Abstract
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) spike protein substitution D614G became dominant during the coronavirus disease 2019 (COVID-19) pandemic1,2. However, the effect of this variant on viral spread and vaccine efficacy remains to be defined. Here we engineered the spike D614G substitution in the USA-WA1/2020 SARS-CoV-2 strain, and found that it enhances viral replication in human lung epithelial cells and primary human airway tissues by increasing the infectivity and stability of virions. Hamsters infected with SARS-CoV-2 expressing spike(D614G) (G614 virus) produced higher infectious titres in nasal washes and the trachea, but not in the lungs, supporting clinical evidence showing that the mutation enhances viral loads in the upper respiratory tract of COVID-19 patients and may increase transmission. Sera from hamsters infected with D614 virus exhibit modestly higher neutralization titres against G614 virus than against D614 virus, suggesting that the mutation is unlikely to reduce the ability of vaccines in clinical trials to protect against COVID-19, and that therapeutic antibodies should be tested against the circulating G614 virus. Together with clinical findings, our work underscores the importance of this variant in viral spread and its implications for vaccine efficacy and antibody therapy.
Similar content being viewed by others
Main
Since the identification of SARS-CoV-2 in China in late 20193, COVID-19 has caused more than 47 million confirmed infections and more than 1.2 million deaths worldwide. Although most infections are mild, SARS-CoV-2 can cause severe, life-threatening pneumonia, particularly in older age groups and those with chronic conditions. The exact pathogenesis of severe COVID-19 remains unclear, but it typically involves a dysregulated, hyperinflammatory response following viral infection4. However, in addition to the host response, variations in the viral strain could contribute to disease severity and efficiency of spread.
Coronaviruses have evolved a genetic proofreading mechanism to maintain their long RNA genomes5. Despite the low sequence diversity of SARS-CoV-26, mutations in the spike protein, which interacts with cellular receptors such as angiotensin-converting enzyme 2 (ACE2) to mediate entry into cells, have a strong influence on host range, tissue tropism and pathogenesis. During the SARS-CoV outbreak in 2002–2003, one such mutation mediated adaptation for infection of the intermediate civet host as well as interhuman transmission7. For SARS-CoV-2, analyses of more than 28,000 spike gene sequences in May 2020 revealed a D614G substitution that was rare before March 2020, but became more common as the pandemic spread1, occurring in over 74% of all published sequences by June 20202. The D614G substitution was accompanied by three other mutations: a C-to-T mutation in the 5′ untranslated region at position 241, a synonymous C-to-T mutation at position 3037, and a nonsynonymous C-to-T mutation at position 14408 in the RNA-dependent RNA polymerase gene8. The frequency of this set of mutations increased not only globally, but also during co-circulation within individual regions during outbreaks, suggesting that the increase was the result of a fitness advantage rather than founder effects and/or genetic drift. The association of spike amino acid substitutions with coronavirus transmissibility suggested that the D614G substitution was critical to this putative selective sweep. The correlation of this mutation with higher nasopharyngeal viral RNA loads in patients with COVID-191,9 also supported a putative advantage of the mutant in transmission. However, direct measurements of fitness are needed to confirm this hypothesis.
Initial phenotypic characterizations of the spike D614G substitution were performed using pseudotyped viruses, in which replication kinetics were studied in vesicular stomatitis virus and lentiviral particles incorporating the SARS-CoV-2 spike protein. The D614G substitution resulted in significantly higher pseudovirus titres in multiple cell types, suggesting that spike G614 might be associated with enhanced entry into cells and enhanced replication in airways1,2. However, these results need to be confirmed in studies with authentic G614 SARS-CoV-2, as well as using in vivo studies with a suitable animal model. Therefore, using an infectious cDNA clone for SARS-CoV-210, we generated the D614G substitution in the USA-WA1/2020 strain11 and performed experiments in cell culture, primary human airway tissue and a hamster infection model12. We also developed D614 and G614 mNeonGreenSARS-CoV-2 viruses for rapid testing of neutralization by serum specimens and monoclonal antibodies. Our study has major implications for understanding the evolution and transmission of SARS-CoV-2 as well as the development of COVID-19 vaccines and therapeutic antibodies.
D614G effect on viral replication and infectivity
We first examined the effect of the spike D614G substitution on viral replication in cell cultures. Site-directed mutagenesis was performed on an infectious cDNA clone of SARS-CoV-2 to prepare a pair of recombinant isogenic viruses with spike D614 or G614 (Fig. 1a). Similar amounts of infectious D614 and G614 viruses were recovered from Vero E6 cells (monkey kidney epithelial cells). The two viruses formed similar plaque morphologies (Fig. 1b). In Vero E6 cells, the G614 virus replicated with a higher infectious titre than D614 at 12 h post-infection (hpi), after which the two viruses replicated at comparable levels (Fig. 1c). A similar trend was observed for extracellular viral RNA production from the infected Vero E6 cells (Fig. 1d). Sequencing of the spike gene of viruses purified from Vero E6 cells did not reveal any further mutations. To compare infectivity, we calculated the genomic RNA/plaque-forming units (PFU) ratio; we found no significant differences between the two viruses (Fig. 1e), indicating that the D614G mutation does not affect viral replication or virion infectivity in Vero E6 cells.
a, Construction of G614 SARS-CoV-2. A single-nucleotide A-to-G substitution was introduced to produce the spike D614G substitution in the infectious cDNA clone of SARS-CoV-2. Numbers on the main schematic refer to open reading frames (ORFs). E, envelope glycoprotein gene; L, leader sequence; M, membrane glycoprotein gene; N, nucleocapsid gene; UTR, untranslated region. b, Plaque morphologies of D614 and G614 viruses, developed on day 2 after infection in Vero E6 cells. c–h, Viral replication and genomic RNA/PFU ratios of D614 and G614 viruses produced from Vero E6 cells (c–e) and from Calu-3 cells (f–h). Cells were infected at a MOI of 0.01. Infectious viral titres (c, f) and genomic RNA levels (d, g) in the culture medium were determined by plaque assay and quantitative RT–PCR (RT–qPCR), respectively. The genomic RNA/PFU ratios (e, h) were calculated to indicate virion infectivity. Dots represent individual samples, bar heights represent means and error bars show s.d. P values were determined by two-tailed Mann–Whitney test from a sample size of n = 6 (two independent experiments performed in triplicate). All P values below 0.05 are shown. i, j, Spike protein cleavage of purified virions from Calu-3 cells (i) and Vero E6 cells (j). Purified D614 and G614 virions were analysed by western blot using polyclonal antibodies against spike and nucleocapsid. Full-length spike20, S1/S2-cleavage form and S2′ protein are indicated. Results from two independent experiments are presented.
