Abstract
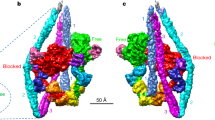
Myosin-2 is essential for processes as diverse as cell division and muscle contraction. Dephosphorylation of its regulatory light chain promotes an inactive, ‘shutdown’ state with the filament-forming tail folded onto the two heads1, which prevents filament formation and inactivates the motors2. The mechanism by which this happens is unclear. Here we report a cryo-electron microscopy structure of shutdown smooth muscle myosin with a resolution of 6 Å in the head region. A pseudo-atomic model, obtained by flexible fitting of crystal structures into the density and molecular dynamics simulations, describes interaction interfaces at the atomic level. The N-terminal extension of one regulatory light chain interacts with the tail, and the other with the partner head, revealing how the regulatory light chains stabilize the shutdown state in different ways and how their phosphorylation would allow myosin activation. Additional interactions between the three segments of the coiled coil, the motor domains and the light chains stabilize the shutdown molecule. The structure of the lever in each head is competent to generate force upon activation. This shutdown structure is relevant to all isoforms of myosin-2 and provides a framework for understanding their disease-causing mutations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The electron density maps for the SmM shutdown heads region and whole molecule have been deposited into EMDB, with accession codes EMD-11069 and EMD-11070, respectively. The pseudo-atomic model of the SmM shutdown heads region has been deposited in PDB, with accession code 6Z47. The following models were used for comparison purposes in our study: PDB 1I84, PDB 3JBH, PDB 5TBY, cardiac IHM model MS01 (http://spudlab.stanford.edu/homology-models) and cardiac IHM model SD232.
References
Lee, K. H. et al. Interacting-heads motif has been conserved as a mechanism of myosin II inhibition since before the origin of animals. Proc. Natl Acad. Sci. USA 115, E1991–E2000 (2018).
Cross, R. A., Cross, K. E. & Sobieszek, A. ATP-linked monomer–polymer equilibrium of smooth muscle myosin: the free folded monomer traps ADP.Pi. EMBO J. 5, 2637–2641 (1986).
McLachlan, A. D. & Karn, J. Periodic charge distributions in the myosin rod amino acid sequence match cross-bridge spacings in muscle. Nature 299, 226–231 (1982).
Craig, R., Smith, R. & Kendrick-Jones, J. Light-chain phosphorylation controls the conformation of vertebrate non-muscle and smooth muscle myosin molecules. Nature 302, 436–439 (1983).
Ikebe, M., Hartshorne, D. J. & Elzinga, M. Identification, phosphorylation, and dephosphorylation of a second site for myosin light chain kinase on the 20,000-dalton light chain of smooth muscle myosin. J. Biol. Chem. 261, 36–39 (1986).
Cross, R. A., Jackson, A. P., Citi, S., Kendrick-Jones, J. & Bagshaw, C. R. Active site trapping of nucleotide by smooth and non-muscle myosins. J. Mol. Biol. 203, 173–181 (1988).
Wendt, T., Taylor, D., Trybus, K. M. & Taylor, K. Three-dimensional image reconstruction of dephosphorylated smooth muscle heavy meromyosin reveals asymmetry in the interaction between myosin heads and placement of subfragment 2. Proc. Natl Acad. Sci. USA 98, 4361–4366 (2001).
Burgess, S. A. et al. Structures of smooth muscle myosin and heavy meromyosin in the folded, shutdown state. J. Mol. Biol. 372, 1165–1178 (2007).
Jung, H. S., Komatsu, S., Ikebe, M. & Craig, R. Head-head and head-tail interaction: a general mechanism for switching off myosin II activity in cells. Mol. Biol. Cell 19, 3234–3242 (2008).
Sellers, J. R. Regulation of cytoplasmic and smooth muscle myosin. Curr. Opin. Cell Biol. 3, 98–104 (1991).
Woodhead, J. L. et al. Atomic model of a myosin filament in the relaxed state. Nature 436, 1195–1199 (2005).
Naber, N., Cooke, R. & Pate, E. Slow myosin ATP turnover in the super-relaxed state in tarantula muscle. J. Mol. Biol. 411, 943–950 (2011).
Stewart, M. A., Franks-Skiba, K., Chen, S. & Cooke, R. Myosin ATP turnover rate is a mechanism involved in thermogenesis in resting skeletal muscle fibers. Proc. Natl Acad. Sci. USA 107, 430–435 (2010).
