Abstract
Antibody affinity maturation depends on positive selection in germinal centres (GCs) of rare B cell clones that acquire higher-affinity B cell receptors via somatic hypermutation, present more antigen to follicular helper T (TFH) cells and, consequently, receive more contact-dependent T cell help1. As these GC B cells and TFH cells do not maintain long-lasting contacts in the chaotic GC environment2,3,4, it is unclear how sufficient T cell help is cumulatively focused onto those rare clones. Here we show that, upon stimulation of CD40, GC B cells upregulate the chemokine CCL22 and to a lesser extent CCL17. By engaging the chemokine receptor CCR4 on TFH cells, CCL22 and CCL17 can attract multiple helper cells from a distance, thus increasing the chance of productive help. During a GC response, B cells that acquire higher antigen-binding affinities express higher levels of CCL22, which in turn ‘highlight’ these high-affinity GC B cells. Acute increase or blockade of TFH cells helps to rapidly increase or decrease CCL22 expression by GC B cells, respectively. Therefore, a chemokine-based intercellular reaction circuit links the amount of T cell help that individual B cells have received recently to their subsequent ability to attract more help. When CCL22 and CCL17 are ablated in B cells, GCs form but B cells are not affinity-matured efficiently. When competing with wild-type B cells in the same reaction, B cells lacking CCL22 and CCL17 receive less T cell help to maintain GC participation or develop into bone-marrow plasma cells. By uncovering a chemokine-mediated mechanism that highlights affinity-improved B cells for preferential help from TFH cells, our study reveals a principle of spatiotemporal orchestration of GC positive selection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data generated during and/or analysed during the current study are available from the corresponding author upon reasonable request. scRNA-seq data of human tonsillar GCs has been deposited in the Genome Sequence Archive for Human under the accession number HRA000540. Source data are provided with this paper.
Change history
15 March 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-03384-8
References
Victora, G. D. & Nussenzweig, M. C. Germinal centers. Annu. Rev. Immunol. 30, 429–457 (2012).
Allen, C. D., Okada, T., Tang, H. L. & Cyster, J. G. Imaging of germinal center selection events during affinity maturation. Science 315, 528–531 (2007).
Shulman, Z. et al. Dynamic signaling by T follicular helper cells during germinal center B cell selection. Science 345, 1058–1062 (2014).
Liu, D. et al. T–B-cell entanglement and ICOSL-driven feed-forward regulation of germinal centre reaction. Nature 517, 214–218 (2015).
Victora, G. D. et al. Germinal center dynamics revealed by multiphoton microscopy with a photoactivatable fluorescent reporter. Cell 143, 592–605 (2010).
Gitlin, A. D. et al. T cell help controls the speed of the cell cycle in germinal center B cells. Science 349, 643–646 (2015).
Mayer, C. T. et al. The microanatomic segregation of selection by apoptosis in the germinal center. Science 358, eaao2602 (2017).
Griffith, J. W., Sokol, C. L. & Luster, A. D. Chemokines and chemokine receptors: positioning cells for host defense and immunity. Annu. Rev. Immunol. 32, 659–702 (2014).
Tang, H. L. & Cyster, J. G. Chemokine up-regulation and activated T cell attraction by maturing dendritic cells. Science 284, 819–822 (1999).
Iellem, A. et al. Unique chemotactic response profile and specific expression of chemokine receptors CCR4 and CCR8 by CD4+CD25+ regulatory T cells. J. Exp. Med. 194, 847–853 (2001).
Sather, B. D. et al. Altering the distribution of Foxp3+ regulatory T cells results in tissue-specific inflammatory disease. J. Exp. Med. 204, 1335–1347 (2007).
Shinnakasu, R. et al. Regulated selection of germinal-center cells into the memory B cell compartment. Nat. Immunol. 17, 861–869 (2016).
Rapp, M. et al. CCL22 controls immunity by promoting regulatory T cell communication with dendritic cells in lymph nodes. J. Exp. Med. 216, 1170–1181 (2019).
Rodriguez, E. A. et al. The growing and glowing toolbox of fluorescent and photoactive proteins. Trends Biochem. Sci. 42, 111–129 (2017).
Holmes, A. B. et al. Single-cell analysis of germinal-center B cells informs on lymphoma cell of origin and outcome. J. Exp. Med. 217, e20200483 (2020).
