Abstract
Gene-expression programs define shared and species-specific phenotypes, but their evolution remains largely uncharacterized beyond the transcriptome layer1. Here we report an analysis of the co-evolution of translatomes and transcriptomes using ribosome-profiling and matched RNA-sequencing data for three organs (brain, liver and testis) in five mammals (human, macaque, mouse, opossum and platypus) and a bird (chicken). Our within-species analyses reveal that translational regulation is widespread in the different organs, in particular across the spermatogenic cell types of the testis. The between-species divergence in gene expression is around 20% lower at the translatome layer than at the transcriptome layer owing to extensive buffering between the expression layers, which especially preserved old, essential and housekeeping genes. Translational upregulation specifically counterbalanced global dosage reductions during the evolution of sex chromosomes and the effects of meiotic sex-chromosome inactivation during spermatogenesis. Despite the overall prevalence of buffering, some genes evolved faster at the translatome layer—potentially indicating adaptive changes in expression; testis tissue shows the highest fraction of such genes. Further analyses incorporating mass spectrometry proteomics data establish that the co-evolution of transcriptomes and translatomes is reflected at the proteome layer. Together, our work uncovers co-evolutionary patterns and associated selective forces across the expression layers, and provides a resource for understanding their interplay in mammalian organs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw and processed Ribo-seq and RNA-seq data are available from ArrayExpress with accession number E-MTAB-7247. All other data are available as supplementary information or are available upon request. The interactive data resource we created, Ex2plorer, is publicly available at https://ex2plorer.kaessmannlab.org/.
Code availability
Custom R scripts used to generate the results reported in the manuscript and processed data are available at https://github.com/evgenyleushkin/translatome.
References
Necsulea, A. & Kaessmann, H. Evolutionary dynamics of coding and non-coding transcriptomes. Nat. Rev. Genet. 15, 734–748 (2014).
Vogel, C. & Marcotte, E. M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Genet. 13, 227–232 (2012).
Khan, Z. et al. Primate transcript and protein expression levels evolve under compensatory selection pressures. Science 342, 1100–1104 (2013).
Brar, G. A. & Weissman, J. S. Ribosome profiling reveals the what, when, where and how of protein synthesis. Nat. Rev. Mol. Cell Biol. 16, 651–664 (2015).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
McManus, J., May, G., Spealman, P. & Shteyman, A. Ribosome profiling reveals post-transcriptional buffering of divergent gene expression in yeast. Genome Res. 24, 422–430 (2014).
Artieri, C. G. & Fraser, H. B. Evolution at two levels of gene expression in yeast. Genome Res. 24, 411–421 (2014).
Stadler, M. & Fire, A. Conserved translatome remodeling in nematode species executing a shared developmental transition. PLoS Genet. 9, e1003739 (2013).
Albert, F. W., Muzzey, D., Weissman, J. S. & Kruglyak, L. Genetic influences on translation in yeast. PLoS Genet. 10, e1004692 (2014).
Wang, Z. et al. Evolution of gene regulation during transcription and translation. Genome Biol. Evol. 7, 1155–1167 (2015).
Wang, S. H., Hsiao, C. J., Khan, Z. & Pritchard, J. K. Post-translational buffering leads to convergent protein expression levels between primates. Genome Biol. 19, 83 (2018).
Hou, J. et al. Extensive allele-specific translational regulation in hybrid mice. Mol. Syst. Biol. 11, 825 (2015).
Csárdi, G., Franks, A., Choi, D. S., Airoldi, E. M. & Drummond, D. A. Accounting for experimental noise reveals that mRNA levels, amplified by post-transcriptional processes, largely determine steady-state protein levels in yeast. PLoS Genet. 11, e1005206 (2015).
Kleene, K. C. A possible meiotic function of the peculiar patterns of gene expression in mammalian spermatogenic cells. Mech. Dev. 106, 3–23 (2001).
Kleene, K. C. Patterns, mechanisms, and functions of translation regulation in mammalian spermatogenic cells. Cytogenet. Genome Res. 103, 217–224 (2003).
Iguchi, N., Tobias, J. W. & Hecht, N. B. Expression profiling reveals meiotic male germ cell mRNAs that are translationally up- and down-regulated. Proc. Natl Acad. Sci. USA 103, 7712–7717 (2006).
Brawand, D. et al. The evolution of gene expression levels in mammalian organs. Nature 478, 343–348 (2011).
Ramm, S. A., Schärer, L., Ehmcke, J. & Wistuba, J. Sperm competition and the evolution of spermatogenesis. Mol. Hum. Reprod. 20, 1169–1179 (2014).
