Abstract
Behavioural experiences activate the FOS transcription factor in sparse populations of neurons that are critical for encoding and recalling specific events1,2,3. However, there is limited understanding of the mechanisms by which experience drives circuit reorganization to establish a network of Fos-activated cells. It is also not known whether FOS is required in this process beyond serving as a marker of recent neural activity and, if so, which of its many gene targets underlie circuit reorganization. Here we demonstrate that when mice engage in spatial exploration of novel environments, perisomatic inhibition of Fos-activated hippocampal CA1 pyramidal neurons by parvalbumin-expressing interneurons is enhanced, whereas perisomatic inhibition by cholecystokinin-expressing interneurons is weakened. This bidirectional modulation of inhibition is abolished when the function of the FOS transcription factor complex is disrupted. Single-cell RNA-sequencing, ribosome-associated mRNA profiling and chromatin analyses, combined with electrophysiology, reveal that FOS activates the transcription of Scg2, a gene that encodes multiple distinct neuropeptides, to coordinate these changes in inhibition. As parvalbumin- and cholecystokinin-expressing interneurons mediate distinct features of pyramidal cell activity4,5,6, the SCG2-dependent reorganization of inhibitory synaptic input might be predicted to affect network function in vivo. Consistent with this prediction, hippocampal gamma rhythms and pyramidal cell coupling to theta phase are significantly altered in the absence of Scg2. These findings reveal an instructive role for FOS and SCG2 in establishing a network of Fos-activated neurons via the rewiring of local inhibition to form a selectively modulated state. The opposing plasticity mechanisms acting on distinct inhibitory pathways may support the consolidation of memories over time.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Ribotag, FFJ snRNA-seq and CUT&RUN data are deposited into the public repository Gene Expression Omnibus (GEO) with accession number GSE158843. All other data will be shared upon reasonable request.
Code availability
Custom code will be provided upon reasonable request.
References
Josselyn, S. A. & Tonegawa, S. Memory engrams: Recalling the past and imagining the future. Science 367, eaaw4325 (2020).
Tanaka, K. Z. et al. The hippocampal engram maps experience but not place. Science 361, 392–397 (2018).
Greenberg, M. E. & Ziff, E. B. Stimulation of 3T3 cells induces transcription of the c-fos proto-oncogene. Nature 311, 433–438 (1984).
Freund, T. F. & Katona, I. Perisomatic inhibition. Neuron 56, 33–42 (2007).
Klausberger, T. et al. Complementary roles of cholecystokinin- and parvalbumin-expressing GABAergic neurons in hippocampal network oscillations. J. Neurosci. 25, 9782–9793 (2005).
Bartos, M. & Elgueta, C. Functional characteristics of parvalbumin- and cholecystokinin-expressing basket cells. J. Physiol. 590, 669–681 (2012).
Yap, E. L. & Greenberg, M. E. Activity-regulated transcription: bridging the gap between neural activity and behavior. Neuron 100, 330–348 (2018).
Ryan, T. J., Roy, D. S., Pignatelli, M., Arons, A. & Tonegawa, S. Engram cells retain memory under retrograde amnesia. Science 348, 1007–1013 (2015).
Glickfeld, L. L. & Scanziani, M. Distinct timing in the activity of cannabinoid-sensitive and cannabinoid-insensitive basket cells. Nat. Neurosci. 9, 807–815 (2006).
Hefft, S. & Jonas, P. Asynchronous GABA release generates long-lasting inhibition at a hippocampal interneuron-principal neuron synapse. Nat. Neurosci. 8, 1319–1328 (2005).
Buzsáki, G. Theta oscillations in the hippocampus. Neuron 33, 325–340 (2002).
Buzsáki, G. & Wang, X. J. Mechanisms of gamma oscillations. Annu. Rev. Neurosci. 35, 203–225 (2012).
Hasselmo, M. E. & Stern, C. E. Theta rhythm and the encoding and retrieval of space and time. Neuroimage 85, 656–666 (2014).
Sørensen, A. T. et al. A robust activity marking system for exploring active neuronal ensembles. eLife 5, e13918 (2016).
Fenno, L. E. et al. Targeting cells with single vectors using multiple-feature Boolean logic. Nat. Methods 11, 763–772 (2014).
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuron 71, 995–1013 (2011).
Roth, B. L. DREADDs for neuroscientists. Neuron 89, 683–694 (2016).