Next, we compared the replication kinetics of D614 and G614 viruses in the human lung epithelial cell line Calu-3. With a multiplicity of infection (MOI) of 0.01, the G614 virus produced modest 1.2-, 2.4- and 1.9-fold more infectious virus than the D614 virus at 24, 36 and 48 hpi, respectively (Fig. 1f), indicating that D614G enhances viral replication. By contrast, the cells infected with G614 virus produced less (at 24 and 36 hpi) or equivalent (at 48 hpi) extracellular viral RNA compared with cells infected with D614 virus (Fig. 1g). The genomic RNA/PFU ratios of D614 virus were therefore 1.9- to 3.0-fold higher than those of G614 virus (Fig. 1h), indicating that the D614G mutation increases the infectivity of SARS-CoV-2 produced from a human lung cell line.
To explore the mechanism of increased infectivity of G614 virus produced from Calu-3 cells, we compared the processing of the spike protein from D614 and G614 viruses. We purified virions from the culture medium of infected Calu-3 cells using ultracentrifugation and a sucrose cushion. The pelleted viruses were analysed for spike protein processing by western blot, using nucleocapsid protein as a loading control. In both viruses, full-length spike was almost completely processed to the S1/S2 cleaved form and S2′, with similar cleavage efficiencies of 93% for D614 and 95% for G614 (Fig. 1i). When virions produced from Vero E6 cells were analysed, less full-length spike protein was processed to the S1/S2 form, with cleavage efficiencies of 73% for D614 and 67% for G614 (Fig. 1j; uncropped images in Supplementary Fig. 1). These results suggest that more spike protein is cleaved to the S1/S2 form in virions produced from Calu-3 cells compared with Vero E6 cells, and that the D614G substitution does not significantly affect the spike cleavage ratio.
Increased fitness in the hamster upper airway
The in vivo relevance of the D614G mutation was evaluated in the Syrian golden hamster model (Extended Data Fig. 1a). Four- to five-week-old hamsters were infected intranasally with 2 × 104 PFU of D614 or G614 virus. Infected hamsters from both groups exhibited similar weight loss (Fig. 2a). On day 2 after infection, infectious viral titres from nasal washes, trachea and various lobes of the lung were consistently higher in the hamsters infected with G614 virus compared with those infected with D614 virus, particularly in the upper airway, although the differences did not reach statistical significance (Fig. 2b, Extended Data Fig. 1b). By day 4 after infection the differences in infectious viral titres became more pronounced in the upper airway, with 1.3 log10 PFU ml−1 and 0.9 log10 PFU g−1 higher titres of G614 virus than D614 virus in the nasal washes and tracheae, respectively (Fig. 2c), whereas the viral loads in the lungs were essentially identical. No infectious virus was detected on day 7 after infection (data not shown).
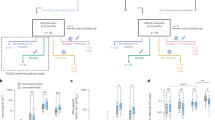
a–i,Three- to four-week-old male Syrian golden hamsters were infected intranasally with 2 × 104 PFU of D614 or G614 SARS-CoV-2 or PBS (mock). All data are from a single experiment. a, Weight loss was monitored for seven days after infection. Data are mean ± s.d.; n = 18 (D614 and G614) and n = 14 (mock) at days 0–2; n = 12 (D614 and G614) and n = 10 (mock) at days 3 and 4; n = 6 (all cohorts) at days 5–7. Weight loss was analysed by two-factor analysis of variance (ANOVA) with Tukey’s post hoc test. P > 0.05 for D614 vs G614 at all time points. b–d, Infectious titres (b, c) and amount of viral genomes (d) were measured in the nasal wash, trachea and lung on days 2 (b, d), 4 (c, d) and 7 (d) after infection. e, f, Genome/PFU ratios on days 2 (e) and 4 (f) after infection were calculated as a measure of infectivity. In b–f, symbols represent individual hamsters (n = 6). Centre lines (b, c, e, f) and bar heights (d) represent means, and error bars represent s.d. Two-factor ANOVA with Sidak’s post hoc test. g–i, Hamsters were inoculated with 1:1 mixtures of D614 and G614 viruses (104 PFU each). Nasal wash, trachea and lungs were collected on days 2 (g), 4 (h) and 7 (i) after infection. Relative amounts of D614 and G614 RNAs were assessed by RT–PCR and Sanger sequencing. In g–i, dots represent individual hamsters (n = 6), the centre line represents the mean, shaded regions represent s.e.m., the width represents the distribution of the model-adjusted means, and heights extend to 99.8% of the distribution; y-axes use a log10 scale. P values are calculated for the group (strain) coefficient for each linear regression model.
We compared the infectivity of the D614 and G614 viruses produced in hamsters by determining their viral RNA levels and viral RNA/PFU ratios. The two viruses produced nearly identical levels of viral RNA across all organs and time points (Fig. 2d). The RNA/PFU ratios of G614 virus were 0.3 log10 to 0.7 log10 lower than those of D614 virus across airway tissues on day 2 after infection (Fig. 2e). On day 4 after infection, the RNA/PFU ratio of G614 was 1.1 log10 lower than that of D614 in nasal wash, whereas the differences in the trachea and lungs were negligible (Fig. 2f). On day 7 after infection, despite there being no detectable infectious virus (detection limit 40 PFU ml−1), more than 108 viral genomes per ml were detected in the nasal washes (Fig. 2d), demonstrating high levels of viral RNA persistence after the clearance of infectious virus. This result recapitulates findings in human patients with COVID-19, who frequently test positive by PCR with reverse transcription (RT–PCR) for several weeks but have low or undetectable infectious virus. One caveat of the above calculations is that the total RNA could include viral RNAs from both virions and intracellular subgenomic RNAs during sample processing.
The above results prompted us to directly compare the finesses of D614 and G614 viruses in a competition experiment. This approach has major advantages over infecting replicate hosts with individual strains in that each competition is internally controlled, eliminating host-to-host variation that can reduce the power of experiments. Also, the ratios of infective strains can be assayed with more precision than individual virus titres. Thus, competition assays have been used for many studies of microbial fitness, including viruses13,14,15,16. We infected hamsters intranasally with equal amounts of the two viruses (104 PFU per virus), collected nasal wash and respiratory organs on days 2, 4 and 7 after infection, and quantified the relative amounts of D614 and G614 RNAs by RT–PCR and Sanger sequencing. This method was validated to reliably quantify the relative amounts of D614 and G614 viruses in a mixed specimen (Extended Data Fig. 2). A G614/D614 ratio greater than one indicates a replication advantage for G614. All respiratory tissues showed G614/D614 ratios of 1.2 to 2.6 on days 2, 4 and 7 after infection, indicating that G614 virus has a consistent advantage over D614 virus (Fig. 2g–i).