Lareau, L. F., Inada, M., Green, R. E., Wengrod, J. C. & Brenner, S. E. Unproductive splicing of SR genes associated with highly conserved and ultraconserved DNA elements. Nature 446, 926–929 (2007).
Zarate, Y. A. & Fish, J. L. SATB2-associated syndrome: mechanisms, phenotype, and practical recommendations. Am. J. Med. Genet. A. 173, 327–337 (2017).
Sharma, K. et al. Cell type- and brain region-resolved mouse brain proteome. Nat. Neurosci. 18, 1819–1831 (2015).
Wang, D. et al. A deep proteome and transcriptome abundance atlas of 29 healthy human tissues. Mol. Syst. Biol. 15, e8503 (2019).
Bartha, I., di Iulio, J., Venter, J. C. & Telenti, A. Human gene essentiality. Nat. Rev. Genet. 19, 51–62 (2018).
Eisenberg, E. & Levanon, E. Y. Human housekeeping genes, revisited. Trends Genet. 29, 569–574 (2013).
Cardoso-Moreira, M. et al. Gene expression across mammalian organ development. Nature 571, 505–509 (2019).
Chen, W. H., Trachana, K., Lercher, M. J. & Bork, P. Younger genes are less likely to be essential than older genes, and duplicates are less likely to be essential than singletons of the same age. Mol. Biol. Evol. 29, 1703–1706 (2012).
Graves, J. A. Evolution of vertebrate sex chromosomes and dosage compensation. Nat. Rev. Genet. 17, 33–46 (2016).
Julien, P. et al. Mechanisms and evolutionary patterns of mammalian and avian dosage compensation. PLoS Biol. 10, e1001328 (2012).
Marin, R. et al. Convergent origination of a Drosophila-like dosage compensation mechanism in a reptile lineage. Genome Res. 27, 1974–1987 (2017).
Turner, J. M. Meiotic silencing in mammals. Annu. Rev. Genet. 49, 395–412 (2015).
Soumillon, M. et al. Cellular source and mechanisms of high transcriptome complexity in the mammalian testis. Cell Rep. 3, 2179–2190 (2013).
Faucillion, M. L. & Larsson, J. Increased expression of X-linked genes in mammals is associated with a higher stability of transcripts and an increased ribosome density. Genome Biol. Evol. 7, 1039–1052 (2015).
Chen, X. & Zhang, J. No X-chromosome dosage compensation in human proteomes. Mol. Biol. Evol. 32, 1456–1460 (2015).
Bader, D. M. et al. Negative feedback buffers effects of regulatory variants. Mol. Syst. Biol. 11, 785 (2015).
Schaefke, B., Sun, W., Li, Y. S., Fang, L. & Chen, W. The evolution of posttranscriptional regulation. Wiley Interdiscip. Rev. RNA 9, e1485 (2018).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Yates, A. et al. Ensembl 2016. Nucleic Acids Res. 44, D710–D716 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Tress, M. L., Abascal, F. & Valencia, A. Alternative splicing may not be the key to proteome complexity. Trends Biochem. Sci. 42, 98–110 (2017).
Gonzàlez-Porta, M., Frankish, A., Rung, J., Harrow, J. & Brazma, A. Transcriptome analysis of human tissues and cell lines reveals one dominant transcript per gene. Genome Biol. 14, R70 (2013).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Chan, P. P. & Lowe, T. M. GtRNAdb 2.0: an expanded database of transfer RNA genes identified in complete and draft genomes. Nucleic Acids Res. 44, D184–D189 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
R Core Team. R: a language and environment for statistical computing (R Foundation for Statistical Computing, 2012).
Yanai, I. et al. Genome-wide midrange transcription profiles reveal expression level relationships in human tissue specification. Bioinformatics 21, 650–659 (2005).
Futschik, M. E. & Carlisle, B. Noise-robust soft clustering of gene expression time-course data. J. Bioinform. Comput. Biol. 3, 965–988 (2005).
Alexa, A., Rahnenführer, J. & Lengauer, T. Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics 22, 1600–1607 (2006).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Bedford, T. & Hartl, D. L. Optimization of gene expression by natural selection. Proc. Natl Acad. Sci. USA 106, 1133–1138 (2009).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Shihab, H. A., Rogers, M. F., Campbell, C. & Gaunt, T. R. HIPred: an integrative approach to predicting haploinsufficient genes. Bioinformatics 33, 1751–1757 (2017).
Shao, Y. et al. GenTree, an integrated resource for analyzing the evolution and function of primate-specific coding genes. Genome Res. 29, 682–696 (2019).
Yang, Z. PAML: a program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 13, 555–556 (1997).
