Abstract
Mechanosensitive channels sense mechanical forces in cell membranes and underlie many biological sensing processes1,2,3. However, how exactly they sense mechanical force remains under investigation4. The bacterial mechanosensitive channel of small conductance, MscS, is one of the most extensively studied mechanosensitive channels4,5,6,7,8, but how it is regulated by membrane tension remains unclear, even though the structures are known for its open and closed states9,10,11. Here we used cryo-electron microscopy to determine the structure of MscS in different membrane environments, including one that mimics a membrane under tension. We present the structures of MscS in the subconducting and desensitized states, and demonstrate that the conformation of MscS in a lipid bilayer in the open state is dynamic. Several associated lipids have distinct roles in MscS mechanosensation. Pore lipids are necessary to prevent ion conduction in the closed state. Gatekeeper lipids stabilize the closed conformation and dissociate with membrane tension, allowing the channel to open. Pocket lipids in a solvent-exposed pocket between subunits are pulled out under sustained tension, allowing the channel to transition to the subconducting state and then to the desensitized state. Our results provide a mechanistic underpinning and expand on the ‘force-from-lipids’ model for MscS mechanosensation4,11.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The cryo-EM maps have been deposited in the Electron Microscopy Data Bank under accession codes EMD-21462 (MscS–PC-18:1), EMD-21463 (MscS–PC-10), EMD-21464 (MscS–PC-18:1–βCD), EMD-21465 (MscS–PC-18:1–βCD-30 min) and EMD-21466 (MscS(Ala106Val)–DDM). The atomic coordinates have been deposited in the PDB under accession codes 6VYK (MscS–PC-18:1), 6VYL (MscS–PC-10) and 6VYM (MscS–PC-18:1–βCD).
Code availability
The custom-written SPIDER scripts used to measure the nanodisc size and the Python codes used to analyse molecular dynamics trajectories are available from the corresponding author on request.
References
Hamill, O. P. & Martinac, B. Molecular basis of mechanotransduction in living cells. Physiol. Rev. 81, 685–740 (2001).
Kung, C. A possible unifying principle for mechanosensation. Nature 436, 647–654 (2005).
Katta, S., Krieg, M. & Goodman, M. B. Feeling force: physical and physiological principles enabling sensory mechanotransduction. Annu. Rev. Cell Dev. Biol. 31, 347–371 (2015).
Cox, C. D., Bavi, N. & Martinac, B. Biophysical principles of ion-channel-mediated mechanosensory transduction. Cell Rep. 29, 1–12 (2019).
Sukharev, S. Purification of the small mechanosensitive channel of Escherichia coli (MscS): the subunit structure, conduction, and gating characteristics in liposomes. Biophys. J. 83, 290–298 (2002).
Levina, N. et al. Protection of Escherichia coli cells against extreme turgor by activation of MscS and MscL mechanosensitive channels: identification of genes required for MscS activity. EMBO J. 18, 1730–1737 (1999).
Sukharev, S. I., Blount, P., Martinac, B., Blattner, F. R. & Kung, C. A large-conductance mechanosensitive channel in E. coli encoded by mscL alone. Nature 368, 265–268 (1994).
Martinac, B., Buechner, M., Delcour, A. H., Adler, J. & Kung, C. Pressure-sensitive ion channel in Escherichia coli. Proc. Natl Acad. Sci. USA 84, 2297–2301 (1987).
Bass, R. B., Strop, P., Barclay, M. & Rees, D. C. Crystal structure of Escherichia coli MscS, a voltage-modulated and mechanosensitive channel. Science 298, 1582–1587 (2002).
Wang, W. et al. The structure of an open form of an E. coli mechanosensitive channel at 3.45 A resolution. Science 321, 1179–1183 (2008).
Reddy, B., Bavi, N., Lu, A., Park, Y. & Perozo, E. Molecular basis of force-from-lipids gating in the mechanosensitive channel MscS. eLife 8, e50486 (2019).
Martinac, B. Mechanosensitive ion channels: molecules of mechanotransduction. J. Cell Sci. 117, 2449–2460 (2004).
