Abstract
Telomeres—repeated, noncoding nucleotide motifs and associated proteins that are found at the ends of eukaryotic chromosomes—mediate genome stability and determine cellular lifespan1. Telomeric-repeat-containing RNA (TERRA) is a class of long noncoding RNAs (lncRNAs) that are transcribed from chromosome ends2,3; these RNAs in turn regulate telomeric chromatin structure and telomere maintenance through the telomere-extending enzyme telomerase4,5,6 and homology-directed DNA repair7,8. The mechanisms by which TERRA is recruited to chromosome ends remain poorly defined. Here we develop a reporter system with which to dissect the underlying mechanisms, and show that the UUAGGG repeats of TERRA are both necessary and sufficient to target TERRA to chromosome ends. TERRA preferentially associates with short telomeres through the formation of telomeric DNA–RNA hybrid (R-loop) structures that can form in trans. Telomere association and R-loop formation trigger telomere fragility and are promoted by the recombinase RAD51 and its interacting partner BRCA2, but counteracted by the RNA-surveillance factors RNaseH1 and TRF1. RAD51 physically interacts with TERRA and catalyses R-loop formation with TERRA in vitro, suggesting a direct involvement of this DNA recombinase in the recruitment of TERRA by strand invasion. Together, our findings reveal a RAD51-dependent pathway that governs TERRA-mediated R-loop formation after transcription, providing a mechanism for the recruitment of lncRNAs to new loci in trans.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Maciejowski, J. & de Lange, T. Telomeres in cancer: tumour suppression and genome instability. Nat. Rev. Mol. Cell Biol. 18, 175–186 (2017).
Azzalin, C. M., Reichenbach, P., Khoriauli, L., Giulotto, E. & Lingner, J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318, 798–801 (2007).
Schoeftner, S. & Blasco, M. A. Developmentally regulated transcription of mammalian telomeres by DNA-dependent RNA polymerase II. Nat. Cell Biol. 10, 228–236 (2008).
Redon, S., Zemp, I. & Lingner, J. A three-state model for the regulation of telomerase by TERRA and hnRNPA1. Nucleic Acids Res. 41, 9117–9128 (2013).
Cusanelli, E., Romero, C. A. P. & Chartrand, P. Telomeric noncoding RNA TERRA is induced by telomere shortening to nucleate telomerase molecules at short telomeres. Mol. Cell 51, 780–791 (2013).
Moravec, M. et al. TERRA promotes telomerase-mediated telomere elongation in Schizosaccharomyces pombe. EMBO Rep. 17, 999–1012 (2016).
Arora, R. et al. RNaseH1 regulates TERRA-telomeric DNA hybrids and telomere maintenance in ALT tumour cells. Nat. Commun. 5, 5220 (2014).
Graf, M. et al. Telomere length determines TERRA and R-loop regulation through the cell cycle. Cell 170, 72–85 (2017).
Porro, A., Feuerhahn, S., Reichenbach, P. & Lingner, J. Molecular dissection of telomeric repeat-containing RNA biogenesis unveils the presence of distinct and multiple regulatory pathways. Mol. Cell. Biol. 30, 4808–4817 (2010).
Larson, D. R., Zenklusen, D., Wu, B., Chao, J. A. & Singer, R. H. Real-time observation of transcription initiation and elongation on an endogenous yeast gene. Science 332, 475–478 (2011).
Hockemeyer, D. et al. Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nat. Biotechnol. 27, 851–857 (2009).
Jensen, R. B., Carreira, A. & Kowalczykowski, S. C. Purified human BRCA2 stimulates RAD51-mediated recombination. Nature 467, 678–683 (2010).
Thorslund, T. et al. The breast cancer tumor suppressor BRCA2 promotes the specific targeting of RAD51 to single-stranded DNA. Nat. Struct. Mol. Biol. 17, 1263–1265 (2010).
Mason, J. M., Chan, Y.-L., Weichselbaum, R. W. & Bishop, D. K. Non-enzymatic roles of human RAD51 at stalled replication forks. Nat. Commun. 10, 4410 (2019).
