Abstract
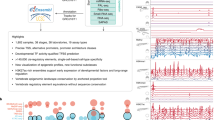
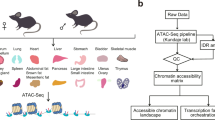
The zebrafish (Danio rerio) has been widely used in the study of human disease and development, and about 70% of the protein-coding genes are conserved between the two species1. However, studies in zebrafish remain constrained by the sparse annotation of functional control elements in the zebrafish genome. Here we performed RNA sequencing, assay for transposase-accessible chromatin using sequencing (ATAC-seq), chromatin immunoprecipitation with sequencing, whole-genome bisulfite sequencing, and chromosome conformation capture (Hi-C) experiments in up to eleven adult and two embryonic tissues to generate a comprehensive map of transcriptomes, cis-regulatory elements, heterochromatin, methylomes and 3D genome organization in the zebrafish Tübingen reference strain. A comparison of zebrafish, human and mouse regulatory elements enabled the identification of both evolutionarily conserved and species-specific regulatory sequences and networks. We observed enrichment of evolutionary breakpoints at topologically associating domain boundaries, which were correlated with strong histone H3 lysine 4 trimethylation (H3K4me3) and CCCTC-binding factor (CTCF) signals. We performed single-cell ATAC-seq in zebrafish brain, which delineated 25 different clusters of cell types. By combining long-read DNA sequencing and Hi-C, we assembled the sex-determining chromosome 4 de novo. Overall, our work provides an additional epigenomic anchor for the functional annotation of vertebrate genomes and the study of evolutionarily conserved elements of 3D genome organization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All the sequencing data are deposited in the NCBI Gene Expression Omnibus under accession code GSE134055. All the genomic data generated in this study can be visualized in the WashU Epigenome Browser (https://epigenome.wustl.edu/zebrafishENCODE/). The human histone-modification ChIP-seq data were downloaded from the ROADMAP Project. The mouse histone modification ChIP-seq data were downloaded from the mouse ENCODE Consortium. The human tissue transcriptome data were downloaded from the GTEx Consortium. The public zebrafish ChIP-seq and ATAC-seq data used in this study are listed in Supplementary Table 6. The human h1-ESC Hi-C data were downloaded from GSE52457. GM12878 and K562 GRO-seq data were downloaded from GSE60456. GM12878 and K562 CTCF ChIP-seq were downloaded from GSE31477. GM12878 and K562 Pol2 ChIP-seq were downloaded from GSE91426 and GSE31477. Source data are provided with this paper.
References
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Gerhard, G. S. et al. Life spans and senescent phenotypes in two strains of Zebrafish (Danio rerio). Exp. Gerontol. 37, 1055–1068 (2002).
Lamason, R. L. et al. SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 310, 1782–1786 (2005).
Vastenhouw, N. L. et al. Chromatin signature of embryonic pluripotency is established during genome activation. Nature 464, 922–926 (2010).
Bogdanovic, O. et al. Dynamics of enhancer chromatin signatures mark the transition from pluripotency to cell specification during embryogenesis. Genome Res. 22, 2043–2053 (2012).
Kaaij, L. J. et al. Enhancers reside in a unique epigenetic environment during early zebrafish development. Genome Biol. 17, 146 (2016).
Aday, A. W., Zhu, L. J., Lakshmanan, A., Wang, J. & Lawson, N. D. Identification of cis regulatory features in the embryonic zebrafish genome through large-scale profiling of H3K4me1 and H3K4me3 binding sites. Dev. Biol. 357, 450–462 (2011).
Vesterlund, L., Jiao, H., Unneberg, P., Hovatta, O. & Kere, J. The zebrafish transcriptome during early development. BMC Dev. Biol. 11, 30 (2011).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Yue, F. et al. A comparative encyclopedia of DNA elements in the mouse genome. Nature 515, 355–364 (2014).
Anderson, J. L. et al. Multiple sex-associated regions and a putative sex chromosome in zebrafish revealed by RAD mapping and population genomics. PLoS ONE 7, e40701 (2012).
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019).
Meuleman, W. et al. Index and biological spectrum of human DNase I hypersensitive sites. Nature 584, 244–251 (2020).
Quillien, A. et al. Robust identification of developmentally active endothelial enhancers in zebrafish using FANS-assisted ATAC-seq. Cell Rep. 20, 709–720 (2017).
Letelier, J. et al. Evolutionary emergence of the rac3b/rfng/sgca regulatory cluster refined mechanisms for hindbrain boundaries formation. Proc. Natl Acad. Sci. USA 115, E3731–E3740 (2018).