Haswell, E. S., Phillips, R. & Rees, D. C. Mechanosensitive channels: what can they do and how do they do it? Structure 19, 1356–1369 (2011).
Booth, I. R. Bacterial mechanosensitive channels: progress towards an understanding of their roles in cell physiology. Curr. Opin. Microbiol. 18, 16–22 (2014).
Akitake, B., Anishkin, A. & Sukharev, S. The “dashpot” mechanism of stretch-dependent gating in MscS. J. Gen. Physiol. 125, 143–154 (2005).
Cox, C. D. et al. Selectivity mechanism of the mechanosensitive channel MscS revealed by probing channel subconducting states. Nat. Commun. 4, 2137 (2013).
Kamaraju, K., Belyy, V., Rowe, I., Anishkin, A. & Sukharev, S. The pathway and spatial scale for MscS inactivation. J. Gen. Physiol. 138, 49–57 (2011).
Edwards, M. D., Bartlett, W. & Booth, I. R. Pore mutations of the Escherichia coli MscS channel affect desensitization but not ionic preference. Biophys. J. 94, 3003–3013 (2008).
Koprowski, P. & Kubalski, A. Voltage-independent adaptation of mechanosensitive channels in Escherichia coli protoplasts. J. Membr. Biol. 164, 253–262 (1998).
Pliotas, C. et al. Conformational state of the MscS mechanosensitive channel in solution revealed by pulsed electron-electron double resonance (PELDOR) spectroscopy. Proc. Natl Acad. Sci. USA 109, E2675–E2682 (2012).
Lai, J. Y., Poon, Y. S., Kaiser, J. T. & Rees, D. C. Open and shut: crystal structures of the dodecylmaltoside solubilized mechanosensitive channel of small conductance from Escherichia coli and Helicobacter pylori at 4.4 Å and 4.1 Å resolutions. Protein Sci. 22, 502–509 (2013).
Pliotas, C. et al. The role of lipids in mechanosensation. Nat. Struct. Mol. Biol. 22, 991–998 (2015).
Zhang, X. et al. Structure and molecular mechanism of an anion-selective mechanosensitive channel of small conductance. Proc. Natl Acad. Sci. USA 109, 18180–18185 (2012).
Brohawn, S. G., Campbell, E. B. & MacKinnon, R. Physical mechanism for gating and mechanosensitivity of the human TRAAK K+ channel. Nature 516, 126–130 (2014).
Guo, Y. R. & MacKinnon, R. Structure-based membrane dome mechanism for Piezo1 mechanosensitivity. eLife 6, e33660 (2017).
Rasmussen, T., Flegler, V. J., Rasmussen, A. & Böttcher, B. Structure of the mechanosensitive channel MscS embedded in the membrane bilayer. J. Mol. Biol. 431, 3081–3090 (2019).
Xue, F. et al. Membrane stiffness is one of the key determinants of E. coli MscS channel mechanosensitivity. Biochim. Biophys. Acta Biomembr. 1862, 183203 (2020).
Nomura, T. et al. Differential effects of lipids and lyso-lipids on the mechanosensitivity of the mechanosensitive channels MscL and MscS. Proc. Natl Acad. Sci. USA 109, 8770–8775 (2012).
Bavi, N. et al. The role of MscL amphipathic N terminus indicates a blueprint for bilayer-mediated gating of mechanosensitive channels. Nat. Commun. 7, 11984 (2016).
Saotome, K. et al. Structure of the mechanically activated ion channel Piezo1. Nature 554, 481–486 (2018).
Reddy, A. S., Warshaviak, D. T. & Chachisvilis, M. Effect of membrane tension on the physical properties of DOPC lipid bilayer membrane. Biochim. Biophys. Acta 1818, 2271–2281 (2012).
Lewis, B. A. & Engelman, D. M. Lipid bilayer thickness varies linearly with acyl chain length in fluid phosphatidylcholine vesicles. J. Mol. Biol. 166, 211–217 (1983).
