tubulin (TUB) was used as the loading control. c–e, Ethylene (C2H4) and Torin2 activate the expression of both overlapped and distinct ERF genes. The 4-d-etiolated wild-type seedlings (c, d) or mutants (e) were treated with ethylene (c) or Torin2 (d, e) for 2 h, then the hypocotyls were collected for RT–qPCR analyses. Data are mean ± s.d.; n = 3 biological repeats. f, g, Representative images (f) and hypocotyl length (g) of wild-type (Col-0) and ERF1-OX seedlings germinated in the dark with different concentrations of Torin2 for 4 days. Scale bar, 1 mm. h, erf-c mutant sequences. The erf-c mutants were generated by CRISPR–Cas9 technology. Green arrows indicate the position of the deletion or insertion of a nucleotide; red letters indicate amino acids resulting from the frameshift translation; * indicates stop codon. i, j, Representative images (i) and hypocotyl length (j) of wild-type (Col-0) and erf-c seedlings germinated in the dark with or without Torin2 (1 μM) for 4 days. Scale bar, 1 mm. k, l, TOR regulation of BZR1 stability is independent from the EIN2–EIN3/EIL1 axis. The 3-d-etiolated seedlings were treated with Torin2 for 2 days. Tubulin was used as the loading control. Representative images (k) and relative BZR1 abundance (l). Data are mean ± s.d., n = 3 biological repeats. m, The non-canonical role of EIN2 in glucose–TOR-signalling-mediated hypocotyl elongation. In g, j, data were analysed from 45 seedlings for each genotype and treatment from 3 experiments, and are expressed as mean ± s.d. Statistical significance was determined by one-way ANOVA with Tukey’s post hoc test (g) or by two-sided Student’s t-test (c, d, e, j, l); n.s. not significant; P values are indicated. In a, b, experiments were repeated three times with similar results. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 4 Glucose–TOR and ethylene–CTR1 regulate the nuclear localization of EIN2 through distinct phosphorylation sites.
a, Thr657 is conserved among EIN2 orthologues of A. thaliana, O. sativa and Z. mays. b, TOR kinase directly phosphorylates OsEIN2-C and ZmEIN2-C, as shown by an in vitro kinase assay. Torin2, 1 μM. Experiments were repeated three times and similar results were obtained. c–j, Different nucleus accumulation patterns and quantification of relative GFP fluorescent signal intensity between the nucleus and the cytosol of OsEIN2–GFP and ZmEIN2–GFP (c, d), EIN2(WT)–GFP (e, f), EIN2(T657A)–GFP and EIN2(T657D)–GFP (g, h), and EIN2(S645D)–GFP and EIN2(S924D)–GFP (i, j) in Arabidopsis leaf mesophyll protoplasts. Mesophyll protoplasts transiently transfected with various EIN2 constructs were treated with or without rapamycin (10 μM) or ACC (10 μM) for 3 h. Scale bars, 5 μm. NLS-mC denotes nuclear HY5–mCherry. In d, f–h data were analysed from the indicated numbers of cells from three experiments, and are expressed as mean ± s.d. Centre line, median; box limits, 25th and 75th percentiles; whiskers extend to 1.5 × IQR from the 25th and 75th percentiles. Statistical significance was determined by one-way ANOVA with Tukey’s post hoc test; n.s, not significant; P values are indicated. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 5 Glucose–TOR- and ethylene signalling regulate hypocotyl elongation through distinct phosphorylation sites in EIN2.
a, b, Different patterns of nuclear accumulation of EIN2(WT)–GFP, EIN2(T657D)–GFP and EIN2(S645D)–GFP in hypocotyl cells. Seedlings were germinated in the dark for 4 days, then treated with Torin2 (1 μM) or ethylene (C2H4, 100 ppm) (a), or rapamycin (10 μM) (b) for 4 h. Arrows indicate nuclei. Scale bars, 10 μm. c, Quantification of the relative GFP fluorescent signal intensity between the nucleus and the cytosol of EIN2(WT)–GFP, EIN2(T657D)–GFP and EIN2(S645D)–GFP in b. See quantification data of a in Fig. 2e. d, Quantification data of Fig. 2f. e, f, EIN2(S645D)–GFP but not EIN2(T657D)–GFP restores ein2-5 sensitivity to rapamycin-inhibited hypocotyl elongation. Seedlings were germinated in the dark with or without rapamycin (10 μM) for 4 days. Representative images (e) and hypocotyl length (f). Scale bar, 1 mm. In c, data were analysed from the indicated numbers of hypocotyl cells from three experiments, and are expressed as mean ± s.d. Centre line, median; box limits, 25th and 75th percentiles; whiskers extend to 1.5 × IQR from the 25th and 75th percentiles. In d, f, data were analysed from 45 (d) and 40 (f) seedlings for each genotype and treatment from three experiments, and are expressed as mean ± s.d. Statistical significance was determined by two-sided Student’s t-test (c) or by one-way ANOVA with post hoc Tukey test (d, f); n.s, not significant; P values are indicated.