Xue, M., Atallah, B. V. & Scanziani, M. Equalizing excitation–inhibition ratios across visual cortical neurons. Nature 511, 596–600 (2014).
Vierbuchen, T. et al. AP-1 transcription factors and the BAF complex mediate signal-dependent enhancer selection. Mol. Cell 68, 1067–1082 (2017
Fleischmann, A. et al. Impaired long-term memory and NR2A-type NMDA receptor-dependent synaptic plasticity in mice lacking c-Fos in the CNS. J. Neurosci. 23, 9116–9122 (2003)
Hrvatin, S. et al. Single-cell analysis of experience-dependent transcriptomic states in the mouse visual cortex. Nat. Neurosci. 21, 120–129 (2018).
Sanz, E. et al. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc. Natl Acad. Sci. USA 106, 13939–13944 (2009).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Mo, A. et al. Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86, 1369–1384 (2015).
Bloodgood, B. L., Sharma, N., Browne, H. A., Trepman, A. Z. & Greenberg, M. E. The activity-dependent transcription factor NPAS4 regulates domain-specific inhibition. Nature 503, 121–125 (2013).
Hensch, T. K. Bistable parvalbumin circuits pivotal for brain plasticity. Cell 156, 17–19 (2014).
Nedivi, E., Hevroni, D., Naot, D., Israeli, D. & Citri, Y. Numerous candidate plasticity-related genes revealed by differential cDNA cloning. Nature 363, 718–722 (1993).
Fischer-Colbrie, R., Laslop, A. & Kirchmair, R. Secretogranin II: molecular properties, regulation of biosynthesis and processing to the neuropeptide secretoneurin. Prog. Neurobiol. 46, 49–70 (1995).
Földy, C., Malenka, R. C. & Südhof, T. C. Autism-associated neuroligin-3 mutations commonly disrupt tonic endocannabinoid signaling. Neuron 78, 498–509 (2013).
Hartzell, A. L. et al. NPAS4 recruits CCK basket cell synapses and enhances cannabinoid-sensitive inhibition in the mouse hippocampus. eLife 7, e35927 (2018).
Weiler, R. et al. Chromogranins in rat brain: characterization, topographical distribution and regulation of synthesis. Brain Res. 532, 87–94 (1990).
Franklin, K. B. J. & Paxinos, G. The Mouse Brain in Stereotaxic Coordinates 3rd edn (Academic Press/Elsevier, 2007).
Miyoshi, G. et al. Genetic fate mapping reveals that the caudal ganglionic eminence produces a large and diverse population of superficial cortical interneurons. J. Neurosci. 30, 1582–1594 (2010).
Kenner, L. et al. Mice lacking JunB are osteopenic due to cell-autonomous osteoblast and osteoclast defects. J. Cell Biol. 164, 613–623 (2004).
Sharma, N. et al. ARNT2 tunes activity-dependent gene expression through NCoR2-mediated repression and NPAS4-mediated activation. Neuron 102, 390–406 (2019).
Lin, Y. et al. Activity-dependent regulation of inhibitory synapse development by Npas4. Nature 455, 1198–1204 (2008).
Ataman, B. et al. Evolution of Osteocrin as an activity-regulated factor in the primate brain. Nature 539, 242–247 (2016).
Mardinly, A. R. et al. Sensory experience regulates cortical inhibition by inducing IGF1 in VIP neurons. Nature 531, 371–375 (2016).
Habib, N. et al. Div-Seq: Single-nucleus RNA-seq reveals dynamics of rare adult newborn neurons. Science 353, 925–928 (2016).
Cembrowski, M. S., Wang, L., Sugino, K., Shields, B. C. & Spruston, N. Hipposeq: a comprehensive RNA-seq database of gene expression in hippocampal principal neurons. eLife 5, e14997 (2016).
Hainer, S. J. & Fazzio, T. G. High-resolution chromatin profiling using CUT&RUN. Curr. Protoc. Mol. Biol. 126, e85 (2019).
Malik, A. N. et al. Genome-wide identification and characterization of functional neuronal activity-dependent enhancers. Nat. Neurosci. 17, 1330–1339 (2014).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Yatsenko, D. et al. DataJoint: managing big scientific data using MATLAB or Python. Preprint at bioRxiv https://doi.org/10.1101/031658 (2015).
