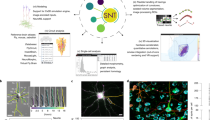
Abstract
Animal nervous system organization is crucial for all body functions and its disruption can lead to severe cognitive and behavioural impairment1. This organization relies on features across scales—from the localization of synapses at the nanoscale, through neurons, which possess intricate neuronal morphologies that underpin circuit organization, to stereotyped connections between different regions of the brain2. The sheer complexity of this organ means that the feat of reconstructing and modelling the structure of a complete nervous system that is integrated across all of these scales has yet to be achieved. Here we present a complete structure–function model of the main neuropil in the nematode Caenorhabditis elegans—the nerve ring—which we derive by integrating the volumetric reconstructions from two animals with corresponding3 synaptic and gap-junctional connectomes. Whereas previously the nerve ring was considered to be a densely packed tract of neural processes, we uncover internal organization and show how local neighbourhoods spatially constrain and support the synaptic connectome. We find that the C. elegans connectome is not invariant, but that a precisely wired core circuit is embedded in a background of variable connectivity, and identify a candidate reference connectome for the core circuit. Using this reference, we propose a modular network architecture of the C. elegans brain that supports sensory computation and integration, sensorimotor convergence and brain-wide coordination. These findings reveal scalable and robust features of brain organization that may be universal across phyla.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The volumetric datasets generated during the current study, associated connectivity databases and associated analysis are available at https://doi.org/10.5281/zenodo.4383277 and http://wormwiring.org/. The raw data for volumetric reconstructions for Figs. 1, 3, Extended Data Fig. 8 and all Supplementary Videos are available at https://doi.org/10.5281/zenodo.4383277. Extracted adjacency data are available in Supplementary Data 1. The reference datasets are available in Supplementary Data 3. The Cytoscape files used to generate the brain map (Fig. 4, Extended Data Fig. 10) and network motifs (Extended Data Fig. 10) are available at https://doi.org/10.5281/zenodo.4383277. The collection of C. elegans nervous system electron micrographs is also available at https://www.wormatlas.org/ and https://wormimage.org. Source data are provided with this paper.
Code availability
The software packages parsetrakem2 (extracting adjacency data) and elegansbrainmap (analysis and visualization software) are available at https://github.com/cabrittin/parsetrakem2 and https://github.com/cabrittin/elegansbrainmap, respectively.
References
Hahamy, A., Behrmann, M. & Malach, R. The idiosyncratic brain: distortion of spontaneous connectivity patterns in autism spectrum disorder. Nat. Neurosci. 18, 302–309 (2015).
Swanson, L. W. & Lichtman, J. W. From Cajal to connectome and beyond. Annu. Rev. Neurosci. 39, 197–216 (2016).
Cook, S. J. et al. Whole-animal connectomes of both Caenorhabditis elegans sexes. Nature 571, 63–71 (2019).
Ryan, K., Lu, Z. & Meinertzhagen, I. A. The CNS connectome of a tadpole larva of Ciona intestinalis (L.) highlights sidedness in the brain of a chordate sibling. eLife 5, e16962 (2016).
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Phil. Trans. R. Soc. B 314, 1–340 (1986).
Hall, D. H. & Russell, R. L. The posterior nervous system of the nematode Caenorhabditis elegans: serial reconstruction of identified neurons and complete pattern of synaptic interactions. J. Neurosci. 11, 1–22 (1991).
Jarrell, T. A. et al. The connectome of a decision-making neural network. Science 337, 437–444 (2012).
Bumbarger, D. J., Riebesell, M., Rödelsperger, C. & Sommer, R. J. System-wide rewiring underlies behavioral differences in predatory and bacterial-feeding nematodes. Cell 152, 109–119 (2013).
Ohyama, T. et al. A multilevel multimodal circuit enhances action selection in Drosophila. Nature 520, 633–639 (2015).
Zheng, Z. et al. A complete electron microscopy volume of the brain of adult Drosophila melanogaster. Cell 174, 730–743 (2018).
Kasthuri, N. et al. Saturated reconstruction of a volume of neocortex. Cell 162, 648–661 (2015).
Motta, A. et al. Dense connectomic reconstruction in layer 4 of the somatosensory cortex. Science 366, eaay3134 (2019).
Varshney, L. R., Chen, B. L., Paniagua, E., Hall, D. H. & Chklovskii, D. B. Structural properties of the Caenorhabditis elegans neuronal network. PLOS Comput. Biol. 7, e1001066 (2011).
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev. Biol. 100, 64–119 (1983).
Barabási, D. L. & Barabási, A.-L. A genetic model of the connectome. Neuron 105, 435–445 (2020).
Albertson, D. G. & Thomson, J. N. The pharynx of Caenorhabditis elegans. Phil. Trans. R. Soc. B 275, 299–325 (1976).
Cook, S. J. et al. The connectome of the Caenorhabditis elegans pharynx. J. Comp. Neurol. 528, 2767–2784 (2020).
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. Factors that determine connectivity in the nervous system of Caenorhabditis elegans. Cold Spring Harb. Symp. Quant. Biol. 48, 633–640 (1983).