Boguslawski, S. J. et al. Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids. J. Immunol. Methods 89, 123–130 (1986).
Sagie, S. et al. Telomeres in ICF syndrome cells are vulnerable to DNA damage due to elevated DNA:RNA hybrids. Nat. Commun. 8, 14015 (2017).
Pfeiffer, V., Crittin, J., Grolimund, L. & Lingner, J. The THO complex component Thp2 counteracts telomeric R-loops and telomere shortening. EMBO J. 32, 2861–2871 (2013).
Balk, B. et al. Telomeric RNA-DNA hybrids affect telomere-length dynamics and senescence. Nat. Struct. Mol. Biol. 20, 1199–1205 (2013).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Crossley, M. P., Bocek, M. & Cimprich, K. A. R-loops as cellular regulators and genomic threats. Mol. Cell 73, 398–411 (2019).
Wahba, L., Gore, S. K. & Koshland, D. The homologous recombination machinery modulates the formation of RNA–DNA hybrids and associated chromosome instability. eLife 2, e00505 (2013).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protocols 8, 2281–2308 (2013).
Grolimund, L. et al. A quantitative telomeric chromatin isolation protocol identifies different telomeric states. Nat. Commun. 4, 2848 (2013).
Feretzaki, M. & Lingner, J. A practical qPCR approach to detect TERRA, the elusive telomeric repeat-containing RNA. Methods 114, 39–45 (2017).
Porro, A. et al. Functional characterization of the TERRA transcriptome at damaged telomeres. Nat. Commun. 5, 5379 (2014).
Porro, A., Feuerhahn, S. & Lingner, J. TERRA-reinforced association of LSD1 with MRE11 promotes processing of uncapped telomeres. Cell Rep. 6, 765–776 (2014).
Špírek, M. et al. Human RAD51 rapidly forms intrinsically dynamic nucleoprotein filaments modulated by nucleotide binding state. Nucleic Acids Res. 46, 3967–3980 (2018).
Acknowledgements
We thank D. Larson (NIH), D. Bishop (Univ. Chicago), V. Simanis (EPFL), P. Gönczy (EPFL) and D. Trono (EPFL) for providing material. We also thank the members of the BIOP core facility at EPFL, members of the Gönczy laboratory and M. Spirek for technical support and advice and J. Cibulka for recombinant mutant RAD51 protein. M.F. was supported by the European Union’s Horizon 2020 research and innovation programme under Marie Skłodowska-Curie grant agreement 702824. J.L.’s laboratory was supported by the Swiss National Science Foundation (SNFS grant 310030_184718), the SNFS-funded National Centres of Competence in Research (NCCR) RNA and disease network (grant 182880), and EPFL. L.K.’s laboratory was supported by the European Structural and Investment Funds, Operational Programme Research, Development and Education ‘Preclinical Progression of New Organic Compounds with Targeted Biological Activity’ (Preclinprogress) (CZ.02.1.01/0.0/0.0/16_025/0007381); a Wellcome Trust Collaborative Grant 206292/E/17/Z; the Czech Science Foundation (GACR 17-17720S); and the National Program of Sustainability II (MEYS CR, project number LQ1605). Both J.L. and L.K. were also supported by the European Union's Horizon 2020 research and innovation programme under grant agreement 812829.
Author information
Authors and Affiliations
Contributions
M.F. and J.L. conceived the study. M.F. and R.V.F. executed all cell and molecular biology experiments. M.P. performed all biochemistry experiments and T.L. some EMSA experiments. L.K. conceived the RAD51-based biochemistry experiments and advised on the text. J.L. and M.F. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Co-localization of transgenic TERRA with telomeres.