Pachitariu, M., Steinmetz, N., Kadir, S., Carandini, M. & Harris, K. D. Kilosort: realtime spike-sorting for extracellular electrophysiology with hundreds of channels. Preprint at bioRxiv https://doi.org/10.1101/061481 (2016).
Rossant, C. et al. Spike sorting for large, dense electrode arrays. Nat. Neurosci. 19, 634–641 (2016).
Barthó, P. et al. Characterization of neocortical principal cells and interneurons by network interactions and extracellular features. J. Neurophysiol. 92, 600–608 (2004).
Berens, P. CircStat: a MATLAB toolbox for circular statistics. J. Stat. Softw. 31, (2009).
Bokil, H., Andrews, P., Kulkarni, J. E., Mehta, S. & Mitra, P. P. Chronux: a platform for analyzing neural signals. J. Neurosci. Methods 192, 146–151 (2010).
Acknowledgements
We thank all members of the Greenberg laboratory for discussions and critical feedback during the course of this work; all members of the Harvey laboratory for discussions and critical feedback; W. Regehr, D. Ginty, R. Wilson, C. Tzeng, J. Green, E. Pollina and T. Vierbuchen for feedback and critical evaluation of the data and/or manuscript; G. Fishell for the Dlx5/6Flp mice; L. Wu and the Harvard Genome Modification Facility for aiding in the generation of the Scg2fl/fl mice; C. Wang and the Boston Children’s Hospital Viral Core for generation of AAVs; the Harvard Neurobiology Imaging Facility (NINDS P30 Core Center grant NS072030) for imaging support; and O. Mazor and P. Gorelik at the Harvard Research Instrumentation Core for technical design and support. This work was supported by NIH grants R01 NS028829 and R01 NS115965 (M.E.G.), NIH grant R01 NS089521 (C.D.H.), T32 NS007473 (C.P.D.), F32 NS112455 (C.P.D.), Stuart H.Q. and Victoria Quan fellowship (E-L.Y. and N.L.P.), Harvard Department of Neurobiology graduate fellowship (E.-L.Y.), and Aramont Fund for Emerging Science Research fellowship (E.-L.Y). In addition the Greenberg laboratory is supported by the Allen Discovery Center program, a Paul G. Allen Frontiers Group advised program of the Paul G. Allen Family Foundation.
Author information
Authors and Affiliations
Contributions
E.-L.Y. and M.E.G. conceived and designed the project. E.-L.Y. designed, executed and analysed all ex vivo electrophysiology, molecular biology, FFJ snRNA-seq and Ribotag experiments with input on all aspects from M.E.G., S.R. and E.C.G. E.-L. Y. designed and validated strategy for generation of Scg2fl/fl. E.-L.Y., N.L.P., C.D.H. and M.E.G. conceived and designed, E.-L.Y. and N.L.P. executed, and N.L.P. analysed in vivo silicon probe recordings. E.-L.Y. and C.P.D. designed and executed, and C.P.D. analysed CUT&RUN. M.A.N. and N.S. assisted in analysis of FFJ snRNA-seq. D.A.H. analysed Ribotag sequencing. E.-L.Y., E.G. and N.L.P. executed and analysed Morris water maze experiments. O.D. designed, executed and analysed NE snRNA-seq. C.L. and N.S. assisted in molecular biology. E.-L.Y. and M.E.G. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Hyung-Bae Kwon, Elly Nedivi and Thomas Sudhof for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of novel environment paradigm, AAV-based mKate2 activity reporter, and intersectional genetic strategy for CCK-INs.