Shih, T. A., Roederer, M. & Nussenzweig, M. C. Role of antigen receptor affinity in T cell-independent antibody responses in vivo. Nat. Immunol. 3, 399–406 (2002).
Wang, Y. et al. Germinal-center development of memory B cells driven by IL-9 from follicular helper T cells. Nat. Immunol. 18, 921–930 (2017).
Shi, J. et al. PD-1 controls follicular T helper cell positioning and function. Immunity 49, 264–274 (2018).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Acknowledgements
We thank M. Nussenzweig for providing the B1-8hi mice. The work was funded in part by the National Key R&D Program of China (Ministry of Science and Technology, 2018YFE0200300 to H.Q.), National Natural Science Foundation of China (grant 81621002, 31830023, 81761128019 to H.Q.), the Tsinghua-Peking Center for Life Sciences, the Beijing Municipal Science & Technology Commission, and the Beijing Frontier Research Center for Biological Structure.
Author information
Authors and Affiliations
Contributions
H.Q. conceptualized and supervised the study. B.L. conducted a majority of the experiments and designed parts of the study. Y.L. helped with mouse experiments. J. Yan, D.L. and W.M. were involved in initial imaging observations. W.L. contributed critical reagents. C.W. and L.Z. contributed to human GC work. J. Yao and J.W. helped with scRNA-seq analyses. H.Q. and B.L. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Charles Mackay, Carola Vinuesa and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The frequency issue in T cell–B cell interactions for GC positive selection and the model of chemoattraction-driven remote sensing of affinity information.
a–c, Conceptualization of the frequency dilemma. Red arrow, a hypothetical unit of T cell help delivered during a cognate T cell–B cell contact; blue shade, a hypothetical range of potential that does not ensure survival; red shade, a hypothetical range of potential that ensure positive selection. a, Receipt of T cell help is necessary to ensure survival (black curve), as GC cells die in its absence (blue curve). b, Frequent T cell contacts raise the positive selection potential (black curve), which decays when T cell contacts and help signals are not as frequent (grey curve). c, Sufficiently frequent T cell contacts help B cells to cross the threshold for positive selection (red curve), leading to cyclic re-entry, memory B cell formation or plasma cell formation as possible outcomes (red arrows). d, The working model of CCL22-mediated dynamic highlighting and remote sensing of B cell affinity information. Cognate T cell–B cell contact and help-signal strength are ultimately determined by the B cell affinity; help signals received during brief cognate contacts with T cells (green) drive CCL22 (cyan) upregulation in the B cell (red); higher-affinity cells produce more CCL22, attracting more TFH cells from a distance; more attempts to contact by TFH cells are translated into more contacts and better help signals experienced by a given B cell, which in turn can produce more CCL22 to attract more help; without experiencing sufficient T cell help, either because of low affinities or by chance, B cells also rapidly lose CCL22 production (not depicted).
Extended Data Fig. 2 CCL22 expression by GC B cells after in vitro stimulation.
a, b, Gating strategies for sorting GC and non-GC B cells after in vivo anti-CD40 stimulation (corresponding to Fig. 1. a, b) and in vitro anti-CD40 or anti-IgM stimulation (corresponding to Extended Data Fig. 2. c–f). c–f, Relative expression of Ccl22 normalized to Cd19 in sort-purified GC B cells (c, d) and non-GC B cells (e, f) after in vitro stimulation with 10 μg ml−1 anti-CD40 (c, e), anti-IgM (d, f) or the respective controls, measured in 2 independent experiments as indicated. Data are mean ± s.e.m. of triplicate wells of indicated treatment.
Extended Data Fig. 3 Schemes for genetic ablation of Ccl22, Ccl17 and Ccr4.
a, b, CRISPR–Cas9-mediated strategy for genomic deletion of Ccr4. The targeting sgRNA (red line), genotyping primers (yellow arrowheads) and the deleted segment (yellow highlight) are shown in the context of genomic sequences (nucleotide coordinates according to NCBI) (a), with representative genotyping PCR products (b), which differ between wild-type and Ccr4-KO alleles by 8 bp and are distinguished by sequencing. c, Surface CCR4 expression on CD4+ T cells of indicated genotypes following in vitro activation with anti-CD3 and anti-CD28 for 4 days. d–g, CRISPR–Cas9 strategies for Ccl17 and Ccl22 ablation. Relevant targeting sgRNAs (red lines), genotyping primers (yellow arrowheads) and deleted segments (purple lines) are shown in the context of genomic sequences (nucleotide coordinates according to NCBI) for Ccl17 (d) and Ccl22 (f), with representative genotyping PCR results shown for Ccl17 (e) and Ccl22 (g).