Durbin, R. M. Studies on the Development and Organisation of the Nervous System of Caenorhabditis elegans. PhD thesis, Univ. Cambridge (1987).
Witvliet, D. et al. Connectomes across development reveal principles of brain maturation in C. elegans. Preprint at https://doi.org/10.1101/2020.04.30.066209 (2020).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008 (2008).
Gray, J. M., Hill, J. J. & Bargmann, C. I. A circuit for navigation in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 102, 3184–3191 (2005).
Kato, S. et al. Global brain dynamics embed the motor command sequence of Caenorhabditis elegans. Cell 163, 656–669 (2015).
Towlson, E. K., Vértes, P. E., Ahnert, S. E., Schafer, W. R. & Bullmore, E. T. The rich club of the C. elegans neuronal connectome. J. Neurosci. 33, 6380–6387 (2013).
Cohen, N. & Denham, J. E. Whole animal modeling: piecing together nematode locomotion. Curr. Opin. Syst. Biol. 13, 150–160 (2019).
Milo, R. et al. Network motifs: simple building blocks of complex networks. Science 298, 824–827 (2002).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (IEEE, 2016).
Thomson, A. M. Neocortical layer 6, a review. Front. Neuroanat. 4, 13 (2010).
Rapti, G., Li, C., Shan, A., Lu, Y. & Shaham, S. Glia initiate brain assembly through noncanonical Chimaerin-Furin axon guidance in C. elegans. Nat. Neurosci. 20, 1350–1360 (2017).
Morgan, J. L. & Lichtman, J. W. An individual interneuron participates in many kinds of inhibition and innervates much of the mouse visual thalamus. Neuron 106, 468–481 (2020).
Chen, X. et al. Brain-wide organization of neuronal activity and convergent sensorimotor transformations in larval zebrafish. Neuron 100, 876–890.e5 (2018).
Stern, S., Kirst, C. & Bargmann, C. I. Neuromodulatory control of long-term behavioral patterns and individuality across development. Cell 171, 1649–1662.e10 (2017).
Wang, L. & Marquardt, T. What axons tell each other: axon-axon signaling in nerve and circuit assembly. Curr. Opin. Neurobiol. 23, 974–982 (2013).
Moyle, M. W. et al. Structural and developmental principles of neuropil assembly in C. elegans. Nature https://doi.org/10.1038/s41586-020-03169-5 (2021).
Ware, R. W., Clark, D., Crossland, K. & Russell, R. L. The nerve ring of the nematode Caenorhabditis elegans: sensory input and motor output. J. Comp. Neurol. 162, 71–110 (1975).
Peachey, L. D. Thin sections. I. A study of section thickness and physical distortion produced during microtomy. J. Biophys. Biochem. Cytol. 4, 233–242 (1958).
Cardona, A. et al. TrakEM2 software for neural circuit reconstruction. PLoS One 7, e38011 (2012).
Xu, M. et al. Computer assisted assembly of connectomes from electron micrographs: application to Caenorhabditis elegans. PLoS One 8, e54050 (2013).
Newman, M. E. & Girvan, M. Finding and evaluating community structure in networks. Phys. Rev. E 69, 026113 (2004).
Rosvall, M. & Bergstrom, C. T. Maps of random walks on complex networks reveal community structure. Proc. Natl Acad. Sci. USA 105, 1118–1123 (2008).
Csardi, G. C. & Nepusz, T. The igraph software package for complex network research. InterJournal Complex Systems 1695 (2006).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Chang, A. J., Chronis, N., Karow, D. S., Marletta, M. A. & Bargmann, C. I. A distributed chemosensory circuit for oxygen preference in C. elegans. PLoS Biol. 4, e274 (2006).
Zimmer, M. et al. Neurons detect increases and decreases in oxygen levels using distinct guanylate cyclases. Neuron 61, 865–879 (2009).
Tomioka, M. et al. The insulin/PI 3-kinase pathway regulates salt chemotaxis learning in Caenorhabditis elegans. Neuron 51, 613–625 (2006).
Hendricks, M., Ha, H., Maffey, N. & Zhang, Y. Compartmentalized calcium dynamics in a C. elegans interneuron encode head movement. Nature 487, 99–103 (2012).
Perkins, L. A., Hedgecock, E. M., Thomson, J. N. & Culotti, J. G. Mutant sensory cilia in the nematode Caenorhabditis elegans. Dev. Biol. 117, 456–487 (1986).
Sawin, E. R., Ranganathan, R. & Horvitz, H. R. C. elegans locomotory rate is modulated by the environment through a dopaminergic pathway and by experience through a serotonergic pathway. Neuron 26, 619–631 (2000).
Kang, L., Gao, J., Schafer, W. R., Xie, Z. & Xu, X. Z. C. elegans TRP family protein TRP-4 is a pore-forming subunit of a native mechanotransduction channel. Neuron 67, 381–391 (2010).
Chalfie, M. & Sulston, J. Developmental genetics of the mechanosensory neurons of Caenorhabditis elegans. Dev. Biol. 82, 358–370 (1981).