Extended Data Fig. 6 Full-length EIN2 is translocated to the nucleus in response to inhibition of TOR.
a, b, EIN2–GFP and GFP–EIN2 complement the response of ein2-5 seedlings to treatment with Torin2 and ethylene (C2H4). Seedlings were germinated on solid half-strength MS medium with or without Torin2 (1 μM) or ethylene (100 ppm) in the dark for 4 days. Representative images (a) and hypocotyl length (b). Scale bar, 1 mm. Data were analysed from 22 seedlings for each genotype and treatment from 3 experiments, and are expressed as mean ± s.d. c, Different nucleus accumulation patterns of EIN2(WT)–GFP and GFP–EIN2(WT); see quantification data in Fig. 2g. Scale bars, 500 μm (left); 10 μm (right). d, EIN2 accumulates in the nucleus after inhibition of TOR. Arabidopsis mesophyll protoplasts transiently co-transfected with mCherry–EIN2–GFP and CFP–HDEL (an ER marker) were treated with or without rapamycin (10 μM) or Torin2 (1 μM) for 3 h. Scale bars, 5 μm. Experiments were repeated three times with similar results. e, f, Full-length EIN2 is translocated to the nucleus in response to inhibition of TOR. Four-day-etiolated pEIN2::EIN2WT-3XHA seedlings were treated with or without Torin2 (1 μM) for the indicated times, then total and nuclear fractions of hypocotyl protein were prepared and subjected to western blot (e). Tubulin (total protein loading control), calnexin homologue 1 or 2 (CNX1/2, ER marker) and histone H3 (nuclear marker) were used as controls to monitor the loading and purity of the total and nuclear fraction. f, Quantification of e. Data are mean ± s.d., n = 3 biological repeats. For b, f, statistical significance was determined by one-way ANOVA with Tukey’s post hoc test; n.s, not significant; P values are indicated. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 7 Role of EIN2 in glucose–TOR-mediated activation of the root meristem.
a, Inhibition of TOR triggers nuclear localization of EIN2–GFP in root meristem cells. Seedlings (4 DAG) were treated with Torin2 (1 μM) for 4 h. Arrows indicate nuclei. Scale bars, 25 μm (left); 10 μm (right). See Fig. 3a for quantification data. b–f, EIN2 is involved in glucose-promoted primary root growth (b, d) and cell proliferation activity indicated by EdU staining (c, e) and S-phase gene expression (f; RT–qPCR analysis; mean ± s.d., n = 3 biological repeats). Scale bars, 1 mm (b); 20 μm (c). g, h, Quantification data of Fig. 3f, g. i, EIN2 Thr657 is required for glucose-promoted S-phase gene expression. RT–qPCR analysis; mean ± s.d.; n = 3 biological repeats. For d, e, g, h, data were analysed from 45 (d), 30 (e), 45 (g) and 30 (h) of roots for each genotype from 3 experiments, and are expressed as mean ± s.d. In all of the bar charts, statistical significance was determined by two-sided Student’s t-test; n.s, not significant; P values are indicated.
Extended Data Fig. 8 EIN2(T657D) and EIN2(S645D) distinguish ethylene- and glucose–TOR-regulated cell proliferation.
a, b, EIN2(T657D) and EIN2(S645D) distinguish ethylene-regulated cell proliferation, indicated by EdU staining of the root apical meristem. Scale bar, 50 μm. Seedlings were germinated in glucose-free liquid half-strength MS medium for 4 days, then transferred to solid half-strength MS medium with glucose (15 mM) and ethylene (C2H4, 100 ppm) for 4 h. Representative EdU staining images (a) and quantification of EdU intensity (b). c–f, EIN2(T657D) and EIN2(S645D) distinguish glucose-TOR-regulated cell proliferation. Scale bars, 50 μm. Seedlings were germinated in glucose-free liquid half-strength MS medium for 4 days, then treated with glucose (15 mM) and Torin2 (1 μM) for 2 h. Representative EdU staining images and quantification of the EdU intensity of ethylene-insensitive mutants (c, d) and pEIN2::EIN2WT-GFP/ein2-5, pEIN2::EIN2T657-GFP/ein2-5 and pEIN2::EIN2S645D-GFP/ein2-5 transgenic plants (e, f). For b, d, f, data were analysed from 30 roots for each genotype and treatment from 3 experiments, and are expressed as mean ± s.d. Statistical significance was determined by one-way ANOVA with Tukey’s post hoc test (b) or by two-sided Student’s t-test (d, f); n.s, not significant; P values are indicated.