Enhanced replication in a human airway model
To further define the effect of the spike D614G mutation in the human respiratory tract, we characterized the replication of D614 and G614 viruses in a primary human airway tissue culture model (Fig. 3a). This model contains human tracheal and bronchial epithelial cells in layers resembling the epithelial tissue of the respiratory tract. The primary tissue is cultured at an air–liquid interface to recapitulate the barrier, microciliary response and infection of human airway tissues in vivo17,18. After infecting the airway tissue at a MOI of 5 (as determined in Vero E6 cells), both D614 and G614 viruses produced increasing infectious titres from day 1 to 5, up to 7.8 × 105 PFU ml−1 (Fig. 3b), demonstrating that the airway tissue supports SARS-CoV-2 replication. The infectious viral titres of G614 virus were significantly higher (2.1- to 8.6-fold) than those of D614 virus (Fig. 3b). By contrast, there were no differences in viral RNA yields between the variants (Fig. 3c). The genomic RNA/PFU ratios of D614 virus were 1.4- to 5.3-fold higher than those of G614 virus (Fig. 3d). Sequencing of the viruses collected on day 5 after infection did not show any acquired mutations. Collectively, these results demonstrate that the spike D614G mutation enhances viral replication through increased virion infectivity in primary human upper airway tissues.
a, Experimental scheme. D614 and G614 viruses were inoculated onto primary human airway tissues at a MOI of 5. After incubation for 2 h, the culture was washed with Dulbecco’s PBS (DPBS), then maintained for 5 days. For collection, DPBS was added at 37 °C for 30 min to elute the virus. b–d, Viral replication and genomic RNA/PFU ratios. The amounts of infectious virus (b) and genomic RNA (c) were quantified by plaque assay and RT–qPCR, respectively. d, The genomic RNA/PFU ratio was calculated as an indication of virion infectivity. In b–d, symbols represent individual samples, bar heights represent means, and error bars represent standard deviations. P values were determined by two-tailed Mann–Whitney test (n = 6, two independent experiments conducted in triplicate). e–g, Competition assay. A mixture of D614 and G614 viruses with initial ratios of 1:1 (e), 3:1 (f) or 9:1 (g) were inoculated onto human airway tissue cultures at a total MOI of 5. Virus ratios after competition were measured by Sanger sequencing. Dots represent individual samples (n = 6, two independent experiments conducted in triplicate); the centre line represents the sample mean; the shaded region represents s.e.m.; the width represents the distribution of the model-adjusted means; and the heights extend to 99.8% of the distribution of the mean; the y-axes use a log10 scale such that the null value is 1. P values are calculated for the group (strain) coefficient for each linear regression model.
Next, we performed competition experiments in the human airway tissue culture model. After infecting the cells with a 1:1 infectious ratio of D614 and G614 viruses, the G614/D614 ratio increased from 1.2 at 1 day post-infection (dpi) to 13.9 at 5 dpi (Fig. 3e). In addition, after infecting the airway culture with 3:1 ratio of D614 and G614 viruses, the G614 virus rapidly overcame its initial deficit to reach a slight advantage with a G614/D614 ratio of 1.2 by 1 dpi, with that advantage increasing to 9.1 by 5 dpi (Fig. 3f). Furthermore, when infecting the airway tissue with a 9:1 ratio of D614 to G614, the G614/D614 ratios increased from1.4 to 5.2 from 1 dpi to 5 dpi (Fig. 3g). Similar results were obtained when the experiment was repeated using a different donor-derived human airway culture (Extended Data Fig. 3). These results confirm that the G614 virus can rapidly outcompete the D614 virus when infecting human airway tissues, even when it is initially present as a minor variant in a mixed population.
Stability of D614 and G614 viruses
To examine the effect of D614G on virus stability, we measured the decay of infectivity of D614 and G614 viruses over time at 33 °C, 37 °C and 42 °C (Extended Data Fig. 4). The infectivity of both viruses decreased at a faster rate at 42 °C than at 37 °C or 33 °C. However, the G614 virus retained higher infectivity than the D614 virus at all temperatures, suggesting that the D614G mutation may increase the stability of SARS-CoV-2.
Susceptibility to neutralization
All of the COVID-19 vaccines currently in clinical trials are based on the original D614 spike sequence19,20. We therefore investigated whether the D614G substitution could reduce the efficacy of these vaccines, assuming that the G614 virus continues to circulate. To address this question, we measured the neutralization titres of a panel of sera collected from hamsters that were previously infected with D614 SARS-CoV-2 (Extended Data Fig. 5). Each serum was analysed using mNeonGreen reporter D614 or G614 SARS-CoV-2 viruses21 (Extended Data Fig. 6). The mNeonGreen gene was engineered in open reading frame 7 of the SARS-CoV-2 genome10. All sera exhibited 1.4- to 2.3-fold higher neutralization titres (mean 1.7-fold) against heterologous G614 virus compared with homologous D614 virus (Figs. 4a–c, Extended Data Fig. 7), suggesting that the D614G mutation may confer higher susceptibility to serum neutralization.
a, Neutralizing activities of hamster sera against D614 and G614 mNeonGreen reporter SARS-CoV-2. Sera from hamsters infected with D614 virus (n = 8) were tested for neutralizing titres against D614- and G614-reporter SARS-CoV-2, and the 1/NT50 values are plotted. Symbols represent sera from individual hamsters. b, Ratio of 1/NT50 between D614 and G614 viruses. Symbols represent sera from individual hamsters (n = 8), the centre line represents the mean, and error bars represent s.d. c, Representative neutralizing curve of serum from an individual hamster (serum 5). Symbols represent individual replicates, the solid line represents the fitted curve, and the dotted line indicates 50% viral inhibition. d, Neutralizing activities of 11 human monoclonal antibodies against D614 and G614 mNeonGreen SARS-CoV-2. Symbols represent individual monoclonal antibodies. Data represent one of two independent experiments. e, Ratio of NT50 between D614 and G614 viruses. The averages of the NT50 ratios from two independent experiments performed in duplicate are shown. Dots represent individual monoclonal antibodies, the centre line represents the mean, and the error bars represent the s.d. f, Representative neutralizing curve of mAb18. Symbols represent individual replicates, the solid line represents the fitted curve, and the dotted line indicates 50% viral inhibition.
Next, we evaluated the activity of a panel of 11 monoclonal antibodies to the SARS-CoV-2 receptor-binding domain (RBD) against the D614 and G614 mNeonGreen SARS-CoV-2 viruses. The details of these antibodies will be reported elsewhere (Z. An et al., submitted). One of the antibodies (mAb18) showed a 2.1-fold higher neutralizing activity against G614 than against D614 virus, whereas the others showed similar neutralization activities against both viruses (Fig. 4d–f, Extended Data Figs. 8, 9). The results suggest that the D614G mutation may modulate spike protein conformation, thereby affecting neutralization by monoclonal antibodies in an epitope-specific manner.