Belyy, V., Anishkin, A., Kamaraju, K., Liu, N. & Sukharev, S. The tension-transmitting ‘clutch’ in the mechanosensitive channel MscS. Nat. Struct. Mol. Biol. 17, 451–458 (2010).
Ohi, M., Li, Y., Cheng, Y. & Walz, T. Negative staining and image classification — powerful tools in modern electron microscopy. Biol. Proced. Online 6, 23–34 (2004).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Cheng, A. et al. High resolution single particle cryo-electron microscopy using beam-image shift. J. Struct. Biol. 204, 270–275 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Jo, S., Kim, T. & Im, W. Automated builder and database of protein/membrane complexes for molecular dynamics simulations. PLoS ONE 2, e880 (2007).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N⋅ log (N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Steinbach, P. J. & Brooks, B. R. New spherical-cutoff methods for long-range forces in macromolecular simulation. J. Comput. Chem. 15, 667–683 (1994).
Nosé, S. & Klein, M. L. Constant pressure molecular-dynamics for molecular-systems. Mol. Phys. 50, 1055–1076 (1983).
Parrinello, M. & Rahman, A. Polymorphic transitions in single-crystals — a new molecular-dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Hoover, W. G. Canonical dynamics: equilibrium phase-space distributions. Phys. Rev. A Gen. Phys. 31, 1695–1697 (1985).
Nosé, S. A molecular-dynamics method for simulations in the canonical ensemble. Mol. Phys. 52, 255–268 (1984).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Roux, B. The membrane potential and its representation by a constant electric field in computer simulations. Biophys. J. 95, 4205–4216 (2008).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Delcour, A. H., Martinac, B., Adler, J. & Kung, C. Modified reconstitution method used in patch-clamp studies of Escherichia coli ion channels. Biophys. J. 56, 631–636 (1989).
Yang, Z., Fang, J., Chittuluru, J., Asturias, F. J. & Penczek, P. A. Iterative stable alignment and clustering of 2D transmission electron microscope images. Structure 20, 237–247 (2012).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Buchan, D. W. A., Minneci, F., Nugent, T. C. O., Bryson, K. & Jones, D. T. Scalable web services for the PSIPRED protein analysis workbench. Nucleic Acids Res. 41, W349–W357 (2013).
Acknowledgements
We thank D. Rees for critical reading of the manuscript, and M. Ebrahim, J. Sotiris and H. Ng at the Evelyn Gruss Lipper Cryo-EM Resource Center of The Rockefeller University for assistance with data collection. We acknowledge the German Research Foundation DFG (SFB 803, Project A03; to B.d.L.G. and C.D.) and the Alexander von Humboldt foundation (to R.-X.G.) for funding. B.M. is supported by a National Health and Medical Research Council of Australia Principal Research Fellowship (APP1135974). C.D.C. is supported by an NSW Health EMCR Fellowship.
Author information
Authors and Affiliations
Contributions
Y.Z. performed the biochemistry and EM experiments. C.D. and R.-X.G. performed the molecular dynamics experiments. B.d.L.G. supervised the molecular dynamics experiments. C.D.C. and B.M. performed the patch-clamp experiments. T.W. conceived and supervised the project. Y.Z., C.D., R.-X.G., C.D.C., B.M., B.d.L.G. and T.W. analysed the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Architecture of MscS in the closed and open states and image processing of MscS in PC-18:1 nanodiscs.
a, b, Crystal structures of MscS in the closed conformation (PDB: 2OAU) (a) and in the open conformation (PDB: 2VV5) (b). Left panels: ribbon representation of the channel with each subunit shown in a different colour. Right panels: a single subunit with domains labelled. Opening of the channel is accompanied by an increase in the tilt of TM1–TM2 and a rotation about the symmetry axis. c, Area of an image of a vitrified sample. Some particles are circled. Scale bar, 50 nm. d, Selected 2D-class averages obtained with RELION-3. Side length of individual averages, 21.6 nm. e, Image-processing workflow for 3D classification and refinement in RELION-3 that resulted in a density map at 3.2 Å resolution. See Methods and Extended Data Table 1 for details.