Liu, G., Wang, W., Hu, S., Wang, X. & Zhang, Y. Inherited DNA methylation primes the establishment of accessible chromatin during genome activation. Genome Res. 28, 998–1007 (2018).
Marlétaz, F. et al. Amphioxus functional genomics and the origins of vertebrate gene regulation. Nature 564, 64–70 (2018).
Meier, M. et al. Cohesin facilitates zygotic genome activation in zebrafish. Development 145, dev156521 (2018).
Torbey, P. et al. Cooperation, cis-interactions, versatility and evolutionary plasticity of multiple cis-acting elements underlie krox20 hindbrain regulation. PLoS Genet. 14, e1007581 (2018).
Paik, E. J. et al. A Cdx4–Sall4 regulatory module controls the transition from mesoderm formation to embryonic hematopoiesis. Stem Cell Reports 1, 425–436 (2013).
Kang, J. et al. Modulation of tissue repair by regeneration enhancer elements. Nature 532, 201–206 (2016).
Kaufman, C. K. et al. A zebrafish melanoma model reveals emergence of neural crest identity during melanoma initiation. Science 351, aad2197 (2016).
Goldman, J. A. et al. Resolving heart regeneration by replacement histone profiling. Dev. Cell 40, 392–404 (2017).
Pérez-Rico, Y. A. et al. Comparative analyses of super-enhancers reveal conserved elements in vertebrate genomes. Genome Res. 27, 259–268 (2017).
Lister, R. et al. Global epigenomic reconfiguration during mammalian brain development. Science 341, 1237905 (2013).
Visel, A. et al. Ultraconservation identifies a small subset of extremely constrained developmental enhancers. Nat. Genet. 40, 158–160 (2008).
Dimitrieva, S. & Bucher, P. UCNEbase–a database of ultraconserved non-coding elements and genomic regulatory blocks. Nucleic Acids Res. 41, D101–D109 (2013).
Visel, A., Minovitsky, S., Dubchak, I. & Pennacchio, L. A. VISTA Enhancer Browser—a database of tissue-specific human enhancers. Nucleic Acids Res. 35, D88–D92 (2007).
Corces, M. R. et al. The chromatin accessibility landscape of primary human cancers. Science 362, eaav1898 (2018).
Neph, S. et al. Circuitry and dynamics of human transcription factor regulatory networks. Cell 150, 1274–1286 (2012).
Yang, T. et al. HiCRep: assessing the reproducibility of Hi-C data using a stratum-adjusted correlation coefficient. Genome Res. 27, 1939–1949 (2017).
Krefting, J., Andrade-Navarro, M. A. & Ibn-Salem, J. Evolutionary stability of topologically associating domains is associated with conserved gene regulation. BMC Biol. 16, 87 (2018).
Lazar, N. H. et al. Epigenetic maintenance of topological domains in the highly rearranged gibbon genome. Genome Res. 28, 983–997 (2018).
Fishman, V. et al. 3D organization of chicken genome demonstrates evolutionary conservation of topologically associated domains and highlights unique architecture of erythrocytes’ chromatin. Nucleic Acids Res. 47, 648–665 (2019).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Smagulova, F. et al. Genome-wide analysis reveals novel molecular features of mouse recombination hotspots. Nature 472, 375–378 (2011).
Canela, A. et al. Genome organization drives chromosome fragility. Cell 170, 507–521 (2017).
Gothe, H. J. et al. Spatial chromosome folding and active transcription drive DNA fragility and formation of oncogenic MLL translocations. Mol. Cell 75, 267–283 (2019).
Canela, A. et al. Topoisomerase II–induced chromosome breakage and translocation is determined by chromosome architecture and transcriptional activity. Mol. Cell 75, 252–266 (2019).
Postlethwait, J. H. et al. Vertebrate genome evolution and the zebrafish gene map. Nat. Genet. 18, 345–349 (1998).
Pedroso, G. L. et al. Blood collection for biochemical analysis in adult zebrafish. J. Vis. Exp. 3865, e3865 (2012).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Wang, L. et al. CPAT: Coding-Potential Assessment Tool using an alignment-free logistic regression model. Nucleic Acids Res. 41, e74 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Maertin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, Z. et al. Identification of transcription factor binding sites using ATAC-seq. Genome Biol. 20, 45 (2019).