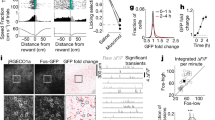
a, Top, representative immunostaining images of FOS and NPAS4 in hippocampus obtained from mice housed under standard (Strd) conditions (left) or exposed to NE (right) for 6 h. Scale, 400 μm. Bottom, higher magnification of insets. Scale, 100 μm. To immunostain for both FOS and NPAS4 proteins in the same sections, mice where FOS or NPAS4 had been endogenously tagged with a Flag-HA tag (Fos-FlagHA19 or Npas4-FlagHA35) were used with a rat anti-HA antibody, while the reciprocal protein was probed with a rabbit polyclonal antibody (Methods). b, Left, number of FOS+ and NPAS4+ nuclei in the CA1 of standard or 6 h NE mice. Strd, N = 6 mice; NE, N = 6 mice. Note that within CA1, significantly fewer NPAS4+ cells were detected, indicating that the AAV-based mKate2 activity reporter mainly labels Fos-activated neurons. Two-sided t-test, ***P = 1.6 × 10−4, *P = 0.033. Right, quantification of number of NPAS4+ cells that are also FOS+. c, Representative images of mKate2+ neurons across different time points and conditions as in d. An AAV encoding GFP was used as a control for the viral injections. Scale, 100 μm. d, Percentages of mKate2+ neurons over total number of DAPI+ cells (left y-axis) or density of mKate2+ neurons (right y-axis). The average percentages of mKate2+ neurons are 1%, 12%, 66% and 96% under Strd (N = 13 mice), 2-3 d NE (N = 10 mice, ***P = 2.7 × 10−4), 7-10 d NE (N = 15 mice, ****P < 1 × 10−15), and 24 h post-KA injection (N = 3 mice, ****P = 7.3 × 10−10), respectively. Ordinary one-way ANOVA, with multiple comparisons correction. Note that data for Strd and 2-3 d NE are replotted from Fig. 1c. e, Bar plots of additional electrophysiological parameters for mKate2— and mKate2+ neurons. n = 30 pairs/4 mice per group. Two-sided t-test, not significant (n.s.) for all parameters. f, Left, schematic of intersectional genetic strategy involving Dlx5/6Flp;CCKCre mice transduced with a dual Cre/Flp recombinase-dependent ChR2EYFP fusion protein necessary to specifically target CCK-INs. Middle, representative immunostaining for PV in magenta; ChR2EYFP in green. Right, percentage of ChR2+ cells in the CA1 field showing overlap with PV expression is low, indicating that the Dlx5/6Flp;CCKCre line is suited for genetic targeting of CCK-INs. N = 4 mice. Scale, 40 μm. g, Representative image of CA1 region of CCKCre mice transduced with AAV encoding Cre-dependent EYFP depicting widespread EYFP expression in the CA1 and underscoring the necessity of the intersectional strategy in f for targeting CCK-INs specifically. N = 2 mice. Scale, 100 μm. In b, d–f, data are mean ± s.e.m. In f, g, schematic images were adapted with permission from Paxinos & Franklin (ref. 32).
Extended Data Fig. 2 IN-to-CA1 PC paired recordings and cell health parameters in 24 h post-KA condition.
a, g, Schematic of genetic strategy to label PV-INs (PVCre;Ai14) or CCK-INs (Dlx5/6Flp;CCKCre;Ai65). b, h, Representative images of tdTomato fluorescence in the CA1 field. Scale, 100 μm. N = 2 mice per line. c, i, Quantification of the fraction of (c) PV- or (i) CCK-to-CA1 PC synaptically-connected pairs from the overall number of pairs recorded in both vehicle (Veh.) and 24 h post-KA mice. (c) Veh., n = 13/22; KA, n = 19/30; (i) Veh., n = 16/40; KA, n = 16/3, where n = number of connections/total pairs. d, j, Quantification of maximum firing rate of (d) PV- or (j) CCK-INs from connected pairs. (d) Veh., n = 10/6; KA, n = 14/7; (j) Veh., n = 15/9; KA, n = 14/4, where n = number of cells/number of mice. e, k, Quantification of spike adaptation ratio of (e) PV- or (j) CCK-INs from connected pairs as in d, j. f, l, Quantification of paired pulse ratios (PPRs) of uIPSCs at the indicated interstimulus intervals (ISI) for (f) PV- (Veh., n = 13/6; KA, n = 19/7) or (l) CCK- (Veh., n = 16/9; KA, n = 16/4) to-CA1 PC connected pairs, where n = number of pairs/number of mice. Two-sided t-tests performed at each ISI or for all ISIs comparing Veh. and 24 h post-KA conditions; *P = 0.039, ****P = 4.4 × 10−5. m, n, Representative hippocampal images from (m) Veh. and (n) 24 h post-KA conditions. Sections were immunostained for NeuN (green) and cleaved-caspase 3 (red), and counterstained with Hoechst (blue). Scale, 200 μm (left); 100 μm (right, CA1 field). N = 2 mice per condition. o–q, Quantification of (o) Hoechst+ nuclei, (p) NeuN+ nuclei, and (q) cleaved-caspase+ cells per 40-μm section in all layers of CA1. Ori, stratum oriens; Pyr, stratum pyramidale; Rad, stratum radiatum; Lac, stratum lacunosum moleculare. Results suggest that KA injection does not induce cell death within 24 h. Veh. and KA, n = 10 sections/2 mice, respectively. In d–f, j–l, o–q, data are mean ± s.e.m.