Extended Data Fig. 2 Cryo-EM structure of nanodisc-embedded MscS in the closed conformation.
a, Cryo-EM map of MscS in a nanodisc formed with PC-18:1. Subunits are coloured as in Extended Data Fig. 1a and nanodisc density is shown as transparent green surface. b, Density representing one subunit of MscS–PC-18:1 filtered to 5-Å resolution (transparent red surface) with the atomic model built into it (cyan) and the crystal structure of MscS in the closed conformation (PDB: 2OAU; red). c, Hydrophobicity surface map of MscS–PC-18:1 with the model displayed as ribbon. TM1–TM2 and TM3b form a hydrophobic pocket that consists of membrane-buried and solvent-exposed regions (yellow dashed box). d, Secondary structure probabilities for MscS residues 1–89 as predicted by PSIPRED62. Residues with a high propensity to form α-helical secondary structure are labelled with ‘H’ and a pink bar on top. The grey level of the bar above indicates the confidence level of the prediction. e, Helical wheel for N-terminal MscS residues 5–20 calculated with Helixator (http://www.tcdb.org/progs/helical_wheel.php). Most hydrophobic residues locate to the upper half of the wheel, while most hydrophilic residues locate to its lower half. f, The density in the MscS–PC-18:1 map representing the N-terminal segment and the model built into the map. Residue Gln21 forms a break between the end of the amphipathic helix and the beginning of TM1. g, The nanodisc density (green transparent surface) shown at different contouring levels. The MscS structure is shown in ribbon representation and is coloured according to hydrophobicity. At a high contouring threshold (0.013), the nanodisc density is reduced to a thin belt at the cytoplasmic bilayer boundary. No such belt is seen at the periplasmic bilayer boundary. Furthermore, the cytoplasmic bilayer boundary remains largely stationary with decreasing threshold levels, whereas the periplasmic bilayer boundary moves further out as more density is included. These observations suggest that the positively charged residues Arg46 and Arg74 may interact with the lipid headgroups of the cytoplasmic leaflet and define/stabilize the position of this leaflet. h, Close-up view of the location of Arg46 and Arg74 with respect to the nanodisc density contoured at 0.013. The side chains of Arg46 and Arg74 are shown in stick representation.
Extended Data Fig. 3 Image processing of MscS in PC-10 nanodiscs.
a, Area of an image of a vitrified sample. Some particles are circled. Scale bar, 50 nm. b, Selected 2D-class averages obtained with RELION-3. Side length of individual averages, 20.8 nm. c, Image-processing workflow for 3D classification and refinement in RELION-3 that resulted in a density map at 3.4 Å resolution. See Methods and Extended Data Table 1 for details.
Extended Data Fig. 4 Treatment of nanodiscs with βCD and image processing of MscS in PC-18:1 nanodiscs after incubation with βCD for 16 h.
a, Negative-stain EM images of empty nanodiscs incubated with βCD at the conditions denoted, indicating that βCD can extract lipids from nanodiscs. Scale bar, 100 nm. Five to ten images were collected for each condition. b, Size-exclusion chromatogram of an MscS–PC-18:1 sample after incubation with 100 mM βCD for 16 h. The insets show negative-stain EM images of the two peak fractions. Scale bar, 50 nm. Five images were collected for each peak fraction. c, Area of an image of a vitrified sample. Some particles are circled. Scale bar, 50 nm. d, Selected 2D-class averages obtained with RELION-3. Side length of individual averages, 21.6 nm. e, Image-processing workflow for 3D classification and refinement in RELION-3 that resulted in a density map at 3.7 Å resolution. See Methods and Extended Data Table 1 for details.