Alamo, L. et al. Effects of myosin variants on interacting-heads motif explain distinct hypertrophic and dilated cardiomyopathy phenotypes. eLife 6, e24634 (2017).
Nag, S. et al. The myosin mesa and the basis of hypercontractility caused by hypertrophic cardiomyopathy mutations. Nat. Struct. Mol. Biol. 24, 525–533 (2017).
Liu, J., Wendt, T., Taylor, D. & Taylor, K. Refined model of the 10S conformation of smooth muscle myosin by cryo-electron microscopy 3D image reconstruction. J. Mol. Biol. 329, 963–972 (2003).
Jung, H. S. et al. Role of the tail in the regulated state of myosin 2. J. Mol. Biol. 408, 863–878 (2011).
Ikebe, M. et al. Function of the NH2-terminal domain of the regulatory light chain on the regulation of smooth muscle myosin. J. Biol. Chem. 269, 28173–28180 (1994).
Kast, D., Espinoza-Fonseca, L. M., Yi, C. & Thomas, D. D. Phosphorylation-induced structural changes in smooth muscle myosin regulatory light chain. Proc. Natl Acad. Sci. USA 107, 8207–8212 (2010).
Persechini, A. & Hartshorne, D. J. Phosphorylation of smooth muscle myosin: evidence for cooperativity between the myosin heads. Science 213, 1383–1385 (1981).
Persechini, A. & Hartshorne, D. J. Ordered phosphorylation of the two 20 000 molecular weight light chains of smooth muscle myosin. Biochemistry 22, 470–476 (1983).
Ikebe, M., Ogihara, S. & Tonomura, Y. Nonlinear dependence of actin-activated Mg2+-ATPase activity on the extent of phosphorylation of gizzard myosin and H-meromyosin. J. Biochem. 91, 1809–1812 (1982).
Collins, J. H. Myoinformatics report: myosin regulatory light chain paralogs in the human genome. J. Muscle Res. Cell Motil. 27, 69–74 (2006).
Kamm, K. E. & Stull, J. T. Signaling to myosin regulatory light chain in sarcomeres. J. Biol. Chem. 286, 9941–9947 (2011).
Bury, L. et al. Next-generation sequencing for the diagnosis of MYH9-RD: predicting pathogenic variants. Hum. Mutat. 41, 277–290 (2020).
Harakalova, M. et al. Incomplete segregation of MYH11 variants with thoracic aortic aneurysms and dissections and patent ductus arteriosus. Eur. J. Hum. Genet. 21, 487–493 (2013).
Kanematsu, T. et al. A case of MYH9 disorders caused by a novel mutation (p.K74E). Ann. Hematol. 95, 161–163 (2016).
Zaninetti, C. et al. MYH9-related thrombocytopenia: four novel variants affecting the tail domain of the non-muscle myosin heavy chain IIA associated with a mild clinical evolution of the disorder. Hamostaseologie 39, 87–94 (2019).
Yang, S. et al. The central role of the tail in switching off 10S myosin II activity. J. Gen. Physiol. 151, 1081–1093 (2019).
Yang, Y. et al. Rigor-like structures from muscle myosins reveal key mechanical elements in the transduction pathways of this allosteric motor. Structure 15, 553–564 (2007).
Alamo, L. et al. Three-dimensional reconstruction of tarantula myosin filaments suggests how phosphorylation may regulate myosin activity. J. Mol. Biol. 384, 780–797 (2008).
Robert-Paganin, J., Auguin, D. & Houdusse, A. Hypertrophic cardiomyopathy disease results from disparate impairments of cardiac myosin function and auto-inhibition. Nat. Commun. 9, 4019 (2018).
Dominguez, R., Freyzon, Y., Trybus, K. M. & Cohen, C. Crystal structure of a vertebrate smooth muscle myosin motor domain and its complex with the essential light chain: visualization of the pre-power stroke state. Cell 94, 559–571 (1998).
Burgess, S. et al. The prepower stroke conformation of myosin V. J. Cell Biol. 159, 983–991 (2002).
Batchelor, M. et al. Myosin tails and single α-helical domains. Biochem. Soc. Trans. 43, 58–63 (2015).
Wolny, M. et al. Stable single α-helices are constant force springs in proteins. J. Biol. Chem. 289, 27825–27835 (2014).
Blankenfeldt, W., Thomä, N. H., Wray, J. S., Gautel, M. & Schlichting, I. Crystal structures of human cardiac β-myosin II S2-Δ provide insight into the functional role of the S2 subfragment. Proc. Natl Acad. Sci. USA 103, 17713–17717 (2006).