Suzuki, H. et al. In vivo imaging of C. elegans mechanosensory neurons demonstrates a specific role for the MEC-4 channel in the process of gentle touch sensation. Neuron 39, 1005–1017 (2003).
Chalfie, M. et al. The neural circuit for touch sensitivity in Caenorhabditis elegans. J. Neurosci. 5, 956–964 (1985).
Li, C. et al. The FMRFamide-related neuropeptide FLP-20 is required in the mechanosensory neurons during memory for massed training in C. elegans. Learn. Mem. 20, 103–108 (2013).
Hukema, R. K., Rademakers, S., Dekkers, M. P. J., Burghoorn, J. & Jansen, G. Antagonistic sensory cues generate gustatory plasticity in Caenorhabditis elegans. EMBO J. 25, 312–322 (2006).
Acknowledgements
We thank J. Hodgkin and J. White for their help in donating archival transmission electron microscopy material from the MRC Laboratory of Molecular Biology to the Hall laboratory for curation. T. Ilett, F. Salfelder and S. L. Braunstein provided useful discussion. We thank M. Zhen for making their synaptic and gap junction data available (https://nemanode.org/). This work was supported by NIH grant NIMH F32MH115438 (S.J.C.), NIHD grant P30HD071593 (S.W.E.), NIMH grant R01MH112689 (S.W.E.), the G. Harold and Leila Y. Mathers Charitable Foundation (S.W.E.), NIH OD 010943 (D.H.H.) and EPSRC EP/J004057/1 (N.C.). C.A.B. was supported by the Leeds International Research Scholarship.
Author information
Authors and Affiliations
Contributions
C.A.B., S.J.C. and S.W.E. conceived the volumetric reconstruction. C.A.B. and S.J.C. segmented the electron micrographs. D.H.H. curated the data. C.A.B. built the software for quantifying membrane contact areas. C.A.B. and N.C. analysed and interpreted the data and wrote the manuscript. S.J.C., D.H.H. and S.W.E. provided critical revisions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Neuron neighbourhoods are bilaterally conserved in size, composition and membrane contact positions.
a, Variability in immediate neighbourhood size (adjacency degree) does not vary with immediate neighbourhood size. Immediate neighbourhood sizes for each neuron in each dataset (adult left, adult right, L4 left, L4 right, n = 80 bilateral cell classes common to L4 and adult) plotted against the immediate neighbourhood size of the corresponding neuron in the adult left. The inset shows the immediate neighbourhood size difference between homologous left and right neurons (vertical spread) as a function of neighbourhood size for the L4 (red) and adult (blue) nerve ring. b, The distributions of immediate neighbourhood size differences between homologous contralateral neurons in the same animal—adult left and right (L/R) and L4 L/R— are statistically indistinguishable from 0 (P values by two-sided Wilcoxon signed-rank test: 0.07 and 0.29, respectively; n = 80 cell classes). Immediate neighbourhood size differences between homologous adult and L4 neurons on the same side of the body are statistically distinguishable from 0 (P = 9.2 × 10−11 by two-sided Wilcoxon signed-rank test; n = 160 cells, but the difference is small—mean degree difference 3.6). c, Similarity between immediate neighbourhood compositions as quantified by the Jaccard index (Supplementary Results), shows higher compositional similarity between homologous contralateral neighbourhoods (n = 80 cell classes) than between proximal ipsilateral neighbourhoods (Supplementary Results; n = 160 cells). d–f, Membrane contact placement along processes is highly reproducible bilaterally and across the adult and L4 datasets. For each process, we mapped each M4 contact to a position along the anterior–posterior (AP) axis, \(\hat{z}\) (see Methods and Supplementary Results). For each M4 contact, we then counted the number of datasets in which the contact was observed at a given \(\hat{z}\) (reproducibility count). d, Demonstration of reproducibility count for a single cell class (RIA): RIA has the longest process in the nerve ring and among the highest average reproducibility counts. Shown is a raster plot of reproducibility counts as a function of \(\hat{z}\), of all M4 contacts made with RIA. Neighbouring processes: rows in alphabetical order. Colour: reproducibility count. We define the maximum spatial reproducibility count, \(\max \,{(\delta )}_{\hat{z}}\), as the highest reproducibility count across all locations, \(\hat{z}\), per cell pair (that is, for every row in the raster). For rasters of all other cell classes, see Supplementary Data 2. e, Fraction of M4 membrane contact sites co-localized in δ datasets (distribution over n = 80 cell classes). f, For each cell class, the fraction of membrane contacts achieved with a maximum spatial reproducibility count, \(\max \,{(\delta )}_{\hat{z}}\) (distribution over n = 80 cell classes). g, h, Comparatively, ℂ4 synaptic contact placement is less reproducible than physical adjacency. For each process, we mapped each ℂ4 contact along the AP axis, \(\hat{z}\). g, Demonstration of synaptic spatial reproducibility count for RIA neurons. h, For each cell class, the fraction of ℂ4 synaptic contacts achieved with a maximum spatial reproducibility count, \(\max \,{(\delta )}_{\hat{z}}\) (distribution over n = 80 cell classes). Box plots: centre line, median; box limits, upper and lower quartiles; whiskers, 1.5 × interquartile range; points, outliers.