Extended Data Fig. 3 Chemogenetic activation of CA1 PCs recapitulated bidirectional changes in perisomatic inhibition while silencing of CA1 PCs led to inverse effects.
a–d, f, g, Top, schematic of recording configuration. Bottom, scatter plots of (a, c, d, f) PV- or (b, g) CCK-IPSCs recorded from untransduced WT and indicated viral-transduced neighbouring CA1 PCs. (a) Veh., n = 16/5; CNO, n = 16/7; (b) Veh., n = 22/5; CNO, n = 21/7; (c) CNO, n = 16/4; (d) Note: pairs of untransduced cells, CNO, n = 8/3; (f) KirMut, n = 18/3; Kir2.1, n = 19/5; (g) KirMut, n = 25/3; Kir2.1, n = 17/4, where n = number of pairs/number of mice and each open circle represents a pair of simultaneously recorded neurons, with closed circles representing mean ± s.e.m. e, Representative trace of spikes detected from a CA1 PC in cell-attached mode in slice after bath application of CNO. As expected, addition of CNO led to firing rate increases in hM3DGq-expressing neurons, providing further confidence that intraperitoneal CNO injection in mice in vivo chemogenetically activates hM3DGq-expressing neurons in the CA1. N = 3 cells/3 mice. Scale, 50 pA, 60 s.
Extended Data Fig. 4 Validation of Fosfl/fl;Fosbfl/fl;Junbfl/fl (FFJ) mouse line and additional electrophysiological parameters in FFJ-WT and KO cells.
a, Schematic representation of the AP-1 members conditionally deleted in FFJ line. b, c, Representative images of smRNA-FISH validating loss of Fos and Fosb (and Junb in c) upon Cre expression in the CA1 field of 1-1.5 h post-KA-injected FFJ mice. Scale, 20 μm. N = 4 mice. d, Normalized pixel intensity for Cre-negative and Cre-positive cells. Each point represents the average for individual sections across N = 4 mice. Two-sided t-tests, Fos, ***P = 7.7 × 10−4; Fosb, *P = 0.031; Junb, *P = 0.047. e, Scatter plots of normalized pixel intensities of Cre signal against Fos, Fosb or Junb signals for each cell. Pearson correlation coefficients (r) shown. Fos, n = 315; Fosb, n = 86; Junb, n = 229 cells from N = 4 mice. f, Representative images of Cre-injected sections immunostained for FOS, FOSB, JUNB, and NPAS4 proteins in the CA1 field of 3 h post-KA-injected FFJ mice. Scale, 100 μm. N = 3 mice. g, j, m, Schematic of stimulus electrode (stim. elec.) placement in stratum pyramidale (s.p.) to stimulate perisomatic inhibitory axons (g), or stratum radiatum (s.r.) to stimulate Schaffer collaterals (j) or proximal dendritic inhibitory axons (m). h, k, n, Scatter plots of recorded pairs of FFJ-WT and FFJ-KO CA1 PCs in 24 h post-vehicle (left) or -KA injected (right) mice, where (h) Veh., n = 26/6; KA, n = 33/7; (k) Veh., n = 18/5; KA, n = 17/4; (n) Veh., n = 30/4; KA, n = 30/6. i, l, o, Quantification of PPRs for indicated currents, where (i) Veh., n = 17/3; KA, n = 18/4; (l) Veh., n = 18/5; KA, n = 17/4; (o) Veh., n = 19/2; KA, n = 26/5. In h, i, k, l, n, o, n = number of pairs/number of mice. In d, h, i, k, l, n, o, data are mean ± s.e.m.