Discussion
We have shown that the spike D614G substitution enhances SARS-CoV-2 replication in the upper respiratory tract through increased virion infectivity. Compared with the original D614 virus, the emergent G614 virus exhibited increased viral replication in the human lung cell line Calu-3 and in primary human upper airway tissues. The differences in replication were higher in the primary human airway culture, with an advantage of up to 13.9-fold in a competition assay. The increased replication fitness was accompanied by increases in specific infectivity and stability of the G614 virions. Previous studies with pseudotyped virus have shown that the efficiency of spike protein cleavage into S1 and S2 fragments modulates SARS-CoV-2 infection22,23; we therefore compared the spike cleavage with D614 and G614 virions. Although virions produced from Calu-3 cells showed more complete S1/S2 cleavage than those produced form Vero E6 cells, no substantial differences in spike cleavage were detectable between the D614 and G614 virions, suggesting that the enhanced virion infectivity is unlikely to be caused by differences in spike cleavage. In contrast with previous studies reporting that the D614G substitution changes the cleavage and shedding of spike protein when expressed alone on pseudotyped virions24,25, we found no such difference in experiments with SARS-CoV-2. Mechanistically, recent studies have shown that the D614G substitution abolishes a hydrogen-bond interaction with T859 from a neighbouring protomer of the spike trimer1, which allosterically shifts the RBD to an ‘up’ conformation, promoting binding with the ACE2 receptor2, leading to enhanced virion infectivity.
The higher viral loads of G614 in the upper airway of patients with COVID-1926 and infected hamsters suggest a role of the D614G substitution in viral transmissibility. The robust replication of SARS-CoV-2 in the human upper airway may be partially conferred by higher expression of the ACE2 receptor in the nasal cavity compared with the lower respiratory tract8,27. Compared with the D614 virus, the G614 virus showed increased replication in the upper airway—but not in the lungs—of hamsters, suggesting that the D614G substitution may select against replication in the lung. Patients infected with G614 virus developed higher levels of viral RNA in nasopharyngeal swabs than those with D614 virus, but did not develop more severe disease1,2,9. Our hamster model of infection recapitulated these clinical findings: the G614 virus developed higher infectious titres than the D614 virus in nasal washes and tracheas, but not in lungs, and there were no differences in weight loss or disease symptoms. If the lower viral RNA/PFU ratio for the G614 virus observed in our hamster and human-airway models could be extrapolated to patients with COVID-19, the modest differences in cycle threshold28 values observed in RT–PCR tests of patients’ nasopharyngeal swabs would translate to more than tenfold higher infectivity for the G614 virus, underscoring the potential for enhanced transmission and spread. Indeed, during the review process for this Article, a non-peer-reviewed manuscript reported that the G614 mutation increased SARS-CoV-2 transmission in hamsters29. This potential is further bolstered by the observation that a patient with COVID-19 who had distinct populations of SARS-CoV-2 in throat swabs and sputum samples transmitted only the throat strain26. Similar nasal-driven transmission was recently reported for human influenza A virus in ferrets30. However, the differential effects of age on G614 SARS-CoV-2 infection, pathogenesis and transmission remain to be studied.
G614 virus was slightly more susceptible to neutralization by sera collected from hamsters infected with D614 virus. This seems counterintuitive, but could be explained by the increase in the ‘up’ conformation of the RBD1,2 in the G614 virus, exposing different epitopes for antibody binding. As current COVID-19 vaccines in clinical trials are based on the original D614 sequence, the results of our neutralization assays mitigate the concern that the D614G mutation might compromise their efficacy. Further studies using human sera collected following inoculation with D614 spike vaccines are needed to eliminate this possibility. We also showed that the neutralizing potency of some monoclonal antibodies could be affected by the D614G mutation. These results underscore the importance of testing therapeutic monoclonal antibodies against G614 virus.
In summary, we have used authentic SARS-CoV-2 to demonstrate that the spike D614G substitution enhances viral replication in the upper respiratory tract and increases the susceptibility of the virus to neutralization by antibodies. These findings have important implications for our understanding of the evolution and spread of the ongoing COVID-19 pandemic, and for vaccine efficacy and therapeutic antibody development. Further studies are needed on other emerging mutations in SARS-CoV-2, including those associated with the D614G substitution.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Ethics statement
Hamster studies were performed in accordance with the guidance for the Care and Use of Laboratory Animals of the University of Texas Medical Branch (UTMB). The protocol was approved by the Institutional Animal Care and Use Committee (IACUC) at UTMB. All the hamster operations were performed under anaesthesia by isoflurane to minimize animal suffering.
Animals and Cells
The Syrian hamsters (HsdHan:AURA strain) were purchased from Envigo. African green monkey kidney epithelial Vero E6 cells (provided by R. Baric) were grown in Dulbecco’s modified Eagle’s medium (DMEM) with 10% fetal bovine serum (FBS; HyClone Laboratories) and 1% antibiotic/streptomycin (Gibco). Human lung adenocarcinoma epithelial Calu-3 cells (ATCC) were maintained in a high-glucose DMEM supplemented with 10% FBS and 1% penicillin–streptomycin at 37 °C with 5% CO2. The EpiAirway system is a primary human airway 3D tissue model purchased from MatTek Life Science. This EpiAirway system was maintained with the provided culture medium at 37 °C with 5% CO2following the manufacturer’s instruction. All other culture media and supplements were purchased from ThermoFisher Scientific. All cell lines were verified and tested negative for mycoplasma. No further authentication of cell lines was performed.
Generation of SARS-CoV-2 spike D614G mutant viruses
One single-nucleotide substitution was introduced into a subclone puc57-CoV-2-F5-7 containing the spike gene of the SARS-CoV-2 wild-type infectious clone10 to convert amino acid 614 from aspartic acid to glycine by overlap fusion PCR. The full-length infectious cDNA clone of SARS-CoV-2 D614G was assembled by in vitro ligation of seven contiguous cDNA fragments following the previously described protocol10. For construction of D614G mNeonGreen SARS-CoV-2, seven SARS-CoV-2 genome fragments (F1 to F5, F6 containing D614G mutation, and F7-mNG containing the mNeonGreen reporter gene) were prepared and ligated in vitro as previously described10. In vitro transcription was then preformed to synthesize full-length genomic RNA. To recover the mutant viruses, the RNA transcripts were electroporated into Vero E6 cells. The viruses from electroporated cells were collected at 40 h after electroporation and served as seed stocks for subsequent experiments. The D614G mutation from the recovered viruses was confirmed by sequence analysis. Viral titres were determined by plaque assay on Vero E6 cells. All virus preparation and experiments were performed in a biosafety level 3 (BSL-3) facilities. Viruses and plasmids are available from the World Reference Center for Emerging Viruses and Arboviruses at the University of Texas Medical Branch.
RNA extraction, RT–PCR, and Sanger sequencing
Cell culture supernatants or clarified tissue homogenates were mixed with a fivefold excess of TRIzol LS Reagent (Thermo Fisher Scientific). Viral RNAs were extracted according to the manufacturer’s instructions. The extracted RNAs were dissolved in 20 μl nuclease-free water. Two microlitres of RNA samples were used for reverse transcription by using the SuperScript IV First-Strand Synthesis System (ThermoFisher Scientific) with random hexamer primers. Nine DNA fragments flanking the entire viral genome were amplified by PCR. The resulting DNAs were cleaned up by the QIAquick PCR Purification Kit, and the genome sequences were determined by Sanger sequencing at Genewiz.