Extended Data Fig. 4 TFH cells exhibited robust migration towards CCL22 and CCL17.
a, b, Transwell migration towards CCL17 and CCL22 by T cell subsets from Foxp3IRES-GFP reporter mice. Representative gating of input cells (a) and migrated cells after 3 h (b). c, Summary data of migration indices, from three independent experiments indicated by three different symbols. Data for each experiment are mean ± s.e.m. of triplicate or quadruplicate wells under each condition.
Extended Data Fig. 5 CCL17 and CCL22 promote TFH cell–GC B cell encounters without changing contact duration.
a, Duration of contact between TFH cells and wild-type or CCL17/22-DKO B cells in follicles. See corresponding Supplementary Videos 1, 2 and Fig. 2a, b. b, Duration of contact between wild-type or CCR4-KO TFH cells and B cells in follicles. See corresponding Supplementary Videos 3, 4 and Fig. 2d, e. c–h, Interacting OT-II T cells and MD4 B cells in GCs 120 h post-immunization. c, Time-lapse images highlighting one CFP-expressing CCL17/22-DKO (blue arrowheads) and one GFP-expressing wild-type (green arrowheads) MD4 B cells being contacted by dsRed-expressing OT-II T cells (red arrowheads), with each incidence of contact marked by a broken white line between the two cells. Also see corresponding Supplementary Video 5. d, Time-lapse images of CFP- and GFP-expressing wild-type MD4 B cells contacted by OT-II T cells, colour-coded as in c. Also see corresponding Supplementary Video 6. e, Summary NHRB statistics. Each dot is one field imaged, red lines denote means. f, Time-lapse images highlighting dsRed-expressing CCR4-KO (red arrowhead) and GFP-expressing wild-type (green arrowheads) OT-II T cells being contacted by CFP-expressing MD4 B cells (blue arrowheads), with each incidence of contact marked by a broken white line between the two cells. Also see corresponding Supplementary Video 7. g, Time-lapse images of dsRed- and GFP-expressing wild-type OT-II T cells contacted by CFP-expressing MD4 B cells, colour-configured as in f. Also see corresponding Supplementary Video 8. h, Summary NHRB statistics. Each dot is one field imaged, red lines denote means. Data are pooled from 5 mice for WT–WT mix and 5 mice for DKO–WT mix or KO–WT mix, imaged in 3 independent experiments. P values by two-tailed t-tests. Scale bars, 20 μm. i, Durations of contacts between TFH cells and wild-type or DKO B cells in GCs, as measured in intravital imaging sessions represented by c, d and Supplementary Videos 5, 6. j, Durations of contacts between wild-type or CCR4KO TFH cells and MD4 B cells in GCs, as measured in intravital imaging sessions represented by f, g and Supplementary Videos 7, 8. Each symbol is one incidence of a T cell–B cell contact, and red lines denote means. Data are pooled from three independent experiments. P values by two-tailed t-tests.
Extended Data Fig. 6 Construction of CCL22-KI-tdTomato mice.
a, The CRISPR–Cas9 strategy for IRES-tdTomato insertion into Ccl22. The targeting sgRNA (red line), genotyping primers (yellow arrowheads) and the insert segment (red block) are shown in the context of genomic sequences (nucleotide coordinates according to NCBI). b, Representative genotyping PCR results for the wild-type and KI mutant allele. c, Gating of CD11c+ dendritic cells in lymph node cells (left) and contour plots to show tdTomato reporter fluorescence from CCL22-KI-tdTomato or control B6 mice (right).