Extended Data Fig. 5 RNA-sequencing to identify CA1 pyramidal neuron-specific FOS targets.
a, Scatter plot showing PV-specific ARGs identified by comparing 6 h post-KA to vehicle-injected conditions. Significantly different genes (green); FDR ≤ 0.005. PV-enriched (IP over input) genes (red). Points represent mean ± s.e.m. n = 9-10 mice per biological replicate; 4 biological replicates per condition. b, UMAP visualization of IN subtypes using only Gad2-expressing (“Inhibitory”) cells from Fig. 3d. c, UMAP visualization of ΔCre+ and respective control nuclei with (left) cell type information or (right) genotype assignments overlaid. Control, ΔCre— in control hemispheres; ΔCre-GFP, ΔCre+ in injected hemispheres; Other, ΔCre— or ΔCre+ in injected or control hemispheres, respectively. n = 25,214 cells/4 mice. d, Quality control metrics for each transcriptionally distinct cell type identified by snRNA-seq in both Cre+ and ΔCre+ (“Del”) samples compared with their respective untransduced controls (“WT”) as in Fig. 3d and c. Top, number of unique genes per cell. Middle, number of RNA molecules per cell. Bottom, percentage of reads that map to mitochondrial genome. CA1, CA1 PCs; Pv, Pvalb+ INs; Cck, Cck+ INs; Sst, Sst+ INs; Vip, Vip+ INs; Nos1, Nos1+ INs; Npy, Npy+ INs; CA3, CA3 excitatory neurons; DG, dentate gyrus neurons; Cck Exc, Cck+ excitatory neurons; OPCs, oligodendrocyte precursor cells; Oligo, oligodendrocytes; Astro, astrocytes; Micro, microglia; Endo, endothelial cells. e, Violin plots depicting CA1 PC-specific expression of Fos (****P = 9.7 × 10−127), Fosb, Junb (****P = 7.2 × 10−26; **P = 0.003), and viral-derived WPRE (****P = 0). Ctrl, untransduced control nuclei. Note that the design of the FFJ line renders snRNA-seq validation of excision of Fosb and Junb suboptimal (see Extended Data Fig. 4b–f and Methods). TPT, tags per ten thousand. f, Strip plot displaying differential gene expression between Cre+ and control samples for each transcriptionally distinct cell type. Colored points represent significant genes (Bonferroni-corrected P < 0.05, with average natural log fold-change (FC) ≥ 20%); grey points represent non-significant genes. g, Heatmap depicting normalized gene expression values from 100 randomly selected cells from each indicated cell type identity. Genes are cell-type-enriched AP-1 targets downregulated by at least 20% with loss of AP-1, and whose expression is detected in at least 25% of untransduced cells. h, Volcano plot of shuffled data where Cre+ and control CA1 excitatory nuclei were randomly assigned between two groups, showing no significant gene expression differences (light grey; Bonferroni-corrected P > 0.05), thus further indicating that the expression differences observed between Cre+ and control were due to presence of Cre. Data are mean ± 2 × s.d. (d, e); two-sided Wilcoxon rank-sum test (e–h).
Extended Data Fig. 6 CaMK2a-SUN1 FOS CUT&RUN revealed FOS binding sites across genome.
a, Pairwise Pearson correlation between CaMK2a-SUN1 FOS CUT&RUN biological replicates for each antibody and stimulus condition. b, Histogram plotting distribution of distances between CaMK2a-SUN1 FOS CUT&RUN peaks and the nearest Refseq transcription start site (TSS). Peaks with a distance of 0 overlap the TSS. As expected42, ~90% of FOS-bound sites are distal to the TSS. c–e, Histograms plotting distributions of distances between the TSS of (c) all Refseq genes, (d) CaMK2a-Ribotag ARGs, or (e) CA1 excitatory genes downregulated with AP-1 loss (FFJ snRNA-seq), and the nearest FOS binding site. A distance of 0 indicates overlap of a FOS peak with the TSS. Notably, both CaMK2a-specific ARGs (d) and putative AP-1 targets downregulated with AP-1 loss in FFJ snRNA-seq (e) are significantly enriched for FOS-bound sites, which are significantly closer to the TSS when compared to all genes (c) (P < 2.2 × 10−16, two-sided Wilcoxon rank-sum test), providing further support that these genes are direct targets of FOS. f, Top three enriched motifs identified by MEME-ChIP from CaMK2a-SUN1 FOS CUT&RUN peaks. E-values and matching transcription factor motifs are displayed to the right of each enriched motif. FOS CUT&RUN peaks identified therefore show significant enrichment for the AP-1 motif. g–k, Tracks displaying FOS or IgG binding under 2-3 h post-vehicle or KA conditions for genomic regions surrounding the (g) Bdnf, (h) Inhba, (i) Rgs2, (j) Nptx2, or (k) Pcsk1 genes (see Fig. 4i for Scg2). y-axis shows spike-in normalized CUT&RUN coverage. Tracks are scaled to the maximum value observed for all samples for the displayed genomic locus, shown in brackets.