The quantify viral RNA samples, quantitative RT–qPCR assays were performed using the iTaq SYBR Green One-Step Kit (Bio-Rad) on the LightCycler 480 system following the manufacturers ’protocols. Primers are listed in Extended Data Table 1. The absolute quantification of viral RNA was determined by a standard curve method using an RNA standard (in vitro transcribed 3,839-bp RNA containing genomic nucleotide positions 26044 to 29883 of SARS-CoV-2 genome).
To quantify D614:G614 ratios for competition assays, a 663-bp RT–PCR product was amplified from extracted RNA using a SuperScript III One-Step RT–PCR kit (Invitrogen). A 20-μl reaction was assembled in PCR 8-tube strips through the addition of 10 μl 2 × reaction mix, 0.4 μl SuperScript III RT/Platinum Taq Mix, 0.8 μl Forward Primer (10 μM) (Extended Data Table 1), 0.8 μl reverse primer (10 μM) (Extended Data Table 1), 4 μl RNA, and 6 μlRnase-free water. Reverse transcription and amplification was completed using the following protocol: (i) 55 °C, 30 min; 94 °C, 2 min; (ii) 94 °C, 15 s; 60 °C, 30 s; 68 °C, 1 min; 40 cycles; (iii) 68 °C, 5 min; (iv) indefinite hold at 4 °C. The presence and size of the desired amplicon was verified with 2 μl of PCR product on an agarose gel. The remaining 18 μl were purified by a QIAquick PCR Purification kit (Qiagen) according to the manufacturer’s protocol.
Sequences of the purified RT–PCR products were generated using a BigDye Terminator v.3.1 cycle sequencing kit (Applied Biosystems). The sequencing reactions were purified using a 96-well plate format (EdgeBio) and analysed on a 3500 Genetic Analyzer (Applied Biosystems). The peak electropherogram height representing each mutation site and the proportion of each competitor was analysed using the QSVanalyser program31.
Plaque assay
Approximately 1.2 × 106 Vero E6 cells were seeded to each well of 6-well plates and cultured at 37 °C, 5% CO2 for 16 h. Virus was serially diluted in either DMEM with 2% FBS (for viral stocks and in vitro-generated samples) or DPBS (for hamster tissues) and 200 μl was transferred to the monolayers. The viruses were incubated with the cells at 37 °C with 5% CO2 for 1 h. After the incubation, overlay medium was added to the infected cells per well. The overlay medium contained either DMEM with 2% FBS and 1% sea-plaque agarose (Lonza) in the case of in vitro samples or Opti-MEM with 2% FBS, 1% penicillin–streptomycin and 0.8% agarose in the case of in vivo samples. After a 2-day incubation, plates were stained with neutral red (Sigma-Aldrich) and plaques were counted on a light box.
Viral infection on cells
Approximately 3 × 105 Vero E6 or Calu-3 cells were seeded onto each well of 12-well plates and cultured at 37 °C, 5% CO2 for 16 h. Either SARS-CoV-2 D614 or G614 virus was inoculated into the cells at a MOI of 0.01. The virus was incubated with the cells at 37 °C for 2 h. After the infection, the cells were washed with DPBS 3 times to remove the unattached virus. One millilitre of culture medium was added into each well to maintain the cells. At each time point, 100 μl of culture supernatants were collected for the RT–qPCR detection and plaque assay. Meanwhile, 100 μl fresh medium was added into each well to replenish the culture volume. The cells were infected in triplicate for each virus. All samples were stored at −80 °C until plaque or RT–qPCR analysis.
Virion purification and spike protein cleavage analysis
Vero E6 or Calu-3 2B4 cells were infected with D614 or G614 viruses at a MOI of 0.01. At 24 (for Vero) or 48 (Calu-3) hours after infection, the culture media were collected and clarified by low speed spin. Virions in the medium were pelleted by ultracentrifugation through a 20% sucrose cushion at 26,000 rpm for 3 h at 4 °C in a BeckmanSW28 rotor. The purified virions were analysed by western blot using polyclonal antibodies against spike protein and nucleocapsid as previously described32.
Viral infection in a primary human airway tissue model
The EpiAirway system is a primary human airway 3D mucociliary tissue model consisting of normal, human-derived tracheal/bronchial epithelial cells. For viral replication kinetics, either D614 or G614 virus was inoculated onto the culture at a MOI of 5 in DPBS. After 2 h infection at 37 °C with 5% CO2, the inoculum was removed, and the culture was washed three times with DPBS. The infected epithelial cells were maintained without any medium in the apical well, and medium was provided to the culture through the basal well. The infected cells were incubated at 37 °C, 5% CO2. From day 1 to day 5, 300 μl DPBS was added onto the apical side of the airway culture and incubated at 37 °C for 30 min to elute the released viruses. All virus samples in DPBS were stored at −80 °C.
Hamster infection
Four- to five-week-old male Syrian golden hamsters, strain HsdHan:AURA (Envigo), were inoculated intranasally with 2 × 104 PFU SARS-CoV-2 in a 100 μl volume. Eighteen hamsters received wild-type D614 virus, 18 received mutant G614 virus, and 18 received a mixture containing 104 PFU of D614virus and 104 PFU of G614 virus. The infected hamsters were weighed and monitored for signs of illness daily. On days 2, 4 and 7 after infection, cohorts of 6 infected hamsters and 4 (days 2 and 4) or 6 (day 7) mock-infected hamsters were anaesthetized with isoflurane and nasal washes were collected in 400 μl sterile DPBS. Hamsters were humanely euthanized immediately following the nasal wash. The trachea and the four lobes of the right lung were collected in maintenance medium (DMEM supplemented with 2% FBS and 1% penicillin–streptomycin) and stored at −80 °C. Samples were subsequently thawed, tissues consisting of individual, intact lung lobes or the trachea were homogenized for 1 min at 26 strokes per s, and debris was pelleted by centrifugation for 5 min at 16,100g. Infectious titres were determined by plaque assay. Genomic RNAs were quantified by quantitative RT–PCR (Extended Data Table 1). Ratios of D614/G614 RNA were determined via RT–PCR with quantification of Sanger peak heights.
Competition assay
For the competition in primary human airway 3D tissue model, the D614 and G614 mutant viruses were mixed and inoculated onto the cells at a final MOI of 5. The initial ratio of D614 and G614 viruses was 1:1, 3:1 or 9:1 based on PFU titres determined on Vero E6 cells. The DPBS with viruses was collected every day from day 1 to 5 following the protocol described above. For the competition in hamsters, 100-μl mixtures of D614 and G614 viruses (total 2 × 104 PFU per hamster) were inoculated intranasally into 4–5-week-old Syrian hamsters. On days 2, 4, and 7 pi, 6 infected hamsters were sampled for competition detection. An aliquot of the inoculum for both hamster and human airway infections was backtitred to estimate the initial ratio of viruses. All samples were stored at −80 °C before analysis.