Extended Data Fig. 2 Contact sizes and reproducibility.
a–f, Small membrane contact areas are less likely to be bilaterally conserved. Membrane contacts were divided into three groups (‘low’, ‘mid’ and ‘high’) on the basis of their membrane contact areas (35% low, 31% mid, 34% high; see Supplementary Results). a, Similarity of homologous (L4 bilateral; adult bilateral; L4 and adult—same side) immediate neighbourhood compositions for low, mid and high membrane contact groups, as measured by the Jaccard index (Supplementary Results; n = 80 cell classes). Box plots: centre line, median; box limits, upper and lower quartiles; whiskers, 1.5 × interquartile range; points, outliers. b, c, Survival (that is, complementary cumulative) distribution of membrane contacts in the adult nerve ring (b, n = 5,179) and the L4 nerve ring (c, n = 4,744). The pie charts show the fraction of total membrane area contact between all processes accounted for by each group. d, Empirical frequency distribution of synaptic (n = 2,433) and gap-junctional (n = 573) contacts broken down by the reproducibility of membrane contacts. The majority of synaptic contacts (77% and 85% of synaptic and gap-junction contacts, respectively) occur at M4 contacts. e, f, Cumulative distribution of ℂδ synaptic contacts (e) and Gδ gap-junction contacts (f) for δ = 1, 2, 3, 4 as a function of membrane contact area (in percentiles). To control for differences in neurite placement, we restrict ℂδ and Gδ to contacts that occur on M4 membrane contacts. The smallest 35% of membrane contacts (dashed line) encompass around 3% of ℂ4 synaptic contacts and around 9% of G4 gap-junction contacts (on M4) with growing fractions for smaller δ (up to around 33% and around 27% of the more variable ℂ1 and G1 contacts). g, Empirical frequency distribution of membrane, synaptic and gap-junctional contacts across the four datasets (δ = 1 to 4). h–j, Survival distribution of contacts as a function of membrane contact area for Mδ (h), ℂδ (i) and Gδ (j) graphs (n given in g), plotting the probability that a membrane, synaptic, or gap-junction contact occurs with a membrane contact area that exceeds some value. Membrane contact areas have been log-normalized and standardized so that the distribution is centred about 0, that is, log-transformed, standardized (by subtracting the mean) and normalized (by dividing by the standard deviation), such that a range of ±1 corresponds to ±1 standard deviation of the distribution of log(membrane contact area).
Extended Data Fig. 3 Core and variable model validations.
a, b, Model fits for the reproducibility of Mδ, ℂδ and Gδ contacts, with membrane contact areas below (a) and above (b) the log-normalized mean (after thresholding; see Methods, Extended Data Fig. 2h). c, d, Reproducibility model fits of inter-cluster (c) and intra-cluster (d) Mδ, ℂδ and Gδ contacts. e, Reproducibility model fits for the complete Mδ, ℂδ and Gδ sets including contacts with the smallest 35% of membrane contact areas (results qualitatively similar to restricted dataset model fit in Fig. 2a; see Methods, ‘Generating reference graphs’). f, Reproducibility model fits for ℂδ excluding synaptic contacts scored in only one electron micrograph (Methods). g, Reproducibility model fits for ℂδ excluding synaptic contacts derived from non-reproducible postsynaptic partners of polyadic synapses (Methods). h, i, Reproducibility model fits for synaptic and gap-junction contact datasets scored by White et al.5 (h) and Witvliet et al.20 (i) limited to our M4 contacts. Black bars, empirical distributions used in this study; grey bars, other empirical distributions5,20; red bars, model fits for the empirical distributions. All data are presented as fractions of the empirical counts (n).
Extended Data Fig. 4 Validation of core-variable model and contact scoring.
a–c, The core-variable model reliably predicts the empirical synaptic and gap-junction contact reproducibility (ℂδ and Gδ) on M2 and M3. To predict synaptic and gap-junctional contact counts on Mj < 4 contacts, ℂδ (or Gδ) contact counts on M4 are scaled by the ratio of 'all ℂ (G) on Mj count':'all ℂ (G) on M4 count' (Methods). For example, in a, the model predicts a ℂ3 count on M3 contacts as 206 × 285/1,474 = 40, where 206 is the empirical ℂ3 count on M4 contacts, 285 is the total empirical synaptic contact count, ℂ, on M3 and 1,474 is the total empirical count of synaptic contacts on M4. The model prediction is consistent with the empirical ℂ3 on M3 count (43). Error bars: ± \(\surd n\), where n is the empirical or predicted count (see Source Data for precise n values). d, e, Chemical synapses (d) and gap junctions (e) also consist of a core and variable circuit. Surrogate model data for ℂδ and Gδ, generated as in Fig. 2b. Across each dataset, around 62% of synaptic contacts and around 59% of gap-junction contacts consist of target contacts (given by fp/[fp + (1 − f) (1 − s)], Methods). f, g, Core synaptic contacts are typically larger than variable ones in both Cook et al.3 and White et al.5. Distribution of ℂδ (f) and Gδ (g) contact counts by electron micrograph sizes (the total number of electron microscopy sections in which a contact was observed)3,7. To check for biases in contact size due to possible differences in synaptic or gap-junction scoring criteria, we compare the distributions of electron micrograph sizes for contacts identified by White et al.5 (orange) and those identified by Cook et al.3 (blue). Because the White et al. report5 does not provide electron micrograph sizes, we used the sizes from Cook et al.3 for all contacts. Although many additional synapses identified by Cook et al.3 occur only in one section, we find no systematic bias towards smaller synaptic contacts by Cook et al.3. h, i, Bidirectional comparison of Cook et al.3 and Witvliet et al.20 synaptic contact reproducibility. h, Fraction of Cook et al.3 synaptic contacts scored by Witvliet et al.20. i, Fraction of Witvliet et al.20 synaptic contacts scored by Cook et al.3. In h, i, data are presented as fractions of the total empirical count of synaptic contacts (n).