Extended Data Fig. 7 Analyses of AP-1-regulated candidate genes to identify molecular effector(s) of bidirectional perisomatic inhibitory plasticity.
a, Table of high-confidence AP-1-regulated candidate genes analysed and their known functions. b, RT–qPCR validation of shRNA efficacy using cultured hippocampal neurons transduced with lentivirus encoding the indicated shRNA. n = 3 biological replicates for each shRNA. Data are mean ± s.e.m. c, Western blot confirmation of the efficacy of the Flp-OFF shRNA strategy, where Bdnf shRNA-containing plasmid was transfected in 293T cells along with BDNF-MYC, and excision of the shRNA expression cassette via introduction of Flp recombinase was confirmed. Loading controls (GAPDH) were run on a separate blot (see Supplementary Fig. 2a for full scans). 100- or 500-ng transfections of indicated u6-plasmid were loaded side-by-side on blot. n = 2 biological replicates. d–f, Scatter plots of recorded PV-IPSC amplitudes from untransduced shRNA— and neighbouring shRNA+ CA1 PCs from mice 24 h post-KA injection. The shRNA target is shown on the y-axis: (d) Control, n = 17/9; Inhba, n = 15/4; Rgs2, n = 20/3; Bdnf, n = 26/10; Nptx2, n = 16/3; Pcsk1, n = 17/6; (e) Scg2 shRNA#2, n = 17/6. Representative traces from a pair of neurons shown; blue marks depict light onset. Scale, 100 pA, 40 ms; (f) Scg2 shRNA#1, Strd, n = 14/5; 7-10 d NE, n = 16/4, where n = number of pairs/number of mice. Each open circle represents a pair of simultaneously recorded neurons; closed circles represent mean ± s.e.m. g, smRNA-FISH scatter plots as in Fig. 4k depicting the correlation between Fos and (left) Scg2 intron or (right) Scg2 mRNA expression. Each point represents the mean number of Scg2 puncta/cell within a bin, with a bin width of 1 Fos punctum/cell. Pearson correlation coefficients (r) are shown. h, Lower magnification images of smRNA-FISH as in Fig. 4j. Scale, 100 μm.
Extended Data Fig. 8 SCG2 is a molecular effector of bidirectional perisomatic inhibitory plasticity.
a, RT–qPCR validation of Scg2fl/fl conditional-knockout line, where normalized (left) Scg2 and (right) Fos RNA levels in cultured hippocampal neurons derived from Scg2fl/fl mice are shown. Cultures were transduced with lentiviral Cre or ΔCre and membrane depolarized with KCl for 0 h or 6 h. n = 3 biological replicates. Data are mean ± s.e.m.; two-sided t-test, **P = 0.002. b, Schematic of intersectional genetic strategy to introduce ChR2 into CCK-INs and sparsely introduce shRNAs specifically into CA1 PCs of Dlx5/6Flp;CCKCre mice. c, Normalized differences in CCK-IPSC amplitudes between pairs of Scg2 shRNA— and shRNA+ PCs depicted in d–f. Strd, n = 30/4; NE, n = 24/3; KA, n = 19/4. Data are mean ± s.e.m. Ordinary one-way ANOVA, with multiple comparisons correction; NE, **P = 0.005; KA, **P = 0.002. d–f, Scatter plots of CCK-IPSC amplitudes of pairs as in c. Representative traces from pairs of neurons shown; blue marks depict light onset. Scale, 100 pA, 40 ms. g, Top, schematic of recording configuration. Scatter plots of (bottom left) PV-IPSC or (bottom right) CCK-IPSC amplitudes recorded from pairs of neurons of which one was untransduced (WT) and the other expressed a Scg2 shRNA#1 with an shRNA-resistant full-length SCG2 rescue construct. Normalized differences in IPSC amplitudes between pairs of neurons shown to the right of each scatter plot. PV, n = 19/6; CCK, n = 19/4. Two-sided one-sample t-test with hypothetical mean of 0, *P = 0.011. In c–g, each open circle represents a pair of simultaneously recorded neurons; closed circles represent mean ± s.e.m.; n = number of pairs/number of mice.