Acknowledgements
We thank K. Harshman and the Lausanne Genomics Technology Facility for high-throughput sequencing support; I. Xenarios and the Vital-IT computational facility for computational support; M. Baumann and S. Richling from the Heidelberg University Computational Center (Universitätsrechenzentrum, URZ) for supporting our computational work on the bwForCluster; M. Sanchez-Delgado for figure assistance; J. VandeBerg for opossum samples; P. Khaitovich for human and macaque samples; P. Jensen and A. Fallahshahroudi for red junglefowl samples; and A. B. Arpat and the Kaessmann group for discussions. This work was primarily supported by a grant (KA 1710/3-1) to H.K. from the German Research Council (DFG) and a European Research Council grant (615253, OntoTransEvol) to H.K. We acknowledge computational support by the state of Baden-Württemberg through bwHPC and the German Research Foundation (DFG) through grant INST 35/1134-1 FUGG. D.G. acknowledges funding through the National Centre of Competence in Research (NCCR) RNA & Disease; F.G. is supported by an Australian Research Council Fellowship; S.O. was supported by the DFG (CRC 1366); S.A. was supported by the DFG (SFB 1036); M.E.G. and A.H.F.M.P acknowledge funding from the Swiss National Science Foundation (31003A_146293) and the Novartis Research Foundation; and B.d.M. was funded by grants from the Centre National pour la Recherche Scientifique (CNRS) and the European Research Council (ERC) Executive Agency under the European Community’s Seventh Framework Programme (FP7/2007-2013 Grant Agreement no. 322788).
Author information
Authors and Affiliations
Contributions
H.K. conceived and designed the original study. The work was supervised by H.K. and E.L. Z.-Y.W. and E.L. processed all data and performed the analyses; A.L., K.M., T.B. and C.R. performed all experimental work. F.G. provided platypus samples and related biological expertise. M.C.-M. provided key analytical ideas and discussions. B.D., B.d.M., M.E.G. and A.H.F.M.P provided purified mouse spermatogenic cell samples. P.J. and D.G. helped to establish the original ribosome-profiling method for solid tissues. D.G. provided key experimental input and guidance during the data production and initial analysis phase. S.O. developed the Ex2plorer app. S.A. provided key statistical advice. Z.-Y.W., E.L. and H.K. wrote the manuscript, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Hunter Fraser, Bo Xia, Itai Yanai and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Information on generated RNA-seq and Ribo-seq data.
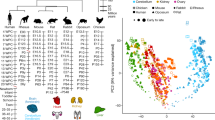
a, Ribosome footprint length distributions across Ribo-seq libraries (nt, nucleotides). b, Fractions of Ribo-seq and RNA-seq reads mapped to 5′-untranslated regions (5′-UTRs), coding sequences (CDSs) and 3′ untranslated regions (3′-UTRs), respectively. c, Distribution of Ribo-seq and RNA-seq reads across the three reading frames in the CDS of dominant splicing isoforms (frame 1: canonical reading frame). d, Mean normalized density of footprints along the coding region of the dominant isoforms of protein-coding genes for the brain Ribo-seq data. The Ribo-seq read (A-site) density for each position is plotted relative to the first nucleotide position of the start codon. e–h, Spearman’s correlation coefficient (ρ) of read counts for protein-coding genes with a mean read count > 1 between the two technical replicates for mouse liver Ribo-seq (e) and RNA-seq (f) data, and for chicken liver Ribo-seq (g) and RNA-seq (h) data. i, Correlations between biological replicates for Ribo-seq and RNA-seq data. Each dot corresponds to Spearman’s correlation coefficient (ρ) in pairs of biological replicates for every species-organ combination. Only one replicate (therefore no pairs) is available for the human liver transcriptome, only two replicates (one pair) are available for the human testis transcriptome and translatome, and only two replicates (one pair) are available for the platypus brain transcriptome. The correlation coefficients between the replicates are similar for the two data types and statistically indistinguishable (P = 0.159) in a Mann–Whitney U test (two-sided). j, Comparisons of gene-expression (rank) changes between the three expression layers. Changes in gene-expression ranks were calculated between expression layers (that is, from transcriptome to translatome, from transcriptome to proteome, and from translatome to proteome), and Spearman’s ρ was calculated to estimate the similarity of rank changes between the different pairs of expression layers. k–n, PCA based on 5,060 robustly expressed (median FPKM > 1 across organ libraries) 1:1 amniote orthologues. Factorial maps represent the relations of PC2 versus PC1 (k), PC3 versus PC1 (l) and PC4 versus PC1 (m). The scree plot (n) indicates the percentage of variance explained by each of the first 10 PCs. o, Expression variation at the two expression layers across mammalian organs for downsampled data. For this analysis data were downsampled to 2.5 million reads in each library. See Fig. 1d for the analysis of the full dataset. Organ and species icons are from a previous study25.