Validation of competition assay by Sanger sequencing
To validate the consistency and accuracy of competition assay by Sanger sequencing, the D614 and G614 viruses were mixed at ratios of 10:1, 5:1, 3:1, 1:1, 1:3, 1:5 and 1:10 based on their PFU titres (total 106 PFU viruses) or mixed with 106, 105, 104, 103 and 102 PFU of the two viruses at a ratio of 1:1. The total RNA of these mixed viruses was isolated and amplified by RT–PCR. The ratios of D614/G614 were calculated by the peak heights of Sanger sequencing. Data were analysed by linear regression with correlation coefficients (r) and significance (P).
Stability assay of SARS-CoV-2
D614 and G614 viruses were diluted to a final titre of 105 PFU ml−1 in DPBS. The diluted viruses were incubated at 42 °C, 37 °C or 33 °C. At indicated time points, 200 μl DPBS containing the viruses were taken out and stored at −80 °C. All the samples were then analysed by plaque assay on Vero E6 cells. The percentages of remaining infectious viruses were calculated by dividing the viral titres at each time point by the mean viral titre at 0 h.
Neutralization assay
Neutralization assays were preformed using D614 and G614 mNeonGreen SARS-CoV-2 as previously described21. In brief, Vero (CCL-81) cells were plated in black CLEAR flat-bottom 96-well plate (Greiner Bio-One). On the following day, sera or monoclonal antibodies were serially diluted from 1/20 starting dilution and nine twofold dilutions to the final dilution of 1/5,120 and incubated with D614 or G614 mNeonGreen SARS-CoV-2 at 37 °C for 1 h. The virus–serum mixture was transferred to the Vero cell plate with the final MOI of 2.0. After 20 h, Hoechst 33342 solution (400-fold diluted in Hank’s Balanced Salt Solution; Gibco) was added to stain cell nucleus, sealed with Breath-Easy sealing membrane (Diversified Biotech), incubated at 37 °C for 20 min, and quantified for mNeonGreen fluorescence using Cytation 7 (BioTek). The raw images (2 × 2 montage) were acquired using a 4× objective, processed and stitched using the default setting. The total cells (indicated by nucleus staining) and mNeonGreen-positive cells were quantified for each well. Infection rates were determined by dividing the mNeonGreen-positive cell number to total cell number. Relative infection rates were obtained by normalizing the infection rates of serum-treated groups to those of non-serum-treated controls. The curves of the relative infection rates versus the serum dilutions (log10 values) were plotted using Prism 8 (GraphPad). A nonlinear regression method was used to determine the dilution fold that neutralized 50% of mNeonGreen fluorescence (NT50). Each serum was tested in duplicates.
Statistics
Male hamsters were randomly allocated into different groups. The investigators were not blinded to allocation during the experiments or to the outcome assessment. A priori power analysis for two independent groups having a 1.0 log10 titre difference with a standard deviation of 0.5 log10, assuming an alpha of 0.05 and a power of 0.8 with a two-tailed t-test, dictated that animal experiments include n = 6 subjects per cohort. Descriptive statistics have been provided in the figure legends. For in vitro replication kinetics, Kruskal–Wallis analysis of variance was conducted to detect any significant variation among replicates. If no significant variation was detected, the results were pooled for further comparison. Differences between continuous variables were assessed with a non-parametric Mann–Whitney test. Hamster weights were analysed by two-factor ANOVA, with the per cent weight change as the dependent variable and the strain and time as fixed factors. Tukey’s post hoc test was used to compare all cohort pairs on days 1–7 after inoculation. log10-tranformed titres were analysed by two-factor repeated measures ANOVA with the organ and strain as fixed factors. Sidak’s post hoc test was used to compare strains within each organ. Genomic RNA/PFU ratios were calculated from non-transformed values, and the resulting ratios were log10-transformed before two-factor repeated measures ANOVA with the organ and strain as fixed factors and Sidak’s post hoc test to compare strains within a given organ. When a sample was below the limit of detection, it was treated as half of the limit of detection value for statistical and graphing purposes. Analysis was performed in Prism v.7.03 (GraphPad).
For virus competition experiments, relative replicative fitness values for G614 strain over D614 strain were analysed according to w = (f0/i0), in which i0 is the initial D614/G614 ratio and f0 is the final D614/G614 ratio after competition. Sanger sequencing (initial time point T0) counts for each virus strain being compared were based on average counts over three replicate samples of inocula per experiment, and post-infection (time point T1) counts were taken from samples of individual subjects. For the primary human airway samples, multiple experiments were performed, so that f0/i0 was clustered by experiment. To model f0/i0, the ratio T0/T1 was found separately for each subject in each strain group, log10-transformed to an improved approximation of normality, and modelled by analysis of variance with relation to group, adjusting by experiment when appropriate to control for clustering within experiment. Specifically, the model was of the form Log10_CountT1overCountT0 ~ Experiment + Group. Fitness ratios between the two groups (the model’s estimate of w = (f0/i0)) were assessed per the coefficient of the model’s Group term, which was transformed to the original scale as 10 to the power of the coefficient. This modelling approach compensates for any correlation due to clustering within experiment similarly to that of corresponding mixed-effect models, and is effective since the number of experiments was small. Statistical analyses were performed using R statistical software (R Core Team, 2019, v.3.6.1). In all statistical tests, two-sided alpha = 0.05. Cat’s-eye plots33, which illustrate the normal distribution of the model-adjusted means, were produced using the catseyes package34.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Data associated with all figures may be accessed via Figshare at https://doi.org/10.6084/m9.figshare.13030430. Source data are provided with this paper.
Change history
15 June 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-03657-2
References
Korber, B. et al. Tracking changes in SARS-CoV-2 spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell 182, 812–827 (2020).
Yurkovetskiy, L. et al. Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell 183, 739–751 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Domingo, P. et al. The four horsemen of a viral apocalypse: the pathogenesis of SARS-CoV-2 infection (COVID-19). eBioMedicine 58, 102887 (2020).
Smith, E. C., Blanc, H., Surdel, M. C., Vignuzzi, M. & Denison, M. R. Coronaviruses lacking exoribonuclease activity are susceptible to lethal mutagenesis: evidence for proofreading and potential therapeutics. PLoS Pathog. 9, e1003565 (2013).
Fauver, J. R. et al. Coast-to-coast spread of SARS-CoV-2 during the early epidemic in the United States. Cell 181, 990–996 (2020).