Extended Data Fig. 5 FSC curves, local-resolution maps and local densities.
a, FSC curves calculated between independently refined half maps for the MscS–PC-18:1 map (red), the MscS–PC-10 map (green), and the MscS–PC-18:1–βCD map (purple). b, Cross-validation FSC curves: purple, refined model versus half map 1 used for refinement (work map); green, refined model versus half map 2 not used for refinement (free map); red, refined model versus the combined final map. The similarity of the ‘work’ and ‘free’ curves suggests no substantial over-fitting. c, Local-resolution maps for MscS–PC-18:1, MscS–PC-10, and MscS–PC-18:1–βCD. d, Local cryo-EM densities for MscS–PC-18:1, MscS–PC-10, and MscS–PC-18:1–βCD.
Extended Data Fig. 6 Distance between the cytoplasmic domain and TM1–TM2 in different MscS structures and βCD perfusion activates reconstituted MscS.
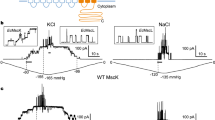
a, The numbers denote the distances between the plane of Leu146 (grey) in the middle β domain and the plane of Asp62 (coloured) at the start of TM2. TM1–TM2 and TM3a are shown as coloured cylinders, and TM3b and the middle β domain as grey ribbons. The trapezoids represent the overall shape of the TMD in the different conformations. The trapezoid shape is defined by the plane connecting Ser58 and Ala28 on TM1. Trapezoids with larger angles would fit better in flatter membranes. b, Set up of excised azolectin proteoliposomes and perfusion of βCD (brown). c, Exemplar trace (black line) showing an initial application of a square wave negative pressure pulse of −45 mmHg (red line) with a high-speed pressure clamp that activates a population of MscS channels at +30 mV pipette potential, followed by the perfusion of 10 mM βCD and the subsequent activation of MscS with no further applied hydrostatic pressure. βCD alone without applied pressure activates the whole cohort of MscS channels in the excised patch with a prolonged plateau indicative of current saturation. Activation is followed by transition of the channel population towards a closed state and the experiment ends with the rupture of the membrane patch.
Extended Data Fig. 7 Two-dimensional class averages and 3D maps of MscS mutant proteins.
a, Two-dimensional class averages of the Gly113Ala MscS mutant in PC-18:1 nanodiscs after incubation with βCD for 16 h obtained with RELION-3. Side length of individual averages, 23.4 nm. While some classes show an extended and defined TMD as seen in 2D averages of MscS in the closed conformation or a compressed and smeared-out TMD or a compressed and asymmetric TMD, none of the classes shows a defined TMD architecture as the one seen in MscS–PC-18:1–βCD. b, Three-dimensional maps of the Gly113Ala MscS mutant in PC-18:1 nanodiscs after incubation with βCD for 16 h obtained from 3D classification in RELION-3. None of the maps shows a TMD architecture similar to the one seen in MscS–PC-18:1–βCD. c, Two-dimensional class averages of the Ala106Val MscS mutant in PC-18:1 nanodiscs obtained with RELION-3. Side length of individual averages, 21.6 nm. While some classes show an extended and defined TMD as seen in 2D averages of MscS in the closed conformation, the TMD in classes with a compressed TMD is smeared out and/or asymmetric. d, Three-dimensional maps of the Ala106Val MscS mutant in PC-18:1 nanodiscs obtained from 3D classification in RELION-3. The only two maps with a defined TMD show MscS in the closed conformation (grey and yellow maps). None of the other maps has a well-defined TMD. Therefore, none of the maps shows MscS in the open conformation seen in the crystal structure of the Ala106Val mutant. e, Two-dimensional class averages of the Ala106Val MscS mutant in DDM obtained with RELION-3. Side length of individual averages, 20.8 nm. f, Three-dimensional maps of the Ala106Val MscS mutant in DDM obtained from 3D classification in RELION-3. The map with the best defined TMD (red) shows MscS in the same open conformation seen in the crystal structure of the Ala106Val mutant. g, Final map at 3.6 Å resolution after refining the map with the best defined TMD (red map in b).