Behrmann, E. et al. Structure of the rigor actin-tropomyosin-myosin complex. Cell 150, 327–338 (2012).
Woodhead, J. L. & Craig, R. The mesa trail and the interacting heads motif of myosin II. Arch. Biochem. Biophys. 680, 108228 (2020).
Alamo, L. et al. Conserved intramolecular interactions maintain myosin interacting-heads motifs explaining tarantula muscle super-relaxed state structural basis. J. Mol. Biol. 428, 1142–1164 (2016).
Straussman, R., Squire, J. M., Ben-Ya’acov, A. & Ravid, S. Skip residues and charge interactions in myosin II coiled-coils: implications for molecular packing. J. Mol. Biol. 353, 613–628 (2005).
Thompson, R. F., Iadanza, M. G., Hesketh, E. L., Rawson, S. & Ranson, N. A. Collection, pre-processing and on-the-fly analysis of data for high-resolution, single-particle cryo-electron microscopy. Nat. Protoc. 14, 100–118 (2019).
Fernandez-Leiro, R. & Scheres, S. H. W. A pipeline approach to single-particle processing in RELION. Acta Crystallogr. D 73, 496–502 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Topf, M. et al. Protein structure fitting and refinement guided by cryo-EM density. Structure 16, 295–307 (2008).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 54, 5.6.1–5.6.37 (2016).
Offer, G., Hicks, M. R. & Woolfson, D. N. Generalized Crick equations for modeling noncanonical coiled coils. J. Struct. Biol. 137, 41–53 (2002).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Pintilie, G. D., Zhang, J., Goddard, T. D., Chiu, W. & Gossard, D. C. Quantitative analysis of cryo-EM density map segmentation by watershed and scale-space filtering, and fitting of structures by alignment to regions. J. Struct. Biol. 170, 427–438 (2010).
Maier, J. A. et al. ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Case, D. A. et al. Amber 2020 (Univ. California, 2020).
Ryckaert, J.-P., Ciccotti, G. & Berendsen, H. Numerical integration of the cartesian equations of motion of a system with constraints: Molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Joosten, R. P., Long, F., Murshudov, G. N. & Perrakis, A. The PDB_REDO server for macromolecular structure model optimization. IUCrJ 1, 213–220 (2014).
Olney, J. J., Sellers, J. R. & Cremo, C. R. Structure and function of the 10 S conformation of smooth muscle myosin. J. Biol. Chem. 271, 20375–20384 (1996).
Brito, R. et al. A molecular model of phosphorylation-based activation and potentiation of tarantula muscle thick filaments. J. Mol. Biol. 414, 44–61 (2011).
Acknowledgements
We thank staff at the Astbury Biostructure Laboratory for their support, namely R. Thompson for advice regarding cryo-EM grid preparation and D. Maskell for image acquisition; and members of the CryoEM community at Leeds for their help and guidance, particularly M. Byrne for advice on structure validation. This work was supported by the Medical Research Council (grant numbers MR/S023593/1 to M.P. and MR/R009406/1 to M.P. and N.A.R.) and by the Wellcome Trust (WT 094231/Z/10/Z to P.J.K., M.P. and N.A.R.). The FEI Titan Krios microscopes were funded by the University of Leeds (UoL ABSL award) and Wellcome Trust (108466/Z/15/Z).
Author information
Authors and Affiliations
Contributions
M.P. and N.A.R. designed the project. J.M.C. purified smooth muscle myosin. C.A.S., D.C.-M. and M.P. performed cryo-EM grid screening and optimization. C.A.S. and D.C.-M. prepared cryo-EM grids for data collection and recorded data. C.A.S., G.C. and D.C.-M. analysed and processed the data. G.C., D.C.-M. and C.A.S. built the model. G.C. performed molecular dynamic simulations. C.A.S., G.C., D.C.-M., M.P. and P.J.K. interpreted the data and the model. M.P. managed the project. M.P., P.J.K. and C.A.S. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Gregory M. Alushin and Robert Cross for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Micrograph, 2-D classification, angular distribution and resolution parameters.
a, Representative micrograph of shutdown SmM molecules from 28,817 micrographs. Scale bar 50 nm. b, Representative 2D classes (representing approximately 25% of particles that contributed to the final reconstruction). Scale bar 10 nm. c, Local resolution of SmM shutdown heads region cryo-EM map. d, FSC curve of the heads region reconstruction, illustrating 6.3 Å resolution at 0.143 FSC. e, Angular distribution of particles in the heads region 3D reconstruction. f, FSC curve of the whole molecule reconstruction. g, Angular distribution of particles contributing to the whole molecule 3D reconstruction.