Extended Data Fig. 5 Robust clustering of nerve-ring processes from M4 spatial population models.
The variability of membrane contacts (Fig. 2, Extended Data Fig. 2) suggests that no single contactome is representative of the population. We estimated the variability among membrane contact areas. a, The log-normalized empirical distribution of M4 membrane contact areas (mean centred at 0; STD, standard deviation; red line, normal distribution with empirical mean and standard deviation; n = 1,258 membrane contacts). We estimated the variability across the four datasets (L4 left, L4 right, adult left and adult right). For each conserved M4 contact, we computed the mean and standard deviation of the membrane contact area across the four datasets (see Methods). b, Plot of the standard deviation versus the mean contact area across the datasets, where each point is one M4 contact. Similar to Extended Data Fig. 1a, we find no dependence of the variability on membrane contact area. Therefore, we estimate membrane contact area variability by the mean variability among all membrane contact areas. c, The distribution of standard deviations of membrane contact area for all M4 contacts. Red dashed line indicates mean standard deviation. d–i, A stochastic spatial population model matches the above distributions by randomly perturbing membrane contact areas in the four datasets with multiplicative white noise with standard deviation (σ) of 0.23 (Methods). d–f, Spatial population data perturb the membrane contact areas while maintaining contact area and variability distributions that are similar to the empirical M4 contact area distributions. g, Perturbed contact areas scale linearly with the empirical contact areas. h, The spread of perturbed contact areas (log of the perturbed contact area as a fraction of the empirical contact area) is mostly uniform across membrane contact areas. i–l, Neurite clusters obtained from a population of 1,000 \(\tilde{{{\mathbb{M}}}^{4}}\) perturbed individuals and 1,000 \(\tilde{{\rm{L}}4}\) and \(\tilde{{\rm{Adult}}}\) perturbed individuals (perturbing left–right conserved contacts in the L4 and adult contact sets). For each perturbed individual in each population we used a multi-level graph-clustering algorithm to identify spatial clusters. Across each population, we computed the frequency that cell pairs cluster together, represented as an n × n cluster-frequency matrix (n = 93). A hierarchical clustering algorithm is used to sort the rows and columns of the cluster-frequency matrix to minimize variation along the diagonal. Hence, cell pairs that frequently cluster together are sorted together on the cluster-frequency matrix (Methods). Five largely overlapping subgroups of neurons emerge across different perturbations (see Fig. 1 and ‘Cluster assignment and validation’). i, Consensus clusters are robust across contact sets. \(\tilde{{\rm{L}}4}\) and \(\tilde{{\rm{Adult}}}\) clusters visualized using row and column colours of the \(\tilde{{{\mathbb{M}}}^{4}}\) population cluster assignments (dashed box). j, The consensus clusters are robust across different noise amplitudes. Clustering applied to populations generated by perturbations to M4 using white noise with standard deviations 0 (empirical data), 0.12, 0.45 and 0.9. k, l, The consensus clusters are robust across different spatial domains. k, Clustering applied to \(\tilde{{{\mathbb{M}}}^{4}}\) populations generated from a more spatially restricted volume of the neuropil34, which excludes its posterior lobe. l, Clustering applied to populations generated by perturbations to all reproducible membrane contact areas after restoring the smallest 35% contact areas to each of the L4, adult and M4 datasets (Extended Data Fig. 2). For all cluster-frequency matrices, matrix element (i, j) corresponds to the frequency that cells i and j cluster together across the 1,000 perturbed individuals. Row and column orders minimize variance along the diagonal (Methods). Cell cluster assignments (colour) follow the perturbed \(\tilde{{{\mathbb{M}}}^{4}}\) dataset (Fig. 1b reproduced in dashed box). Top, dendrogram of the hierarchical clustering.