Extended Data Fig. 8 Two-dimensional class averages and 3D maps of MscS in PC-18:1 nanodiscs after incubation with βCD for 30 min and 4 h.
a, Two-dimensional class averages obtained with RELION-3 for MscS in PC-18:1 nanodiscs after a 30-min incubation with βCD. Side length of individual averages, 21.6 nm. Most classes show an extended and defined TMD as seen for MscS in the closed conformation. b, Three-dimensional maps for the MscS in PC-18:1 nanodiscs incubated for 30 min with βCD obtained from 3D classification in RELION-3. The map with the best defined TMD (purple) shows MscS in the closed conformation but with weaker density for TM1. None of the maps shows MscS in the open conformation seen in the crystal structure of the Ala106Val mutant. c, Final map at 3.5 Å resolution after refining the map with the best defined TMD (purple map in b). d, Two-dimensional class averages obtained with RELION-3 for MscS in PC-18:1 nanodiscs after a 4-h incubation with βCD. Side length of individual averages, 21.6 nm. The class averages show different thicknesses for the TMD. e, Three-dimensional maps for the MscS in PC-18:1 nanodiscs incubated for 4 h with βCD obtained from 3D classification in RELION-3. The map with the best defined TMD (purple) shows MscS in the closed conformation but with weaker density for TM1. Another map with a well-defined TMD (grey) shows MscS in the closed conformation with slightly tilted TM1–TM2. None of the maps shows MscS in the open conformation seen in the crystal structure of the Ala106Val mutant.
Extended Data Fig. 9 Lipid densities in the map of MscS in PC-18:1 nanodiscs and snapshots of MD simulations.
a, Cut-away side view of MscS–PC-18:1 colour-coded as in Fig. 4a. The side chains of gating residues Leu105 and Leu109 are shown as purple sticks. Densities for the pore lipids and the inverted gatekeeper lipids are shown in red and light green, respectively. b, Top view of MscS–PC-18:1 showing that the headgroups of the gatekeeper lipids (shown as green sticks in transparent green surface) are shielded from the membrane. c, d, Side view (c) and top view (d) of the area indicated by the dashed box in b, showing the residues interacting with the acyl chains (c) and headgroup (d) of the gatekeeper lipid. The protein residues are coloured by hydrophobicity as in Extended Data Fig. 2c. e, The protein is shown as transparent white ribbon with two subunits forming a hydrophobic pocket shown in solid white and orange. For lipids other than those in the pockets, only the phosphorous atoms are shown as olive-green spheres. The lipids occupying the pocket formed by the orange subunits are shown in cyan stick representation with the phosphorous atoms as olive-green spheres. On average, each pocket holds 3.3 ± 0.2 PC-18:1 lipids in the closed conformation, 1.5 ± 0.1 PC-10 lipids in the open state, and 0.6 ± 0.1 PC-10 lipid in the subconducting state.
Extended Data Fig. 10 Estimating the membrane tension in MscS-containing nanodiscs induced by βCD treatment.
a, The 2D-class averages of MscS in PC-18:1 nanodiscs before (upper panel) and after (lower panel) incubation with βCD obtained with the iterative stable alignment and clustering algorithm. Side length of individual averages, 22 nm. b, Distribution of nanodisc diameters measured from the averages shown in a. The average diameter of the MscS-containing nanodiscs is 93.6 ± 5.9 Å before incubation with βCD and 90.4 ± 6.2 Å after incubation with βCD. c, Surface areas occupied by MscS and PC-18:1 lipids in nanodiscs before (top) and after (bottom) incubation with βCD. On the basis of the measured diameters of MscS-containing nanodiscs before and after incubation with βCD, their surface areas (black circles) are 6,877 Å2 [(46.8 Å)2 × 3.14] and 6,415 Å2 [(45.2 Å)2 × 3.14], respectively. The areas occupied by