Extended data Fig. 2 Interaction of segment 3 with blocked-head RLC and analysis of RLC photo-crosslink.
a, Segmented map highlighting latch and mortar regions (boxed), which are shown in more detail in panels b and c respectively. The position of phosphorylated serine (S20) in the latch (b) and in the mortar (c) are shown. This illustrates how phosphorylation would disrupt latch-segment 3 and mortar-RLC interactions respectively. d, Segmented map showing region of interest for panels e-g. e, ionic interactions between segment 3 and blocked-head RLC inter-lobe linker and blocked-head heavy chain. f, Hydrophobic interface where segment 3 crosses the blocked head RLC. g, Same view as e and f showing photo-crosslinker benzophenone-4-acetamide attached to C109 of blocked-head RLC and in close proximity to the aliphatic side chain of K1579 of segment 3 chain H. K1579 lies within crosslinked peptide L1554-E158361. h, Segmented map (face view) to show the location (boxed) of i, which shows an alternative view of photo-crosslinker interaction and the ionic interaction between blocked head RLC E112 and segment 3 R1584. For clarity the non-involved chain of the coiled coil has been omitted in the close-up panels.
Extended Data Fig. 3 Annotated sequence alignment of heavy chains of human non-muscle myosin 2A (NM2A) and smooth muscle myosin (SmM) with Turkey SmM, showing sites of mutation.
HsNM2A: Homo sapiens MYH9; MgSmM: Meleagris gallopavo MYH11; HsSmM: Homo sapiens MYH11. Boxes delimit structural features. Grey background identifies residues in the d position of the coiled coil heptad repeat. Skip residues and bend positions are highlighted. Arrowheads and magenta letters denote known sites of mutation, obtained from the Human Genome Mutation Database (HGMD), Variant of unknown significance: retrieved from the Leiden mutation database.
Extended data Fig. 4 Comparisons of IHM pseudo-atomic models, free- and blocked-heads and the SAH-domain character of start of segment 1.
a-f, Comparison of pseudo-atomic models of the IHM with LCs omitted to allow comparison of LCD heavy-chain structure. a, Pseudo-atomic model of SmM from the present study. b, Model (PDB id 1I84) produced from SmM heavy meromyosin IHM 2D crystal 20Å map7; tentative assignment of tail omitted. c, Model (PDB id 3JBH) produced from IHM of tarantula thick filament 20Å map40, fitted with tarantula sequence. d, Model (PDB id 5TBY) proposed for human cardiac IHM by homology modelling the cardiac amino acid sequences on the tarantula model14. e, Model (downloaded from Spudich lab website as MS01.pdb) of human cardiac myosin IHM (produced by use of homology models fitted to tarantula IHM model 3DTP15). f, Model (downloaded as Supplementary Data 2) for cardiac IHM produced by use of a cardiac motor domain-ELC crystal structure (5N69) and homology model of RLC fitted into the tarantula thick filament IHM 20Å map31. g, Superposition of free- and blocked-head motors (up until pliant region) showing how the LCD regions differ between the heads (without light chains shown), segmented maps and model for motor domains shown. h, Blocked-head motor, model in map, i morph between blocked-head and free-head model and maps (shown without light chains), j Free-head model in map. Map contour level 0.28 (Supplementary Video 3). k, SmM heavy-chain sequence at the start of the predicted coiled coil. Coiled-coil seam a and d residues marked by grey stripes; acidic residues red, basic residues blue. l, Heptad net projection of sequence35 in which the dashed line shows the path of the polypeptide backbone as α-helix, circles and squares indicate the a and d positions of the heptad repeat and every seventh residue is repeated (in brackets) to allow all ionic interactions to be mapped.
Extended Data Fig. 5 Interaction between loop 2 (blocked-head motor) and segment 1, the motor-motor interface of the IHM and fit of the SmM IHM structure into filament cryo-EM density maps for tarantula and cardiac myosin filaments.