Korhonen, J., Martinmäki, P., Pizzi, C., Rastas, P. & Ukkonen, E. MOODS: fast search for position weight matrix matches in DNA sequences. Bioinformatics 25, 3181–3182 (2009).
Kulakovskiy, I. V. et al. HOCOMOCO: towards a complete collection of transcription factor binding models for human and mouse via large-scale ChIP-seq analysis. Nucleic Acids Res. 46 (D1), D252–D259 (2018).
Gerstein, M. B. et al. Integrative analysis of the Caenorhabditis elegans genome by the modENCODE project. Science 330, 1775–1787 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liu, T. Use model-based Analysis of ChIP-Seq (MACS) to analyze short reads generated by sequencing protein-DNA interactions in embryonic stem cells. Methods Mol. Biol. 1150, 81–95 (2014).
Hiller, M. et al. Computational methods to detect conserved non-genic elements in phylogenetically isolated genomes: application to zebrafish. Nucleic Acids Res. 41, e151 (2013).
Lee, H. J. et al. Regenerating zebrafish fin epigenome is characterized by stable lineage-specific DNA methylation and dynamic chromatin accessibility. Genome Biol. 21, 52 (2020).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Zhou, X., Li, D., Lowdon, R. F., Costello, J. F. & Wang, T. methylC Track: visual integration of single-base resolution DNA methylation data on the WashU EpiGenome Browser. Bioinformatics 30, 2206–2207 (2014).
Burger, L., Gaidatzis, D., Schübeler, D. & Stadler, M. B. Identification of active regulatory regions from DNA methylation data. Nucleic Acids Res. 41, e155 (2013).
Wu, H. et al. Detection of differentially methylated regions from whole-genome bisulfite sequencing data without replicates. Nucleic Acids Res. 43, e141 (2015).
Hansen, K. D., Langmead, B. & Irizarry, R. A. BSmooth: from whole genome bisulfite sequencing reads to differentially methylated regions. Genome Biol. 13, R83 (2012).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44 (W1), W160–W165 (2016).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Robinson, J. T. et al. Juicebox. js provides a cloud-based visualization system for Hi-C data. Cell Syst. 6, 256–258 (2018).
Abdennur, N. & Mirny, L. A. Cooler: scalable storage for Hi-C data and other genomically labeled arrays. Bioinformatics 36, 311–316 (2020).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Darling, A. E., Mau, B. & Perna, N. T. progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 5, e11147 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Johansen, N. & Quon, G. scAlign: a tool for alignment, integration, and rare cell identification from scRNA-seq data. Genome Biol. 20, 166 (2019).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Acknowledgements
This work was supported by NIH grants R35GM124820, R01HG009906, R24DK106766 (R.C.H. and F.Y.) and R01DK107735 (G.S.G.). F.Y. is also supported by U01CA200060. T.W. is supported by NIH grants R01HG007175, R01HG007354, R01ES024992, U24ES026699 and U01HG009391. We thank J. A. Stamatoyannopoulos for discussion and suggestions; H. Lyu for proof reading and other Yue lab members for discussion; and E. DeForest, S. Stella, P. Hubley and Penn State Zebrafish Functional Genomics Core for fish husbandry and embryo collection.
Author information
Authors and Affiliations
Contributions
F.Y. conceived and supervised the project. H.Y. and T.L. collected tissue and conducted experiments. Y.L. led the data analysis. Y.L., H.Y., H.J.L., Y.W., X.W., B.Z., L.F. and J.W. conducted analyses. D.L. and T.W. provided the website for data presentation. K.C.A. and K.C.C. provided animal support. Q.J., X.X., J.X., F.S., I.S., C. K., T.S., M.N.K.C., J.T., K.W., G.S.G., R.C.H., T.W. and K.C.C. helped with data interpretation. H.Y., Y.L., T.L. and F.Y. prepared the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Michael Beer, Jesse Dixon and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
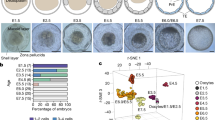
Extended Data Fig. 1 Tissue-specific gene expression in zebrafish.
a, Clustering analysis of transcripts from RNA-seq data in embryonic and adult tissues (n = 31,842). b, c, Gene Ontology and KEGG pathway analysis for the tissue-specific genes in adult brain, heart and testis (the number of tissue-specific genes in these two figures are, Brain = 3,693, Heart = 392, Testis = 1,605). d, Distribution of H3K4me3 signals surrounding the known and predicted novel transcripts. e, Human orthologues of zebrafish tissue-specific genes were more tissue-specific compared to human orthologues of non-tissue-specific zebrafish genes (n = 14,764, 3,739, 6,043, Mann–Whitney U Test, two-sided, ***P < 2.2 × 10−16).