the TMD of MscS (blue areas) in the closed and desensitized states (estimated on the basis of the position of Leu35 in the middle of the transmembrane region of TM1) are very different, namely 2,238 Å2 [(26.7 Å)2 × 3.14] for MscS in the closed conformation and 3,337 Å2 [(32.6 Å)2 × 3.14] for MscS in the desensitized conformation. Therefore, the areas occupied by lipids (brown areas) in nanodiscs containing MscS in the closed conformation are 4,639 Å2 [6,877 Å2 – 2,238 Å2] and 3,078 Å2 [6,415 Å2 – 3,337 Å2] in nanodiscs containing MscS in the desensitized conformation. In summary, incubation with βCD reduces the total area of the nanodiscs (black circles) by only 6.7%, but because the area occupied by MscS increases by 49.1%, the area occupied by the lipids shrinks by 33.6%. d, Representative NanoDrop spectra recorded for MscS-containing nanodiscs with Cy5-labelled PC-18:1 before (blue trace) and after (red trace) incubation with βCD. The ratio of absorbance at 280 nm (MscS) and at 650 nm (Cy5-labelled PC-18:1) was measured to estimate lipid removal. e, Changes in the Cy5-lipid:protein ratio upon incubation of MscS-containing nanodiscs with βCD in three independent experiments. The five individual measurements are shown as green dots. The columns show the averages and the error bars represent the standard deviations. f, Graph of the membrane tension as a function of the percentage of removed lipids determined by MD simulations. Trajectories were divided into blocks of 0.1 μs to evaluate the average and the standard error of the mean (s.e.m.). The s.e.m. in the plot is smaller than the size of the dots. g, Selected 2D-class averages of MscL-containing nanodiscs before (top) and after (bottom) βCD treatment. Side length of individual averages, 20.8 nm.
Supplementary information
Supplementary Information
This file contains further discussion and supportive results for the findings in this study.
Video 1
Morphs between MscS in the closed, open, sub-conducting and desensitized states viewed parallel to the membrane (side view).
Video 2
Morphs between MscS in the closed, open, sub-conducting and desensitized states viewed perpendicular to the membrane (top view).
Video 3
Lipids inside the MscS channel in the open conformation move to the periphery of the channel, where they associate with the transmembrane helices. PC-18:1 lipids were placed into the center of the MscS channel in the open conformation as shown at the beginning of the movie. During the equilibration phase of the MD simulations, the lipids moved towards the periphery of the channel and associated with the transmembrane domains, where they remained during the production phase of the simulations. MscS is shown in white ribbon representation and the lipid acyl chains in cyan spheres representation.
Video 4
For MscS in the closed conformation, lipids diffuse into the solvent-exposed regions of the hydrophobic pockets. MscS in the closed conformation was embedded into a PC-18:1 lipid bilayer with no lipids occupying the solvent-exposed regions of the hydrophobic pockets between neighboring subunits. During MD simulations, lipids spontaneously diffused into these regions and remained there for the remainder of the simulations. The protein is shown as transparent white ribbon with two subunits forming a hydrophobic pocket shown in solid white and orange. For the pore lipids inside the center of the MscS channel and the lipids that remain in the leaflets of the PC-18:1 bilayer during the course of the MD simulation, only the phosphorous atoms are shown as olive-green spheres. For the lipids that move from the cytoplasmic leaflet to the solvent-exposed region of the hydrophobic pocket in between the white and orange subunits, the carbon atoms are shown in blue, oxygen atoms in red, and the phosphorous atoms in tan.