Extended Data Fig. 9 A series of rescue and overexpression analyses suggest a critical role for the processing of SCG2.
a, b, Scatter plots of PV-IPSC (a) and CCK-IPSC (b) amplitudes recorded from mKate2+ pairs that are either Cre— (WT) or Cre+ (KO). Scg2-KO neurons also expressed a Cre-dependent full-length SCG2 construct (Rescue WT) to rescue the loss of Scg2. PV, n = 22/5; CCK, n = 27/3. c, d, As in a, b but using a Cre-dependent non-cleavable SCG2 mutant (Rescue 9AA) instead, which failed to rescue the loss of Scg2. PV, n = 23/4; CCK, n = 23/4. e, f, Scatter plots of PV-IPSC (e) and CCK-IPSC (f) amplitudes recorded from untransduced (WT) and neighbouring full-length SCG2-overexpressing CA1 PCs (OE WT), showing that gain-of-function of SCG2 is sufficient to induce bidirectional perisomatic inhibitory plasticity in the absence of neural activity. PV, n = 20/5; CCK, n = 25/3. g, Western blot confirmation of stable expression of SCG2 and the non-cleavable SCG2 mutant (9AA-Mutant) constructs containing an HA-tag in 293T cells. Expression levels were measured by immunoblot analysis with HA antibody. Loading controls (GAPDH) were run on a separate blot (see Supplementary Fig. 2b for full scans). n = 2 biological replicates. h, i, As in e, f but with overexpression of the non-cleavable SCG2 mutant (9AA Mutant) instead, which failed to induce changes in inhibition. PV, n = 19/4; CCK, n = 16/3. In a–f, h, i, each open circle represents a pair of simultaneously recorded neurons; closed circles represent mean ± s.e.m; n = number of pairs/number of mice.
Extended Data Fig. 10 Silicon probe recordings in Scg2-WT and Scg2-KO mice to assess effects on network oscillations.
a, Left, schematic of stereotaxic injection and recording site in CA1 pyramidal layer. Right, representative image of silicon probe placement in CA1 pyramidal layer with Cre-GFP (green) and Dil (red). N = 4 mice. Scale, 200 μm. b, Normalized power spectra of network oscillations in Scg2-WT or Scg2-KO mice during stationary periods. Average across Scg2-WT (grey, N = 4) or Scg2-KO (green, N = 5) mice, one session per mouse. Data are mean ± s.e.m. c, Mean of the normalized power spectra within theta, slow-gamma, and fast-gamma bands during stationary periods as shown in b. Two-sided t-test, *P = 0.037. Data are mean ± s.e.m. d, Cumulative histogram of mean firing rate for all Scg2-WT and Scg2-KO units. Mean firing rate is not significantly different (two-sided t-test, P = 0.2138). Scg2-WT (n = 67 units) and Scg2-KO (n = 103 units). e, Example local field potential (LFP), single-unit activity, and running speed in a Scg2-WT mouse. From top to bottom: Denoised and downsampled LFP, 4-12 Hz bandpass filtered LFP, population spiking activity raster plot, and smoothed running speed. f, Expanded snippet of data from the example in e. From top to bottom: Denoised and downsampled LFP, 4-12 Hz bandpass filtered LFP, and population spiking activity raster plot. g, As in f but with example data from a Scg2-KO mouse. Schematic image in a (left) adapted with permission from Paxinos & Franklin (ref. 32).
Supplementary information
Supplementary Figures
This file contains Supplementary Figure 1: Gating strategy for flow cytometry analysis of data shown in Figure 3f,g; and Supplementary Figure 2: Full scans of blots shown in Extended Data Figs. 7c and 9g.
Supplementary Table
This file contains Supplementary Table 1: Gene lists from data shown in Figure 3.
Rights and permissions
About this article
Cite this article
Yap, EL., Pettit, N.L., Davis, C.P. et al. Bidirectional perisomatic inhibitory plasticity of a Fos neuronal network. Nature 590, 115–121 (2021). https://doi.org/10.1038/s41586-020-3031-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-3031-0
This article is cited by
-
Spatial transcriptomics reveal neuron–astrocyte synergy in long-term memory
Nature (2024)
-
Control of neuronal excitation–inhibition balance by BMP–SMAD1 signalling
Nature (2024)
-
A concerted neuron–astrocyte program declines in ageing and schizophrenia
Nature (2024)
-
Synaptic configuration and reconfiguration in the neocortex are spatiotemporally selective
Anatomical Science International (2024)
-
Promoting effect and immunologic role of secretogranin II on bladder cancer progression via regulating MAPK and NF-κB pathways
Apoptosis (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.