Extended Data Fig. 2 Comparative analysis of zebrafish cis-regulatory elements.
a, Comparison of the predicted regulatory elements identified with previous data. Enhancers were based on H3K27ac signals in the same four tissues (brain, heart, intestine, testis) from Perez-Rico et al. 2017. The data we generated are from Tübingen zebrafish strain and the published results were from the AB strain. b, Number of predicted cis-regulatory elements in each tissue. E-brain stands for 1 dpf embryonic neuron cells. E-trunk stands for 1 dpf zebrafish whole trunk region. c, An example showing genes with active promoters have higher expression level. Blue hollow bar indicates the known mrpl39 promoter. Orange hollow bar indicates the potential novel promoter. The mrpl39 promoter has H3K4me3 peaks in both muscle and brain, but only has strong H3K27ac signals in muscle and its expression is higher (4.43-fold). d, Gene Ontology results for the muscle-specific enhancers and skin-specific enhancers. We used the GREAT tool for this analysis (the numbers of tissue-specific enhancers used in this figure are muscle = 813, skin = 512).
Extended Data Fig. 3 Enhancer reporter assay for tissue-specific enhancers.
In total, 28 of 32 predicted tissue-specific enhancers showed consistent GFP signals in the corresponding tissues. For the eight brain enhancers tested, 63/95, 51/86, 85/119, 112/143, 27/45, 34/48, 27/41, 62/77, and 37/45 embryos, respectively, had green signals in the brain region. For the six tested heart enhancers, 64/94, 52/85, 79/121, 20/41, 51/95, 32/55 and 20/31 embryos, respectively, had green signals in the heart region. For the six tested muscle enhancers, 52/57, 26/30, 107/124, 53/63, 93/114, 61/67 and 66/78 embryos, respectively, had green signals in the trunk muscle. For the four selected kidney enhancers, 47/82, 35/67, 44/62, 15/42 and 56/110 embryos, respectively, had green signals in the kidney region.
Extended Data Fig. 4 Single-cell ATAC-seq in zebrafish brain.
a, Barcode selection of single cell ATAC-seq. The x-axis represents the log value of the number of unique molecular identifiers (UMI); the y axis represents the ratio of fragments in promoter regions; the red lines represent threshold, and the grey shadows represent that the barcode passed the filter. b, Genomic distribution of all differentially accessible (DA) peaks. c, Overlap of all differentially accessible peaks with enhancers predicted in bulk brain. d, Top, the cluster distribution in the tSNE projection. Bottom left, pileups of differentially accessible ATAC-seq signals for each cluster. Shown in the figure is the +/− 10kb flanking region surrounding peak centres. Bottom right, most significantly enriched transcription factor motif for each cluster. e, t-SNE projection of all scATAC-seq cells colored by Z-score of peak enrichment. f, Motif enrichment of known neuron-specific TFs in scATAC-seq predicted clusters (n = 19,955).
Extended Data Fig. 5 Heterochromatin annotation in adult tissues.
a, WashU Epigenome Browser screenshot of H3K9me3 and H3K9me2 histone ChIP-seq signals in 11 zebrafish adult tissues. The values on the y-axis were input-normalized. b, Distribution of H3K9me3 and H3K9me2 sites in the zebrafish genome. c, Venn diagram shows the overlap between H3K9me3 and H3K9me2 sites in zebrafish genome. d, Overlapping percentile of H3K9me3 and H3K9me2 peaks in adult tissues. e, H3K9me3 and H3K9me2 sites were depleted of ATAC-seq, H3K4me3 and H3K27ac ChIP-seq signals (n = 68,789 H3K9me3 sites and n = 73,777 H3K9me2 sites). f, Overlap of H3K9me3 sites, H3K9me2 sites, and ATAC-seq peaks with repetitive elements (The total number of each bar, from left to right, 68,789, 73,777 and 436,036). g, Examples of H3K9me3 sites in one tissue found to be active regions in other tissues. Horizontal scale 0-20 for H3K27ac and H3K4me3, 0-10 for RNA-seq, 0-5 for H3K9me3 and H3K9me2.