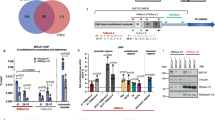
a, Quantification of co-localization of transgenic RNA foci expressed from plasmids with telomeres. b, Strategy for targeting the AAVS1 locus through CRISPR–CAS9 to integrate chimaeric TERRA constructs into the genome. LHA and RHA, left and right homology arms. c, Quantification of co-localization of transgenic RNA foci with telomeres when TERRA was expressed from the AAVS1 locus. d, Confocal images obtained when TERRA was expressed from the AAVS1 locus. For a, c, n = 3 biologically independent experiments; data are mean ± s.d.; two-tailed unpaired t-test: ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 2 TERRA associates preferentially with short telomeres.
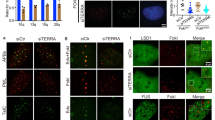
a, TRF analysis of HeLa clones used for transient expression of TERRA. b, d, TERRA expression levels measured by RT–qPCR relative to the levels of TERRA in the clone with the shortest telomeres. n = 3 biologically independent experiments; data are means ± s.d. c, TRF analysis of HeLa clones expressing transgenic TERRA from the AAVS1 locus. e, HeLa clones of different telomere lengths were transiently transfected with the PP7–15qTERRA construct and co-localization was assessed by immunofluorescence with FISH (bottom). White arrows indicate co-localization of PP7 foci with the telomeric signal. The percentage of TERRA foci that co-localized with telomeres is plotted as a function of telomere length (top). Random colocalization events (roughly 3%) were not subtracted. f, Co-localization events as in e but for PP7–15qTERRA expressed from the AAVS locus on chromosome 19. n = 3 biologically independent experiments; data are means ± s.d. Significant differences are indicated. Two-tailed unpaired t-tests were used to calculate P-values: *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 3 Depletion of factors regulating TERRA trafficking.
a, Timeline of transfections and cell harvesting. mRNA levels were determined by RT–qPCR for long-telomere and short-telomere cell lines. b, For the second replicate of each siRNA screen, the level of depletion was also evaluated on western blots. Vinculin and hnRNPA1 were used as loading controls. c, Numbers of TERRA foci per cell are plotted for each depleted factor. n = 2 biologically independent experiments; at least 40 nuclei were analysed per condition. d, Quantification of endogenous TERRA stemming from the ends of chromosomes 15q, 13q and 10q upon depletion of the indicated factors relative to the negative control (siCONTROL). n = 2 biologically independent experiments.
Extended Data Fig. 4 Depletion of RAD51 and BRCA2, which regulate the association of TERRA with telomeres.
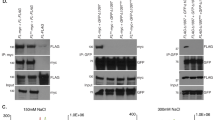
a, HeLa clones with long and short telomeres were transfected with siRAD51 and siBRCA2, and then with chimaeric TERRA constructs. Representative immunoblots show RAD51 and BRCA2 depletion, with vinculin as a loading control. Upon depletion, co-localization of TERRA with telomeres was assessed with immunofluorescence–FISH. n = 2 biologically independent experiments; at least 54 nuclei were analysed per condition; data are means ± s.d. One-way ANOVA with Dunnett’s multiple comparisons test was used, comparing all conditions to siCONTROL: *P < 0.05; **P < 0.01; ***P < 0.001. b, Detection of endogenous and transgenic RAD51 on a western blot. Endogenous RAD51 was depleted with siRNA, and wild-type (WT) RAD51 or the RAD51 II3A mutant was expressed from plasmids containing complementary DNA.
Extended Data Fig. 5 RNaseH1-regulated formation of telomeric R-loops in trans.
a, RNH1 depletion and overexpression (OE) was assessed by western blotting in cell lines with long and short telomeres and upon TERRA overexpression. b, Representative example of one DRIP assay followed by qPCR for the end of chromosome 13q. Left, quantification of RNA–DNA hybrids is expressed as a fraction of input. Right, the amplified DNA was run on a gel, then isolated and sequenced.
Extended Data Fig. 6 Transgenic TERRA expressed from plasmids induces telomere fragility.
a, Representative examples of metaphase chromosomes stained with FISH to visualize telomeres. DNA is stained with DAPI. Fragile telomeres are indicated by white arrowheads. b, Quantification of telomere fragility. c, Quantification of telomere fragility in cells expressing PP7–15qTERRA in which expression of RNaseH1 (RNH1) and RAD51 was manipulated as indicated. For b, c, the numbers of metaphases scored over three biologically independent experiments are indicated for each condition as n.