Song, H. D. et al. Cross-host evolution of severe acute respiratory syndrome coronavirus in palm civet and human. Proc. Natl Acad. Sci. USA 102, 2430–2435 (2005).
Hou, Y. J. et al. SARS-CoV-2 reverse genetics reveals a variable infection gradient in the respiratory tract. Cell 182, 429–446 (2020).
Lorenzo-Redondo, R. et al. A unique clade of SARS-CoV-2 viruses is associated with lower viral loads in patient upper airways. Preprint at https://doi.org/10.1101/2020.05.19.20107144 (2020).
Xie, X. et al. An infectious cDNA clone of SARS-CoV-2. Cell Host Microbe 27, 841–848 (2020).
Harcourt, J. et al. Severe acute respiratory syndrome coronavirus 2 from patient with coronavirus disease, United States. Emerg. Infect. Dis. 26, 1266–1273 (2020).
Chan, J. F. et al. Simulation of the clinical and pathological manifestations of coronavirus disease 2019 (COVID-19) in golden Syrian hamster model: implications for disease pathogenesis and transmissibility. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa325 (2020).
Wiser, M. J. & Lenski, R. E. A comparison of methods to measure fitness in Escherichia coli. PLoS ONE 10, e0126210 (2015).
Grubaugh, N. D. et al. Genetic drift during systemic arbovirus infection of mosquito vectors leads to decreased relative fitness during host switching. Cell Host Microbe 19, 481–492 (2016).
Bergren, N. A. et al. “Submergence” of Western equine encephalitis virus: evidence of positive selection argues against genetic drift and fitness reductions. PLoS Pathog. 16, e1008102 (2020).
Coffey, L. L. & Vignuzzi, M. Host alternation of chikungunya virus increases fitness while restricting population diversity and adaptability to novel selective pressures. J. Virol. 85, 1025–1035 (2011).
Bai, J. et al. Phenotypic responses of differentiated asthmatic human airway epithelial cultures to rhinovirus. PLoS ONE 10, e0118286 (2015).
Jackson, G. R., Jr, Maione, A. G., Klausner, M. & Hayden, P. J. Prevalidation of an acute inhalation toxicity test using the EpiAirway in vitro human airway model. Appl. In Vitro Toxicol. 4, 149–158 (2018).
Mulligan, M. J. et al. Phase I/II study of COVID-19 RNA vaccine BNT162b1 in adults. Nature 586, 589–593 (2020).
Jackson, L. A. et al. An mRNA vaccine against SARS-CoV-2—preliminary report. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2022483 (2020).
Muruato, A. E. et al. A high-throughput neutralizing antibody assay for COVID-19 diagnosis and vaccine evaluation. Nat. Commun. 11, 4059 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280 (2020).
Hoffmann, M., Kleine-Weber, H. & Pohlmann, S. A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol. Cell 181, 779–784 (2020).
Zhang, L. et al. The D614G mutation in the SARS-CoV-2 spike protein reduces S1 shedding and increases infectivity. Preprint at https://doi.org/10.1101/2020.06.12.148726 (2020).
Daniloski, Z., Guo, X. & Sanjana, N. E. The D614G mutation in SARS-CoV-2 spike increases transduction of multiple human cell types. Preprint at https://doi.org/10.1101/2020.06.14.151357 (2020).
Wölfel, R. et al. Virological assessment of hospitalized patients with COVID-2019. Nature 581, 465–469 (2020).
Sungnak, W. et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 26, 681–687 (2020).
Wagner, C. et al. Comparing Viral Load and Clinical Outcomes in Washington State Across D614G Substitution in Spike Protein of SARS-CoV-2 https://github.com/blab/ncov-wa-d614g (2020).
Yixuan, J. et al. SARS-CoV-2 D614G variant exhibits enhanced replication ex vivo and earlier transmission in vivo. Preprint at https://doi.org/10.1101/2020.09.28.317685 (2020).
Richard, M. et al. Influenza A viruses are transmitted via the air from the nasal respiratory epithelium of ferrets. Nat. Commun. 11, 766 (2020).
Carr, I. M. et al. Inferring relative proportions of DNA variants from sequencing electropherograms. Bioinformatics 25, 3244–3250 (2009).
Johnson, B. A. et al. Furin cleavage site is key to SARS-CoV-2 pathogenesis. Preprint at https://doi.org/10.1101/2020.08.26.268854 (2020).
Cumming, G. The new statistics: why and how. Psychol. Sci. 25, 7–29 (2014).
Andersen, C. Catseyes: create catseye plots illustrating the normal distribution of the means. R package version 0.2.3 (2019).
Reznik, G. et al. Clinical Anatomy of the European Hamster Cricetus cricetus, L. (Supt of Docs, US Govt. Print. Off., 1978).
Acknowledgements
This research was supported by grants from NIA and NIAID of the NIH (AI153602 and AG049042 to V.D.M.; R24AI120942 (WRCEVA) to S.C.W.) and by STARs Award provided by the University of Texas System to V.D.M. X.X. was partially supported by NIH5UC7AI094660. P.-Y.S. was supported by NIH grants AI134907, AI145617 and UL1TR001439, and awards from the Sealy & Smith Foundation, the Kleberg Foundation, the John S. Dunn Foundation, the Amon G. Carter Foundation, the Gilson Longenbaugh Foundation, and the Summerfield Robert Foundation. A.E.M. is supported by a Clinical and Translational Science Award NRSA (TL1) Training Core (TL1TR001440) from NIH. J.L. and C.R.F.-G. were supported by the McLaughlin Fellowship at the University of Texas Medical Branch.
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.L., V.D.M., X.X., K.S.P., S.C.W. and P.-Y.S.; methodology: J.A.P., Y.L., J.L., H.X., B.A.J., K.G.L., X.Z., A.E.M., J.Z., C.R.F.-G., A.N.F., V.D.M., X.X., K.S.P., S.C.W. and P.-Y.S.; investigation: J.A.P., Y.L., J.L., H.X., B.A.J., K.G.L., X.Z., A.E.M., J.Z., C.R.F.-G., D.M., D.S., J.P.B., B.K., A.N.F., V.D.M., X.X., K.S.P., S.C.W. and P.-Y.S.; resources: Z.K. and Z.A.; data curation: J.A.P., Y.L., J.L., H.X., B.A.J., K.G.L., X.Z., A.E.M., J.Z., A.N.F., V.D.M., X.X., K.S.P., S.C.W. and P.-Y.S.; writing, original draft: J.A.P., Y.L., J.L., X.X., K.S.P., S.C.W. and P.-Y.S; writing, review and editing: J.A.P., Y.L., J.L., H.X., B.A.J., K.G.L., X.Z., A.E.M., J.Z., A.N.F., V.D.M., X.X., K.S.P., S.C.W. and P.-Y.S.; supervision: A.N.F., V.D.M., X.X., K.S.P., S.C.W. and P.-Y.S.; and funding acquisition: A.N.F., V.D.M., S.C.W. and P.-Y.S.