Extended Data Fig. 6 DNA methylation level and distribution in adult tissues.
a, Fraction of total CpGs with low (<25%), medium (≥25% and <75%), and high (≥75%) methylation levels and mean CpG methylation levels (mCG/CG) in zebrafish adult tissues (the mCG/CG ratio, from left to right, 0.788, 0.859, 0.790, 0.777, 0.791, 0.797, 0.781, 0.777, 0.804, 0.789, 0.781). b, Distribution of CpG methylation levels across zebrafish adult tissues. c, The distribution of non CpG methylation in 11 adult tissues. d, Mean methylation levels of the tissue-specific gene promoters. n represents the number of tissue-specific gene promoter. e, Mean methylation level of CpGs overlapping different genomic features or repetitive element classes. CDS, coding sequence. f, Number of UMRs and LMRs in zebrafish tissues and their overlap with enhancer and promoters (left panel) (number of UMR and LMR, from top to bottom, 14,990, 10,569, 14,569, 14,587, 14,831, 14,289, 13,842, 13,569, 14,424, 14,374, 13,908, 30,009, 7,916, 19,038, 21,411, 22,591, 16,796, 14,961, 16,268, 17,481, 15,932, 15,665) and ATAC-seq peaks (right panel)(numbers of UMR and LMR are the same with left panel). g, Clustering of tissue-specific hypoDMRs. Values in the heat map are mean methylation levels of hypoDMRs (n = 17,654, number of tissue-specific hypoDMRs).
Extended Data Fig. 7 De novo assembly of zebrafish chromosome 4 of the Tübingen strain.
a, WashU Epigenome Browser snapshot showing that heterochromatic marks H3K9me2 and H3K9me3 signals were enriched on chromosome 4 in zebrafish testis. The values on the y-axis were input-normalized. b, H3K9me2, H3K9me3, and DNA methylation level on chr4 long arm are significantly higher than other regions in all tissues (n = 11, two-sided, t-test). c, Overall strategy of de novo assembly of the Tübingen chr4 by integrating 10X, Nanopore, Bionano, and Hi-C data. d, Bionano long molecule sequencing data shows that there were many SVs on chr4 when mapped to the GRCz11 reference genome. e, SVs on chr4 detected by Bionano when the data were mapped to the de novo assembled chr4.
Extended Data Fig. 8 Conservation of cis-regulatory elements from zebrafish to other vertebrates.
a, Percentage of zebrafish enhancers whose sequences were conserved in human (the number of each bar, from left to right, 13,307, 7,018, 11,940, 7,499, 14,783, 14,272, 8,995, 13,777, 10,757, 15,505, 1,734, 4,011, 5,247). b, c, Similar to Fig. 4a. Percentage of zebrafish exons and cis-regulatory elements that have orthologous sequences in mouse and other fish species. Total number of each bar, from left to right: 1,000, 25,593, 58,065, 1,000. For exons and random, we randomly sample 1000 elements and computed their conservation percentage. The simulations were performed 20 times and the average percentage was presented. d, Another example of ultra-conserved noncoding element (UCNE). This element (FOXP1_Finn_1) is predicted to be a muscle enhancer in zebrafish, mouse, and human. Grey vertical bar marks the ultra-conserved region. Red vertical bar is the enhancer sequence in the human genome that was validated as a limb enhancer by transgenic mouse reporter assay in the VISTA Enhancer Browser (#hs956).
Extended Data Fig. 9 Distal ATAC-seq peak-to-gene pairs, enhancer-to-gene pairs, and transcriptional regulation network.
a, b, Distance distribution of cis-regulatory elements to their linked gene TSS. c, Correlation of ATAC-seq peak-to-gene pairs and Enhancer-to-gene pairs (n from left to right = 3,292, 3,827, 3,544, 3,281, 3,008, 2,795, 2,357, 2,001, 1,106). d, Validation of predicted enhancer-to-gene pairs by Hi-C interaction counts in muscle. e, mef2d is a regulator in both zebrafish muscle and heart, but it regulates different downstream targets by motif prediction analysis. f, The overall structure of the regulatory network is conserved between human and zebrafish. FFL connection analysis was performed, in this analysis, there are three types of nodes: A, driver node that regulates B and C; B, middle node, regulated by A but regulating node C; C, passenger node, regulated by both A and B.