Extended Data Fig. 6 Variable contacts obscure the organization of the nerve ring.
a, Cluster analysis of unperturbed membrane contact datasets M1, M2, M3 and M4. Clustering results for membrane contacts predicted to combine core and variable contacts (M3) and overwhelmingly variable contacts (M2, M1) significantly and increasingly diverge from five consensus clusters, indicated by large numbers of small clusters. b, Cluster analysis of (unperturbed) L4 and adult datasets. Both the unperturbed M4 and adult contact sets yield six clusters rather than the five clusters found in the perturbed population models (Fig. 1c, Extended Data Fig. 5). The additional cluster results from a split of the taxis cluster into two. This split of the taxis cluster is not observed in either the perturbed M4 or the perturbed adult contact sets, even with half of the noise levels observed empirically, indicating that the split is unlikely to be robust across a population of animals. For all cluster-frequency matrices, row and column ordering and colours are the same as the perturbed \(\tilde{{{\mathbb{M}}}^{4}}\) population dataset (Extended Data Fig. 5i). Matrix element (i, j) is 1 if cells i and j cluster together and 0 otherwise. Top: dendrogram of the hierarchical clustering.
Extended Data Fig. 7 Distribution of core and variable synapses among neighbourhoods.
a, Membrane contacts of the L4, adult and M4 reference all have similar membrane contact profiles. For L4 and adult, only bilaterally conserved contacts are included. b, Synaptic contacts on M4 membrane contacts broken down by degree of synaptic contact reproducibility (ℂ1, ℂ2, ℂ3 and ℂ4). Most (56%) of conserved synapses (ℂ4) occur within clusters near the main diagonal, whereas variable synapses (ℂ1) are spread across clusters. c, Gap-junction contacts on M4 membrane contacts broken down by degree of reproducibility (G1, G2, G3 and G4). For all matrices, row and column ordering is the same as the perturbed \(\tilde{{{\mathbb{M}}}^{4}}\) dataset (Extended Data Fig. 5i). Row and column colours correspond to final clusters assignments (Fig. 1d), where unclassified cells are coloured grey. Matrix element (i, j) corresponds to the fraction of cell i’s membrane contact with cell j, with rows normalized to sum to 1.
Extended Data Fig. 8 Subcellular structures support local and nonlocal connectivity, and RIA and AIB processes exhibit synaptic compartmentalization.
a, b, Volumetric rendering of RIAL and its synapses (cuboids) coloured by synaptic polarity (a) or intra-/inter-cluster (b). The synaptic input and output segments in a correspond to changes in neighbourhood composition in b. Changes in RIA neighbourhood correspond to the three neurite segments (nV, nD and loop) which exhibit independent calcium dynamics that encode head movement46. c, d, AIB processes change neighbourhood at the dorsal midline18. The ipsilateral segment (†) of the AIB process is surrounded by cells in the taxis cluster, whereas the contralateral segment (††) makes contact with cells in every other cluster. c, AIB-process segments alternate between synaptic inputs on the ipsilateral side and synaptic outputs on the contralateral side. d, The alternating synaptic inputs and outputs correspond to a change in neighbourhood occurring at the dorsal midline. e–h, Flattened protrusions link processes to adjacent cells in adjacent clusters. e, The flattened protrusion strategy as demonstrated by RIM processes (♦). f, The RMDV processes show how flattened protrusions are used to locally expand synaptic polarity. On the contralateral side, the main process trajectory is postsynaptic whereas the contralateral protrusion is presynaptic. Both AVA (g) and SAAV (h) neurons exhibit flattened protrusions that appear to turn into small branches. The small AVA branch extends into a neighbourhood of cells from a different cluster (*). SAAV ipsilateral branches receive synaptic inputs, whereas its main process trajectory on the contralateral side is mostly presynaptic. Spine-like features extend from RMEV/D processes to cells in a different cluster. i, j, Two longer RMED extensions (i) and three shorter RMEV spine-like extensions (j) are postsynaptic to the sublateral cluster. In all images, the pharynx is shown for spatial reference. R, right; A, anterior; V, ventral. Note that for visual clarity, synapses have been offset from the cell process. k, Schematic of neighbourhood changes of selected cells (labelled in the colour of the cluster assignment). P, proximal and D, distal to cell body. Each trajectory is scaled to the length of the reconstructed left L4 process. Black boxes denote sections in which the process makes contact with at least two clusters.
Extended Data Fig. 9 Network features of the brain map.