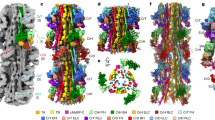
a-b, EM density map reveals an interaction interface between segment 1 and density attributable to the blocked-head motor loop 2 (dashed line). E913 in segment 1 is highlighted for reference. Map contour level 0.17. c. Overall model to show region of interest, the motor-motor interface (boxed). d, Ionic interactions across this interface, involving Helix N and HCM loop of the blocked-head motor with 3 successive residues (168-170) close to Helix E in the free-head motor (Supplementary Video 5). e, The same region of interest from a recent cardiac IHM pseudo-atomic model15. f, Superimposition of the motor domains for our structure, and for the cardiac IHM pseudo-atomic model15 aligned on the free-head (red/pink) with view as in d. The blocked head of the cardiac IHM (cyan) is rotated counter-clockwise compared to that for SmM (blue) g, h, Fit of the paired heads of the pseudo-atomic model of shutdown SmM into the IHM motifs of reconstructions of thick filaments, using Chimera. The filament axis is vertical and the filament tip is at the top of the page. g, The fit to 20 Å map of tarantula thick filaments (EMD-195062, contour level 28.6). h. The fit to 28 Å map of cardiac thick filaments (EMD-224014, contour level 0.14).
Extended Data Fig. 6 Flexibility of SmM tail ribbon.
a, Representative class averages showing flexibility of the tail of myosin molecules (Supplementary Video 7), after alignment of the heads. Numbers at lower left indicate the chord angle of the tail at a position 29.7 nm along the tail from its emergence point from the head (taken to be the fulcrum). The angle is defined in relation to the path of the tail between bend 2 and the fulcrum, see the cartoon at the right. 4895 face view particles were classified into 50 classes of which 5 are shown to represent the range of motion of the tail. b, Cumulative frequency plot of tail angles for myosin with fitted Gaussian curve, yielding a variance value of 34°. Scale bar in a: 20 nm.
Extended Data Fig. 7
Diagram to show the dynamic relationship of the shutdown state (10S) to the open ‘active’ state (6S) and filament assembly. The shutdown state and filaments can both compete for 6S.
Supplementary information
Video 1
: Shutdown state of SMM (a) Cryo-EM density of SmM (grey) shutdown heads region at an average resolution of 6 Å. Segmented map representation showing heavy chain (blue, red), ELC (purple, orange), and RLC (magenta, green) in complex with segments 2 and 3 (light blue, pink) and fitted pseudo-atomic model. (b) Closeup view of blocked-head RLC latch (lilac) with potential interactions with ring of acidic residues (red spheres) in segment-3 and D132 in its own RLC highlighted. (c) Close up view of the free-head RLC mortar (dark green) with potentially interacting acidic residues (red spheres) in both RLCs.
Video 2
: Interactions between Segment-2 and the blocked head motor (a) Segmented map density with fitted quasi-atomic model showing ionic interactions between segment-2 (chain G, light blue) and the blocked head motor (blue). (b) Ionic interactions between segment-2 (chain H, pink) and the blocked head ELC (orange). (c) Ionic interactions between segment-2 (chain H, pink) and segment-3 (chain G, light blue) around bend 2.
Video 3
: Morphing of the free-head motor domain to blocked-head motor domain This shows changes in LCD and converter position (extended fig 4g-j). Motors aligned up until the pliant point (residues 30-787).
Video 4
: Closeup view of the head-tail junction This shows nearly unbroken α-helices (red, blue) between the LCD, ending at P849, and the coiled-coil tail.
Video 5
: Motor-motor domain interactions that stabilise the shutdown state Related to Extended Data Fig. 5.
Video 6
: Whole Molecule SMM 6 Å map fitted to our 9 Å map, which reveals more of the tail domain. Full length model of SmM fitted to 9Å map, with skip residues (yellow) showing the perturbation to the super-helical twist of the coiled coil.
Video 7
: Flexing of whole molecule SMM 2D class averages of the whole molecule SmM arranged to show the range of flexing of the tail in plane. Particles in each class shown bottom left. Scale bar 20 nm.
Rights and permissions
About this article
Cite this article
Scarff, C.A., Carrington, G., Casas-Mao, D. et al. Structure of the shutdown state of myosin-2. Nature 588, 515–520 (2020). https://doi.org/10.1038/s41586-020-2990-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2990-5
This article is cited by
-
A simple and rapid preparation of smooth muscle myosin 2 for the electron microscopic analysis
Applied Microscopy (2024)
-
Cryo-EM structure of the folded-back state of human β-cardiac myosin
Nature Communications (2023)
-
Review on the structural understanding of the 10S myosin II in the era of Cryo-electron microscopy
Applied Microscopy (2022)
-
Alpha and beta myosin isoforms and human atrial and ventricular contraction
Cellular and Molecular Life Sciences (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.