Corresponding authors
Ethics declarations
Competing interests
X.X., V.D.M. and P.-Y.S. have filed a patent on the reverse genetic system and reporter SARS-CoV-2. The other authors declare no competing interests.
Additional information
Peer review information Nature thanks Stanley Perlman, K. Y. Yuen and the other, anonymous, reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Experimental design of hamster infection and sample collection.
a, Graphical overview of experiment to assess the impact of G614 mutation on replication in the respiratory system of hamsters. b, Schematic organs collected from hamsters killed on days 2, 4, and 7 post-infection. Illustration of hamster lung adapted from ref. 35.
Extended Data Fig. 2 Validation of competition assay by Sanger sequencing.
a, The correlation between input PFU ratios and output RT–PCR amplicon ratios determined by Sanger sequencing. D614 and G614 viruses were mixed at PFU ratios of 10:1, 5:1, 3:1, 1:1, 1:3, 1:5, or 1:10. Total RNA of the virus mixtures were extracted and amplified by RT–PCR. The D614/G614 ratios were calculated by the peak heights of Sanger sequencing. Data were analysed by linear regression with correlation coefficients (r) and significance (p). Symbols represent individual replicates and the solid line represents the fitted line. Data are derived from a single experiment conducted in duplicate. b, Assay range evaluation. The ratio of D614/G614virus mixture calculated from Sanger sequencing was consistent when using a wide range of virus amounts. The D614/G614 viruses were mixed at 1:1PFU ratio. The total titres of the mixed viruses were 102, 103, 104, 105, and 106 PFU. The total RNA of virus mixture was isolated and amplified by RT–PCR. The D614/G614 ratios were calculated by the peak heights from Sanger sequencing. Symbols represent individual replicates, bar heights represent the mean, and error bars represent the standard deviation. Data are derived from a single experiment conducted in triplicate.
Extended Data Fig. 3 D614G substitution significantly enhances SARS-CoV-2 replication in primary human airway tissues from a different donor.
D614 and G614 viruses were equally mixed and inoculated onto the airway tissue at a total MOI of 5. This airway tissue was produced from a different donor than that used in Fig. 3. The tissues were washed by DPBS to collect the secreted viruses every day from days 1 to 5. The total RNAs were isolated and amplified by RT–PCR. The ratio of D614 and G614 viruses after competition were measure by Sanger sequencing. Circles represent individual samples (n = 6, two independent experiments conducted in triplicate). The midline represents the sample mean, and the shaded region represents s.e.m. The width of the catseye plot represents the distribution of the model-adjusted means, and the heights extend to span 99.8% of the distribution of the mean. The y-axis is displayed on the log10 scale such that the null value is 1. P values are calculated for the group (strain) coefficient for each linear regression model, and are reported for all instances of P < 0.05.
Extended Data Fig. 4 SARS-CoV-2 G614 is more stable than D614.
a–c, Equal amounts (105 PFU/ml) of D614 and G614 viruses were incubated in DPBS at 42 °C (a), at 37 °C(b), or 33 °C (c), respectively. At 0 h, 0.5 h, 1 h, 2 h, 4 h, and 8 h, the viruses were quantified for their infectious levels by plaque assay on Vero E6 cells. The detect limitation of plaque assay is 10 PFU/ml. The percentage of remaining infectious viruses were normalized by the average titres at 0 h. Symbols represent individual replicates, bar heights represent the mean, and error bars represent the standard deviation. P values were determined by two-tailed Mann–Whitney test (n = 9, from three independent experiments, each conducted in triplicate), and results of P < 0.05 are indicated.
Extended Data Fig. 5 Scheme for preparing the D614 SARS-CoV-2-infected hamster sera for neutralization assay.
Eight sera were collected: Four sera (number 1-4) collected on day 28 post infection and another four sera (number 5-8) collected on day 49 after the second viral infection.
Extended Data Fig. 6 Construction of G614 mNeonGreen SARS-CoV-2.
a, Diagram of the construction. The D614G mutation was introduced into a mNeonGreen reporter SARS-CoV-2 using the method as previously described10. b, Plaque morphologies of D614 and G614 mNeonGreen SARS-CoV-2.
Extended Data Fig. 7 Neutralization activities of hamster sera against D614 and G614 mNeonGreen SARS-CoV-2.
a, Neutralizing curves of eight hamster sera against D614 and G614 mNeonGreenSARS-CoV-2. The neutralizing curve for serum 5 is shown in Fig. 4c. Symbols represent individual samples and the solid line represents the fitted curve. Data are derived from a single experiment conducted in duplicate. b, Calculated NT50 values and ratios of 1/NT50 for all eight hamster sera. The mean ratios were determined by (D614 1/NT50)/(G614 1/NT50).
Extended Data Fig. 8 Neutralization activities of human mAbs against D614 and G614 mNeonGreen SARS-CoV-2 in experiment I.
a, Neutralizing curves of eleven mAbs against D614 and G614 reporter SARS-CoV-2. The neutralizing curve for mAb18 is shown in Fig. 4f. Symbols represent individual replicates and the solid line represents the fitted curve. Data are derived from a single experiment conducted in duplicate. b, Calculated NT50 values for all eleven mAbs.
Extended Data Fig. 9 Neutralization activities of human mAbs against D614 and G614 mNeonGreen SARS-CoV-2 in experiment II.
a, Neutralizing curves of eleven mAbs against D614 and G614 reporter SARS-CoV-2. Symbols represent individual replicates and the solid line represents the fitted curve. Data are derived from a single experiment conducted in either duplicate or quadruplicate. b, Calculated NT50 values for all eleven mAbs. (c) Summary of NT50 ratios from two independent experiments. The ratios were determined by (D614 NT50)/(G614 NT50).
Supplementary information
Supplementary Figure 1
Uncropped chemiluminescent, colorimetric, and merged images of Western blots from Figures 1i and 1j.
Rights and permissions
About this article
Cite this article
Plante, J.A., Liu, Y., Liu, J. et al. Spike mutation D614G alters SARS-CoV-2 fitness. Nature 592, 116–121 (2021). https://doi.org/10.1038/s41586-020-2895-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2895-3
This article is cited by
-
SARS-CoV-2 outbreak: role of viral proteins and genomic diversity in virus infection and COVID-19 progression
Virology Journal (2024)
-
Positive selection underlies repeated knockout of ORF8 in SARS-CoV-2 evolution
Nature Communications (2024)
-
Mucosal vaccine-induced cross-reactive CD8+ T cells protect against SARS-CoV-2 XBB.1.5 respiratory tract infection
Nature Immunology (2024)
-
Detection of SARS-CoV-2 spike protein D614G mutation using μTGGE
Molecular Biology Reports (2024)
-
Evaluation of the PrimeFlow RNA assay as a method of detection of SARS-CoV-2 single and dual Infections
Cytotechnology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.