Video 5
The conformational cycle of MscS, seen parallel to the membrane. In the resting state, MscS (shown in gray with two subunits in blue and yellow) is closed and its TMD is extended, matching the thickness of a relaxed membrane. The TM1–TM2 domain has a tilt angle of 32° relative to the membrane normal and aligns with the TM3b helix of the neighboring subunit. Membrane tension (indicated by black arrows) induces the gatekeeper lipids (light green) to dissociate and to either flip or diffuse into the periplasmic membrane leaflet. As a result, MscS can transition from the closed to the open conformation, which is characterized by a ~50° clockwise rotation (when viewed from the periplasm) of the TM1–TM2 domains about the TM3a helices, so that the TM1–TM2 domains align with the TM3b helices of the same subunits. The rotation is accompanied by an increase in the tilt of the TM1–TM2 domain from 32° to 50°, but only by a subtle decrease in the thickness of the TMD. This defined open conformation captured by the X-ray crystal structure appears to exist only in the context of a detergent micelle, whereas the TMD appears to be highly dynamic in the context of a lipid bilayer. With persisting membrane tension, the first pocket lipids (dark green) leave the solvent-exposed regions of the hydrophobic pockets between neighboring subunits. The tilt of the TM1–TM2 domains increases from 50° to 56°, accompanied by a ~5° anti-clockwise rotation, resulting in a somewhat compressed TMD. This conformation of MscS in a sub-conducting state was artificially captured by reconstituting the channel in an unphysiologically thin membrane. The TMD of the channel in the sub-conducting state would likely be also highly dynamic similar to the TMD of the channel in the open state. With persisting membrane tension, the remaining pocket lipids will leave the solvent-exposed pockets (not shown) and MscS will adopt the desensitized state. This transition is characterized by a further anti-clockwise rotation of the TM1–TM2 domains by ~51°, which now align again with the TM3b helices of the neighboring subunits. The tilt of the TM1–TM2 domains increases further from 56° to 67° and the distance from the TM1–TM2 domains to the cytoplasmic domain shortens from 17.2 Å to 11.3 Å, resulting in a very compressed TMD. The very thin TMD in the desensitized state does not match the hydrophobic thickness of the surrounding membrane, and this hydrophobic mismatch provides a driving force for MscS to return to the relaxed closed conformation once membrane tension subsides. This transition resets the tilt of the TM1–TM2 domains from 67° to 32°, resulting in the extended TMD in the resting state. As MscS relaxes into the resting state, it would pick up fresh pore, pocket and gatekeeper lipids, re-sensitizing the channel to membrane tension.
Video 6
The conformational cycle of MscS, seen perpendicular to the membrane from the periplasm. In the resting state, MscS (shown in gray with two subunits in blue and yellow) is closed and the TM3a helices surround a central pore with a diameter of 5 Å. This diameter does allow some ion conduction, requiring pore lipids (red) to completely close the pore. Membrane tension (indicated by black arrows) induces the gatekeeper lipids (light green) to dissociate and the pore lipids to move to the periphery of the pore. The transition from the closed to the open conformation is associated with a rotation of the TM1–TM2 domains that pulls the TM3a helices away from the center of the pore, increasing the diameter of the pore to 13 Å and allowing ions to permeate the pore. The dynamic nature of the TMD in the open conformation may allow the pore lipids to leave the inside of the channel. With persisting membrane tension, as pocket lipids (dark green) leave the solvent-exposed regions of the hydrophobic pockets between neighboring subunits, MscS transitions into a sub-conducting state. The associated conformational change of the TM1–TM2 domains pushes the TM3a helices towards the center of the pore, reducing the pore diameter to 10 Å. With persisting membrane tension, the remaining pocket lipids leave the solvent-exposed pockets (not shown) and MscS adopts the desensitized state. The associated conformational change of the TM1–TM2 domains pushes the TM3a helices further towards the center of the pore and reduces the pore diameter to 6 Å. When membrane tension subsides, the channel returns to the closed state. While the pore diameter does not change much (from 6 Å to 5 Å), the channel will pick up new pore lipids that block the channel in the closed conformation.
Rights and permissions
About this article
Cite this article
Zhang, Y., Daday, C., Gu, RX. et al. Visualization of the mechanosensitive ion channel MscS under membrane tension. Nature 590, 509–514 (2021). https://doi.org/10.1038/s41586-021-03196-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03196-w
This article is cited by
-
Membrane phospholipids control gating of the mechanosensitive potassium leak channel TREK1
Nature Communications (2023)
-
A mechanical-coupling mechanism in OSCA/TMEM63 channel mechanosensitivity
Nature Communications (2023)
-
Open structure and gating of the Arabidopsis mechanosensitive ion channel MSL10
Nature Communications (2023)
-
Mechanosensitive aquaporins
Biophysical Reviews (2023)
-
Structural insights into the Venus flytrap mechanosensitive ion channel Flycatcher1
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.