Extended Data Fig. 7 Characteristics of GC B cells and serum antibody affinities in the absence of B cell CCL17 and CCL22.
a, Representative contour plots of NIP-binding GC B cells of μMT (80%):wild-type (20%) or μMT (80%):DKO (20%) bone marrow chimaera. b, Summary statistics of fractions of NIP-binding GC B cells at indicated time points. Each dot indicates one mouse, and lines denote means. Data are pooled from three independent experiments. P values by two-tailed t-tests. c, Representative contour plots of IgM+ and IgG1+ GC B cells of μMT (80%):wild-type (20%) or μMT (80%):DKO (20%) bone marrow chimaera. d, e, Summary statistics of IgM+ (d) and IgG1+ (e) GC B cells at day 13 post-immunization. Each dot indicates one chimaeric mouse, and lines denote means. Data are pooled from three independent experiments. P values by two-tailed t-tests. f, Representative ELISA titration curves of serum IgG binding to NP4- or NP30-BSA 21 days post-immunization. g, Ratios between NP4-binding and NP30-binding EC50. Each symbol indicates one mouse, and lines denote means. Data are pooled from three independent experiments. The P value by two-tailed t-test.
Extended Data Fig. 8 Compromised GC formation and output and retarded affinity maturation in the absence of T cell CCR4.
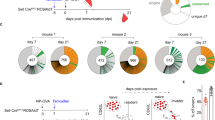
Analyses of GCs and BMPCs in Tcrb−/−Tcrd−/− (TCR-KO) bone marrow chimaeric mice. a, b, Representative contour plots of GC B cells (a) and BMPCs (b) of TCR-KO (80%):wild-type (20%) or TCR-KO (80%):CCR4-KO(20%) bone marrow chimaera. c, d, Summary statistics of GC B cells (c) and BMPCs (d) at indicated time points after immunization. Each dot indicates one chimaeric mouse, and lines denote means. Data are pooled from three independent experiments. P values by two-tailed t-tests. e, f, Numbers of W33L mutations among unique VH186.2 sequences of GC B cells (e) and BMPCs (f) at indicated time points. Total numbers of VH186.2 sequences analysed are given at the centre of each pie chart, and total numbers of nucleotide mutations per sequence are given under each pie chart as mean ± s.e.m. Data are pooled from three independent experiments. P values by two-tailed Fisher’s exact tests.
Extended Data Fig. 9 Gating strategy for generating competitive competency in CD45.1:CD45.2 wild-type and CD45.1:CD45.2 DKO groups.
a, Representative gating of total B cells, GC B cells in the spleen and BMPCs in the bone marrow. b, c, Representative contour plots of GC B cells and BMPCs of CD45.1 WT:CD45.2 DKO or CD45.1 WT:CD45.2 WT bone marrow chimaera 13 (b) or 21 (c) days after NP-KLH immunization.
Extended Data Fig. 10 The CCR4–CCL22 axis in human TFH and GC B cells.
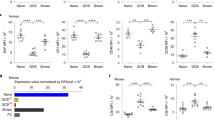
a, Representative gating of non-TFH cells, CXCR5+ T cells and CXCR5+PD1+ tonsillar TFH cells. b, Histograms of CCR4 expression overlaid with isotype control staining (iso). c, CCR4 mean fluorescent intensities (MFI), with respective isotype control background subtracted. Data are mean ± s.e.m. of four donors. P values by two-tailed t-tests. d, Representative gating of human tonsillar GC B cells (left), further divided into light-zone (LZ) and dark-zone (DZ) fractions (right). e, CCL22 upregulation by light-zone GC B cells after 4 h anti-CD40 or control hamster IgG treatment. Data are pooled from nine independent donors. P values by two-tailed Mann–Whitney U-tests. f, Violin plots of CD40 signature genes, identified by Holmes et al.15, as probed by scRNA-seq. The top and bottom rows present cells characterized with CCL22 expression >0 and =0, respectively. Single-cell data were collected from three healthy donors. P values were calculated on the basis of hypergeometric distribution (see Methods for details).
Supplementary information
Supplementary Table
Supplementary Table 1. Primers for RT-PCR detection of chemokine expression. All the primers for detection of chemokine expression are listed.
Video 1
Interaction dynamics between OT-II T cells and CFP-expressing CCL17/22 DKO or GFP-expressing wildtype MD4 B cells in the follicle. CFP-expressing CCL17/22 DKO (blue arrowheads) and GFP-expressing CCL17/22 WT (green arrowheads) MD4 cells interacting with dsRed-expressing TFH cells (red arrowheads), visualized 96 h post immunization. Broken white lines mark contact incidences. See corresponding time-lapse images in Fig. 2a. Scale bar, 20 μm.