a, Schematics of network features (from left to right): Feed-forward loop (FF) motif defined by a triplet of nodes with connectivity: Source → Intermediary → Target and Source → Target; network hub (high-degree node); fan-in (high in-degree node); fan-out (high out-degree node); and rich club (highly connected hubs). b, FF triplets within the brain map support the ResNet architecture of the nerve ring. All 101 FF instances among ℂ4 synaptic contacts (all edges in Fig. 4, Extended Data Fig. 10) are shown. Black arrows, synaptic contacts forming FF motifs within the ResNet architecture (Fig. 4); grey arrows, additional synaptic contacts forming FF motifs (Extended Data Fig. 10). A total of 72 out of 93 cell classes participate in at least one FF motif. Prominent FF targets include AIA, AIB, AIZ, AVA, AVB, AVE, RIA, RIC, RIM, RIP, RMDV and SMDV. Additional contacts superimposed on the ResNet come mostly from cross-sensory module connectivity (Extended Data Fig. 10b). c, RIP, the only synaptic link between the somatic and pharyngeal nervous systems, is a major FF target cell for papillary sensory source cells and URA intermediaries. d, AIA is a major taxis layer-2 intermediary cell pair regulating information flow from layer-1 taxis sensory cells onto the layer-3 AIB taxis target cell. e, AIZ, major layer-3 cells that support nonlocal connectivity (Fig. 3a), serve both to integrate information flow from layer-1 and layer-2 taxis source cells (fan-in) and as intermediary connections to various layer-3 target cells in other modules (fan-out). f, Primary locomotion-regulating interneurons—AVA, AVB and AVE—are major layer-3 FF targets and connect extensively onto motor neurons of the ventral nerve cord. Connectivity among these cells occurs in the ventral nerve cord (but is not observed in the nerve ring), suggesting that the regulation of locomotion down the body occurs posteriorly to the nerve ring. g, The cell pair RIM, a major hub that supports nonlocal connectivity, serves a triple purpose as a source, intermediary and target of FF motifs within layer 3. h, The nonlocal supporter, multi-compartment cell pair RIA is a major FF target for layer-1 sensory (primarily avoidance) source cells and for layer-2 and layer-3 (taxis and avoidance) intermediary cells as well as intermediaries that control layer-3 head motor neurons. In addition, RIA neurons are major targets for feedback from lateral (RMD, RMDD, RMDV) and sublateral (SMDD, SMDV) head motor neurons, consistent with their roles in spatially encoding dorso-ventral head movement to coordinate turning behaviours46. i, Major targets of FF motifs (11 neuron classes acting as a target of more than three FF motifs, including five classes of rich club neurons) form a highly interconnected subnetwork. Note the frequent representation of some cells in multiple motifs (c–i). j, Aggregated synaptic contacts of layer-3. Layer-3 FF motifs within and among the modules show strong recurrence and no clear feed-forward directionality or hierarchy of layer-3 connectivity, consistent with highly distributed computation. Sublaterals are merged into the lateral module node. Layer-3 anterior cells form FF motifs with only one other module (taxis). All network schematics were generated with Cytoscape v.3.7.1.
Extended Data Fig. 10 Around 17% of ℂ4 contacts are not accounted for by the ResNet model.
a, Layer-1 synaptic connectivity across information-processing modules in ℂ4 could support distributed sensory computation and integration. Eight (2% of ℂ4) contacts occur between sensory cells across different modules. b, These contacts include: (i) ADE→OLL; (ii) ALM→CEPD/V; and (iii) reciprocal contacts between ASH, ADL and ADF. (i) The mechanosensitive47,48 anterior neurons OLL loop around intermediate processes, whereas the processes of ADE extend towards the OLL loop, suggesting a functional role for the more elaborate loop morphology. (ii) Both CEPD and CEPV processes loop around intermediate processes and extend flattened protrusions to meet the ALM processes, in which ALM are postsynaptic. CEPD and CEPV neurons respond to head touch49, whereas ALM neurons respond to both gentle50 and harsh51 body touch, inhibit backward locomotion52 and have been implicated in the habituation of tap response53. (iii) Nociception: ASH, ADE and ADF neurons may coordinate avoidance behaviours between the taxis and avoidance modules54. c, Layer-1 to layer-3 inter-module feed-forward synaptic connectivity in ℂ4. A total of 54 (12% of ℂ4) contacts are inter-module, originate in layer 1 and target layer-3 neurons directly. A small number of taxis and avoidance sensory neurons (ADF, ADL, ASH, URX and BAG) project to all but laterals in layer 3; this contrasts with extensive anterior sensory neuron projections that almost exclusively target (sub)lateral layer-3 interneurons and motor neurons, probably mediating rapid sensorimotor transformations. d, Layer-2 and inter-module feed-forward ℂ4 synaptic connectivity. Three contacts (1% of ℂ4) are inter-module and originate in layer 2. Notably, the layer-2 taxis AIY neurons synapse onto RIA—layer-3 anterior multi-compartment neurons. e, Inter-module synaptic feedback connectivity in ℂ4. Nine (3% of ℂ4) contacts provide inter-module feedback. Black arrows, synaptic contacts between cells in the same neighbourhood; grey arrows, synaptic contacts between layer-3 cells in different neighbourhoods; red arrows, synaptic contacts not accounted for by the ResNet model; solid arrows, feed-forward or recurrent (intra-layer) synaptic contacts; dashed arrows: feedback synaptic contacts.
Supplementary information
Supplementary Information
This file contains Supplementary Results including a more technical treatment of: 1 - Analysis to establish bilateral homology (Extended Data Figs. 1 and 2); 2 - Similarity between homologous and proximal immediate neighbourhoods (Extended Data Figs 2a); 3 - Validation of core-variable synaptic model (Extended Data Fig. 3); 4 - Functional identification of neuron process clusters (Fig 1c,d and Extended Data Fig 5); 5 - Rich-club neurons exhibit meso-connectivity properties (Fig. 3, Extended Data Figs. 8 and 9) and 6 - Network motifs and architectural features of the brain map (Extended Data Fig. 9). It also contains Supplementary References and full descriptions for Supplementary Data files 1-5.