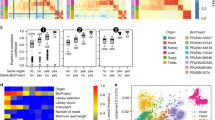
Extended Data Fig. 2 Correlations of gene-expression levels between sequenced libraries.
The heat map of the pairwise Spearman’s correlation coefficient (ρ) is based on the set of 5,060 robustly expressed (median FPKM > 1 across organ libraries) 1:1 amniote orthologues for perfectly aligned regions (see Methods). It represents the degree of similarity of gene-expression profiles between data types (translatome, transcriptome), species (human, macaque, mouse, opossum, platypus, chicken) and tissues (brain, liver, testis).
Extended Data Fig. 3 Quality assessment and analysis of mouse spermatogenesis data.
a, Ribosome footprint length distributions across Ribo-seq libraries (nt, nucleotides). b, Fractions of Ribo-seq and RNA-seq reads mapped to 5′-UTRs, CDSs and 3′-UTRs, respectively. c, Distribution of Ribo-seq and RNA-seq reads across the three reading frames in the CDS of dominant splicing isoforms (frame 1: canonical reading frame). d, PCA based on 11,057 genes robustly expressed (median FPKM > 1) across mouse spermatogenesis libraries. The scree plot (inset) indicates the percentage of variance explained by each of the first 10 PCs. e, Expression variation at the translatome layer calculated for simulated scenarios with different amounts of translational contribution (see Methods for details). Dashed line corresponds to IQR calculated at the transcriptome layer. f, g, Spearman’s ρ between transcription abundance and translational efficiency was calculated for 5,060 robustly expressed (median FPKM > 1 across organ libraries) 1:1 amniote orthologues in bulk testis across the amniotes (f) and across spermatogenesis stages in mouse (g). h, i, Translational efficiency (h) and translational shift (i) for clusters of genes (gene numbers in parentheses) with distinct translational efficiency patterns (Mfuzz clustering). Arrows indicate translational efficiency increases or decreases compared to the respective global pattern (Fig. 1e). *indicates a cluster of genes, which escape expression repression and delay at the translatome layer. j, Expression of individual genes, representing each of the five translational efficiency clusters, at the transcriptome and translatome layers (left column); shift in expression timing between expression layers for the corresponding genes (right column) with crosses representing the centres of mass of gene expression across spermatogenesis. k, Tissue-specificity (tissue Tau) across translational efficiency clusters. Cluster I, highlighted in colour, is dominated by testis-specific genes. Box plots represent the median ± 25th and 75th percentiles; whiskers are at 1.5 × IQR. l, Gene-expression divergence at the two expression layers for genes with stage-specific expression across spermatogenesis among 8,109 1:1 orthologues robustly expressed (FPKM > 1) in macaque, mouse and opossum.
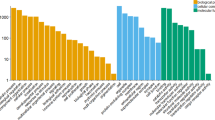
Extended Data Fig. 4 GO enrichment analyses.
a–d, Top five significantly enriched GO terms among genes with high (a, b) and low (c, d) translational efficiency for brain (a, c; blue) and liver (b, d; green) in mouse. e, Top ten significantly enriched GO terms for each of the mouse spermatogenesis translational efficiency trajectory cluster (Extended Data Fig. 3h–k). f–h, Significantly enriched GO terms (biological processes) among genes that changed significantly more at the translatome compared to the transcriptome layer in brain (f), liver (g) and testis (h). Significance was estimated in Fisher’s exact test (P < 0.05), with P values adjusted for multiple testing using the Benjamini–Hochberg method.
Extended Data Fig. 5 Normalization procedures in the evolutionary expression analyses.
a, Illustration of the normalization approach used in our study to globally assess gene-expression evolution. In this approach, evolutionary changes in gene expression are based on the assessment of expression differences across 1:1 orthologues between species. Specifically, we quantify the differences across orthologues as the variance (var) of their log2-transformed fold expression changes between species (left column), which is then divided (normalized) by the expression variation, calculated as the variance (var) of expression levels across genes, averaged across all studied species (right column). This procedure provides the expression divergence estimate (d). We note that the variance is similar across species for a given organ and expression layer (Fig. 1d). The example shown illustrates changes between human and each of the other five species in brain at the transcriptome layer. b, Illustration of the normalization procedure used to assess the expression evolution of individual genes. The normalization coefficient k is calculated as the ratio of the variances (var) across genes between the translatome and the transcriptome layer. The brain is shown as an example. Organ and species icons are from a previous study25.