Video 2
Interaction dynamics between OT-II T cells and CFP-expressing wildtype or GFP-expressing wildtype MD4 B cells in the same follicle. CFP-expressing CCL17/22 WT (blue arrowheads) and GFP-expressing CCL17/22 WT (green arrowheads) MD4 cells interacting with dsRed-expressing TFH cells (red arrowheads), visualized 96 h post immunization. Broken white lines mark contact incidences. See corresponding time-lapse images in Fig. 2b. Scale bar, 20 μm.
Video 3
Interaction dynamics between MD4 B cells and GFP-expressing CCR4 KO or mAmetrine-expressing wildtype OT-II T cells in the same follicle. GFP-expressing CCR4 KO (green arrowheads) and mAmetrine-expressing CCR4 WT (blue arrowheads) T cells interacting with dsRed-expressing MD4 B cells (white arrowheads), visualized 96 h post immunization. Broken white lines mark contact incidences. See corresponding time-lapse images in Fig. 2d. Scale bar, 20 μm.
Video 4
Interaction dynamics between MD4 B cells and GFP-expressing wildtype or mAmetrine-expressing wildtype OT-II T cells in the same follicle. GFP-expressing CCR4 WT (green arrowheads) and mAmetrine-expressing CCR4 WT (blue arrowheads) interacting with dsRed-expressing MD4 B cells (white arrowheads), visualized 96 h post immunization. Broken white lines mark contact incidences. See corresponding time-lapse images in Fig. 2e. Scale bar, 20 μm.
Video 5
Interaction dynamics between OT-II T cells and CFP-expressing CCL17/22 DKO or GFP-expressing wildtype MD4 B cells in the GC. CFP-expressing CCL17/22 DKO (blue arrowheads) and GFP-expressing CCL17/22 WT (green arrowheads) interacting with dsRed-expressing TFH cells (red arrowheads), visualized 120 h post immunization. Broken white lines mark contact incidences. See corresponding time-lapse images in Extended Data Fig. 5c. Scale bar, 20 μm.
Video 6
Interaction dynamics betweem OT-II T cells and and CFP-expressing wildtype or GFP-expressing wildtype MD4 B cells in the GC. CFP-expressing CCL17/22 WT (blue arrowheads) and GFP-expressing CCL17/22 WT (green arrowheads) interacting with dsRed-expressing TFH cells (red arrowheads), visualized 120 h post immunization. Broken white lines mark contact incidences. See corresponding time-lapse images in Extended Data Fig. 5d. Scale bar, 20 μm.
Video 7
. Interaction dynamics between MD4 B cells and dsRed-expressing CCR4 KO or GFP-expressing wildtype OT-II T cells in the GC. dsRed-expressing CCR4 KO (red arrowheads) and GFP-expressing CCR4 WT (green arrowheads) interacting with CFP-expressing GC MD4 B cells (blue arrowheads), visualized 120 h post immunization. Broken white lines mark contact incidences. See corresponding time-lapse images in Extended Data Fig. 5f. Scale bar, 20 μm.
Video 8
Interaction dynamics between MD4 B cells and dsRed-expressing CCR4 WT or GFP-expressing WT OT-II T cells in the GC. dsRed-expressing CCR4 WT (red arrowheads) and GFP-expressing CCR4 WT (green arrowheads) interacting with CFP-expressing GC MD4 B cells (blue arrowheads), visualized 120 h post immunization. Broken white lines mark contact incidences. See corresponding time-lapse images in Extended Data Fig. 5g. Scale bar, 20 μm.
Rights and permissions
About this article
Cite this article
Liu, B., Lin, Y., Yan, J. et al. Affinity-coupled CCL22 promotes positive selection in germinal centres. Nature 592, 133–137 (2021). https://doi.org/10.1038/s41586-021-03239-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03239-2
This article is cited by
-
STAT6 mutations enriched at diffuse large B-cell lymphoma relapse reshape the tumor microenvironment
International Journal of Hematology (2024)
-
The link between circulating follicular helper T cells and autoimmunity
Nature Reviews Immunology (2022)
-
CCL22 mutations drive natural killer cell lymphoproliferative disease by deregulating microenvironmental crosstalk
Nature Genetics (2022)
-
Single-cell Atlas of common variable immunodeficiency shows germinal center-associated epigenetic dysregulation in B-cell responses
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.