Extended Data Fig. 9 E2Fa acts downstream of EIN2 to promote primary root growth.
a, RT–qPCR analyses of E2Fa expression in glucose-depleted roots of wild-type (Col-0), ein2-5, ein3 eil1 and etr1-1 seedlings. Data are mean ± s.d.; n = 3 biological repeats. b–g, Overexpression of E2Fa in oestradiol-inducible E2Fa-GFP transgenic seedlings (b) resulted in longer roots (c, e), increased cell proliferation activities indicated by EdU staining (d, f) and S-phase gene expression revealed by RT–qPCR analysis (g; mean ± s.d.; n = 3 biological repeats). Scale bars, 20 μm (b); 1 mm (c); 25 μm (d). Data were analysed from 45 (#1 −Est), 43 (#1 +Est), 38 (#2 −Est) and 37 (#2 +Est) roots (e) or from 38 (#1 −Est), 38 (#1 +Est), 34 (#2 −Est) and 33 (#2 +Est) roots (f) from 3 experiments, and are expressed as mean ± s.d. In all of the bar charts, statistical significance was determined by two-sided Student’s t-test; P values are indicated.
Extended Data Fig. 10 The glucose–TOR–EIN2 axis mediates nuclear signalling to modulate plant growth.
a, RT–qPCR analyses of transcriptome variation in response to TOR inhibition and ethylene treatments. Wild-type seedlings and various mutants and transgenic lines were germinated in 0.5% glucose (w/v) for 4 days, then treated with or without Torin2 (1 μM) or ethylene (C2H4, 100 ppm) for 2 h. The numbers in the heat map indicate the relative fold changes. b, Representative images and hypocotyl length of wild type (Col-0) and various mutants germinated in the dark with or without AZD8055 (1 μM) or Torin2 (1 μM) for the indicated times. Scale bars, 1 mm. Data were analysed from 30 seedlings for each genotype and treatment from 3 experiments, and are expressed as mean ± s.d. Statistical significance was determined by two-sided Student’s t-test; n.s, not significant; P values are indicated. c, Proposed model for the glucose–TOR–EIN2 axis in plant growth and development. Glucose-activated TOR directly phosphorylates EIN2 to inhibit its nuclear shuttling. In glucose-depleted conditions, dephosphorylated EIN2 moves into the nucleus, and controls root meristem activation and hypocotyl cell elongation through different downstream effectors.
Supplementary information
Supplementary Figure
This file contains Supplementary Figure 1, the source gel data.
Supplementary Table 1
Glucose-controlled genes. The data includes both glucose up-and down-regulated (FDR<0.01) genes, which are visualized in Fig. 3b-e.
Supplementary Table 2
Glucose-EIN2 controlled genes. The data includes glucose insensitive and less sensitive genes in ein2-5, which are visualized in Fig. 3b-e.
Supplementary Table 3
Differential regulated genes between ein2-5 and WT under glucose-depletion condition, which are visualized in Fig. 3b-e.
Supplementary Table 4
Transcriptome comparison between glucose/ethylene-regulated genes. The data includes ethylene specific, glucose/TOR specific, and ethylene/glucose common-regulated genes, which are visualized in Fig. 3h.
Supplementary Table 5
Primers used for plasmid constructs, RT-qPCR and mutagenesis.
Source data
Rights and permissions
About this article
Cite this article
Fu, L., Liu, Y., Qin, G. et al. The TOR–EIN2 axis mediates nuclear signalling to modulate plant growth. Nature 591, 288–292 (2021). https://doi.org/10.1038/s41586-021-03310-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03310-y
This article is cited by
-
Transcription factor TERF1 promotes seed germination through HEXOKINASE 1 (HXK1)-mediated signaling pathway
Journal of Plant Research (2023)
-
Don’t TOR at the TOP
Nature Plants (2022)
-
TOP1α fine-tunes TOR-PLT2 to maintain root tip homeostasis in response to sugars
Nature Plants (2022)
-
TOR signaling is the potential core of conserved regulation of trichome development in plant
Acta Physiologiae Plantarum (2022)
-
Development of potent promoters that drive the efficient expression of genes in apple protoplasts
Horticulture Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.