Supplementary Data 1
Membrane adjacency data scored in the L4 and adult nerve rings – see Supplementary Information document for full description.
Supplementary Data 2
Membrane contact localization data for all bilateral cells – see Supplementary Information document for full description.
Supplementary Data 3
\({\mathbb{M}}\)δ, ℂδ and \({\mathbb{G}}\)δ reference graphs – see Supplementary Information document for full description.
Supplementary Data 4
Cluster assignment and features of process morphology and synaptic organization – see Supplementary Information document for full description.
Supplementary Data 5
Validation of our membrane contact data against membrane contacts scored by White et al. (1983)18– see Supplementary Information document for full description.
Supplementary Table 1
Summary of volumetric reconstruction and extracted reference datasets.
Supplementary Table 2
Error analysis of reconstruction.
Supplementary Table 3
Lateral synaptic contacts are not distinguishable from random contacts.
Supplementary Table 4
Classification of brain map ℂ4 contacts.
Video 1
A fly through of the segmented L4 (JSH) serial sectioned electron micrograph (EM) series. Neurites are manually segmented with TrakEM2. Segmented neurites are coloured based on final clusters assignment (Fig. 1d). The EM series starts at the most anterior section of the nerve ring neuropil and proceeds posteriorly.
Video 2
A fly through of the segmented Adult (N2U) serial sectioned EM series. Neurites are manually segmented with TrakEM2. Segmented neurites are coloured based on final clusters assignment (Fig. 1d). The EM series starts at the most anterior section of the nerve ring neuropil and proceeds posteriorly.
Video 3
Volumetric rendering of the complete L4 nerve ring neuropil. Neuronal processes are coloured by final cluster assignment (Fig. 1d). The video starts with the anterior part of the neuropil facing left, the left side of the neuropil facing the viewer and the dorsal part of the neuropil facing up. The neuropil is then rotated 360o about the vertical (dorsoventral) axis. Note that the wider anterior regions correspond to the nerve ring commissure shown in Fig 1a. The narrower region corresponds to the posterior lobe of the nerve ring neuropil.
Video 4
Volumetric reconstruction of representative (L4) neurons SMBVL and SMBVR. Rotating images of the SMBVL and SMBVR neurons illustrate the bilateral symmetry of their respective processes and of specialized topographical features along these processes. At the start of the video, both left and right neurons are colored by final cluster assignment (sublateral, Fig. 1d). The pharynx (gray volume, ~16 μm) is included for spatial context. Midway through the video the pharynx is removed and the left and right cells are assigned different colors to distinguish between the cells.
Video 5
Volumetric reconstruction of representative (L4) neurons RIBL and RIBR. Rotating images of the RIBL and RIBR neurons illustrate the bilateral symmetry of their respective processes and of specialized topographical features along these processes. At the start of the video, both left and right neurons are coloured by final cluster assignment (taxis, Fig. 1d). The pharynx (grey volume, ~16μm) is included for spatial context. Midway through the video the pharynx is removed and the left and right cells are assigned different colours to distinguish between the cells.
Video 6
Volumetric reconstruction of representative (L4) neuron class IL1. Rotating images of all six IL1 class members (IL1L/R, IL1VL/R,IL1DL/R) illustrate the bilateral symmetry of more complex processes. IL1 processes are closely aligned on both sides of the body. At the start of the video, all cells are coloured by final cluster assignment (anterior, Fig. 1d). The pharynx (grey volume, ~16μm) is included for spatial context. Midway through the video the pharynx is removed and each of the 6 neurons is assigned a different colour to distinguish between the cells.
Video 7
Volumetric reconstruction of representative (L4) neurons RIML and RIMR. Rotating images of RIML and RIMR neurons illustrate the bilateral symmetry of their respective processes and of specialized topographical features along these processes. At the start of the video, both left and right neurons are coloured by final cluster assignment (lateral, Fig. 1d). The pharynx (grey volume, ~16μm) is included for spatial context. Midway through the video the pharynx is removed and the left and right cells are coloured differently to distinguish between the cells.
Source data
Rights and permissions
About this article
Cite this article
Brittin, C.A., Cook, S.J., Hall, D.H. et al. A multi-scale brain map derived from whole-brain volumetric reconstructions. Nature 591, 105–110 (2021). https://doi.org/10.1038/s41586-021-03284-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03284-x
This article is cited by
-
Understanding neural circuit function through synaptic engineering
Nature Reviews Neuroscience (2024)
-
Comparative connectomics of dauer reveals developmental plasticity
Nature Communications (2024)
-
The evolution of Big Data in neuroscience and neurology
Journal of Big Data (2023)
-
Communities in C. elegans connectome through the prism of non-backtracking walks
Scientific Reports (2023)
-
What if worms were sentient? Insights into subjective experience from the Caenorhabditis elegans connectome
Biology & Philosophy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.