Extended Data Fig. 6 Simulation of gene-expression divergence across expression layers.
a, Simulation of gene-expression divergence across different evolutionary scenarios. Expression divergence at the translatome layer between macaque and mouse brain was modelled over parameters of compensation and translational efficiency change (see ‘Modelling gene-expression divergence’ section in the Methods for details). Red (blue) corresponds to simulated scenarios with expression divergence higher (lower) than in actual data. Black line corresponds to simulated scenarios demonstrating expression divergence values observed in actual data. b, Contrast in evolutionary rates between the two expression layers for simulated data. Δ was calculated for simulated datasets with different amounts of compensation and different amounts of ϑ, corresponding to expression variation between individuals and measurement errors (see Methods for details).
Extended Data Fig. 7 Contrast in evolution between transcriptome and translatome layers for individual genes in downsampled data.
a–c, Δ was calculated on the basis of datasets downsampled to 0.5 million in each library for brain (a), liver (b) and testis (c). See Fig. 2e–f in the main text for the analysis of the full dataset.
Extended Data Fig. 8 Screenshot of the SATB2 gene in our Ex2plorer application.
SATB2 is an example of a gene that changes significantly less at the translational layer compared to the transcriptional layer in mammalian brain. Organ icons are from a previous study25.
Extended Data Fig. 9 Evolution at the proteome layer between human and mouse brain for genes with slower or faster evolution at the translatome compared to the transcriptome layer.
Absolute rank changes of proteome expression levels were calculated for genes with slower (olive) and faster (purple) evolution at the translatome compared to the transcriptome layer. The difference of the distributions between the two gene sets is statistically significant (****P < 0.0001, Mann–Whitney U test, two-sided). Box plots represent the median ± 25th and 75th percentiles; whiskers are at 1.5 × IQR.
Extended Data Fig. 10 Mammalian lineage-specific changes between expression layers.
a–c, Number of genes with lineage-specific patterns of slower (olive) or faster (purple) evolution at the translational layer, potentially driven by stabilizing and directional selection, respectively, for brain (a), liver (b) and testis (c). Owing to the lack of a biological replicate, the branch leading to human was omitted in the liver phylogeny for the transcriptional layer. d, e, Examples of individual genes with potential patterns of stabilizing (d) or directional (e) evolution. Species names with significant changes are marked by corresponding colours. Organ and species icons are from a previous study25.
Extended Data Fig. 11 Compensatory evolution of X-linked genes.
a, b, Examples of upregulation for the dosage reduction at the transcriptome layer. Species affected by upregulation are shown in olive, with arrows representing compensatory changes at the translatome layer. c, Median ratio of X-linked gene-expression values in mouse spermatogenic cell types to expression values of their 1:1 orthologues in chicken testis. In all cases log2 ratio at the translatome layer is significantly (P < 0.05, Mann–Whitney U test, two-sided) higher than at the transcriptome layer (marked in bold). Solid vertical lines correspond to expression levels expected under no dosage reduction (that is, log2 ratio = −1). d, Median present-day to ancestral gene-expression ratios at two expression layers for 1:1 orthologous autosomal genes located on chromosome 4 in chicken for brain, liver and testis. Chicken orthologues were used as a proxy for ancestral expression. See Fig. 4a and main text for details. e, Normalized translational efficiencies for 1:1 orthologues of eutherian X-linked and autosomal genes across amniote organs. Mann–Whitney U tests (two-sided) were performed for statistical comparisons (non-significant, ns: P > 0.05, ***P = 0.00003, ****P < 0.0001). P values were adjusted for multiple testing using Bonferroni method. Box plots represent the median ± 25th and 75th percentiles; whiskers are at 1.5 × IQR. Organ and species icons are from a previous study25.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-10.
Rights and permissions
About this article
Cite this article
Wang, ZY., Leushkin, E., Liechti, A. et al. Transcriptome and translatome co-evolution in mammals. Nature 588, 642–647 (2020). https://doi.org/10.1038/s41586-020-2899-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2899-z
This article is cited by
-
Simple but powerful interactive data analysis in R with R/LinekdCharts
Genome Biology (2024)
-
Evolution of chemosensory tissues and cells across ecologically diverse Drosophilids
Nature Communications (2024)
-
Analysis of the mechanism of curcumin against osteoarthritis using metabolomics and transcriptomics
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Evolution of tissue-specific expression of ancestral genes across vertebrates and insects
Nature Ecology & Evolution (2024)
-
Transcriptome and histological analyses on the uterus of freckle egg laying hens
BMC Genomics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.