Extended Data Fig. 10 Compartment and TADs in zebrafish.
a, Heat map of genome-wide Hi-C interaction matrices in zebrafish brain (blue) and muscle (red). b, Active marks (H3K4me3, H3K27ac, and ATAC-seq) were enriched in compartment A and depleted in compartment B. Repressive marks (H3K9me2 and H3K9me3) were enriched in compartment B. Error bands represent standard error of the mean. c, Genome browser snapshot of A/B compartment in brain and muscle. The blue vertical shaded area marks a region that is located in compartment B in brain but in compartment A in muscle. As expected, A compartment which is associated with more ATAC-seq peaks, H3K27ac and RNA-seq signals. d, Examples of shared TADs between zebrafish brain and muscle. e, Average DI scores surrounding TAD boundaries identified in brain (upper panel) and muscle (lower panel). f, ChIP-seq data shows that CTCF binding sites were enriched at TAD boundaries. g, Footprint analysis of ATAC-seq peaks in the TAD boundaries shows enrichment of CTCF binding motif (number of each bar, from left to right, 0.213, 0.24, 0.22, 0.237, 0.251, 0.232, 0.24, 0.262, 0.271, 0.281, 0.37, 0.27, 0.253, 0.25, 0.252, 0.253, 0.26, 0.23, 0.238, 0.24, 0.22). h, Repetitive elements enriched at TAD boundaries (left panel) and loop anchors (right panel).
Extended Data Fig. 11 Comparing zebrafish evolutionary breakpoints with TAD annotation.
a. Similar to Fig. 5d. Enrichment of evolutionary breakpoints at TAD boundaries. Relative positions of evolutionary breakpoints to TADs in 15 vertebrates. In all cases, we found that the evolutionary breakpoints were enriched at zebrafish TAD boundaries and depleted from the centre of TADs. Grey vertical bar labels the TAD body area. b, By comparing zebrafish with 17 vertebrates, H3K4me3 signals were found to be more enriched at TAD boundaries with breakpoints than those without breakpoints. Orange vertical bar labels the TAD boundaries. c, Higher H3K4me3 levels at breakpoint-containing TAD boundaries when using TADs annotation from zebrafish muscle were found as well, similar to Fig. 5g. d, H3K4me3 enrichment in human ESCs (H1) TAD boundaries with or without zebrafish-to-human breakpoints. e, H3K4me3 enrichment in mouse ESCs TAD boundaries with or without zebrafish-to-mouse breakpoints. f, H3K4me3 enrichment in human ESCs (H1) TAD boundaries with or without mouse-to-human breakpoints.
Extended Data Fig. 12 TADs with and without breakpoints.
a, H3K27ac and ATAC-seq signals do not show differences at TAD boundaries with breakpoints compared to those without breakpoints. Orange vertical bar labels the TAD boundaries. b, Sizes of TADs with and without evolutionary breakpoints were similar (n = 573, 777, two-sided, t-test). c, Enrichment of transcription at breakpoints (BP) that overlap with CTCF TAD boundaries in K562 cells (the number of breakpoints in blue line is 639, red line is 625). d, In 17 vertebrates, TADs without evolutionary breakpoints (bottom panel) have stronger interaction frequencies in the middle than TADs with evolutionary breakpoints (upper panel). Breakpoints in these 17 vertebrates were defined by comparing their genomes to the zebrafish genome. e, Distribution of correlations between the expression pattern of each pair of paralogs across 11 adult zebrafish tissues. f, Correlations between pairs of paralogs located on the same chromosome. Among them, 17 pairs were located within the same TAD, and the rest of the 65 pairs were located in different TADs. As a control, we randomly sampled 100 genes. Number of each bar, from left to right, 17, 65, 100.
Supplementary information
Supplementary Information
This file contains a guide to the Supplementary Tables 1-19.
Supplementary Tables
This zip file contains Supplementary Tables 1-19.
Supplementary Data
This zip file contains Supplementary Dataset 1.
Source data
Rights and permissions
About this article
Cite this article
Yang, H., Luan, Y., Liu, T. et al. A map of cis-regulatory elements and 3D genome structures in zebrafish. Nature 588, 337–343 (2020). https://doi.org/10.1038/s41586-020-2962-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2962-9
This article is cited by
-
Pan-3D genome analysis reveals structural and functional differentiation of soybean genomes
Genome Biology (2023)
-
Unraveling the whole genome DNA methylation profile of zebrafish kidney marrow by Oxford Nanopore sequencing
Scientific Data (2023)
-
Upf3a but not Upf1 mediates the genetic compensation response induced by leg1 deleterious mutations in an H3K4me3-independent manner
Cell Discovery (2023)
-
Comparative analysis of genome-scale, base-resolution DNA methylation profiles across 580 animal species
Nature Communications (2023)
-
Long-read sequencing of the zebrafish genome reorganizes genomic architecture
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.