Extended Data Fig. 7 RAD51 binds TERRA and promotes R-loop formation.
a, Detection of endogenous telomeric R-loops on dot-blots as in Fig. 2a. Cells were transfected with siCONTROL (siC), siRNH1 or siRAD51. b, Possible models for the roles of RAD51 in mediating TERRA–telomere associations. Upper row, RAD51 binds telomeric single-stranded DNA, inducing strand invasion of another telomere. TERRA hybridizes to the displaced strand upon branch migration. Lower row, RAD51 binds TERRA directly and initiates homology search and strand invasion by TERRA at a telomere. c, Western blot analysis of RAD51 and hnRNPA1 upon immunoprecipitation (IP) of native RNA (see Fig. 3a for RNA analysis). SN, supernatant. d, Native Immunoprecipitation of RNA was performed in U2OS cell extracts, demonstrating the association of TERRA with RAD51 and hnRNPA1. n = 3 biologically independent experiments; data are means ± s.d. e, Affinity of RAD51 for TERRA and non-telomeric-RNA (Non-TelRNA) oligonucleotides. Quantification is shown on the right. n = 3 independent experiments; data are means ± s.d. f, Top, stability EMSA assay. Bottom, 20 nM TERRA (lanes 2-7) or TelDNA (lanes 9-14) oligonucleotides were pre-bound with RAD51 (2.2 µM or 8.8 µM) and challenged with increasing concentrations of unlabelled competitor ssDNA (cDNA; 0.28 μM, 0.56 μM, 1.13 μM, 2.27 μM or 4.54 μM). Quantification is shown on the right. n = 3 independent experiments; data are means ± s.d.
Extended Data Fig. 8 RAD51 catalyses the formation of canonical R-loops.
a, Left, the RAD51 II3A mutant (lines 2–5; 325 nM, 650 nM, 1,300 nM or 2,600 nM) or wild-type RAD51 (lines 6 and 7; 325 nM or 650 nM) was incubated with TERRA oligonucleotide substrate (50 nM), and then plasmid containing the homologous region was added. Right, quantification of R-loops in the presence of wild-type RAD51 or RAD51 II3A. n = 2 independent experiments; data are means ± s.d. b, Top, the R-loop and D-loop assays. After RAD51-mediated R-loop or D-loop formation, proteins were digested with proteinase K (PK), which was then inactivated with PMSF and EGTA. Bottom left, R-loops or D-loops were detected on native gels as indicated. Treatment with RNaseH1, which degrades the RNA moiety in RNA–DNA hybrid structures, eliminates R-loops but has no effect on D-loops. RNaseH2, which cleaves ribonucleotides in DNA, has no effect, as expected. Right, quantification. n = 3 independent experiments; data are means ± s.d. c, Top, the R-loop and D-loop assays. Middle and bottom, RAD51-mediated R-loops (middle; n = 3 independent experiments; data are means ± s.d.), unlike D-loops (n=1 experiment), are recognized by the S9.6 antibody and supershifted in presence of both S9.6 and anti-mouse IgG. Quantification is shown on the right.
Supplementary information
Supplementary Figures
This file contains a source file for figures.
Rights and permissions
About this article
Cite this article
Feretzaki, M., Pospisilova, M., Valador Fernandes, R. et al. RAD51-dependent recruitment of TERRA lncRNA to telomeres through R-loops. Nature 587, 303–308 (2020). https://doi.org/10.1038/s41586-020-2815-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2815-6
This article is cited by
-
R-loop and diseases: the cell cycle matters
Molecular Cancer (2024)
-
TERRA-LSD1 phase separation promotes R-loop formation for telomere maintenance in ALT cancer cells
Nature Communications (2024)
-
Unveiling the distinctive traits of functional rye centromeres: minisatellites, retrotransposons, and R-loop formation
Science China Life Sciences (2024)
-
Genome maintenance meets mechanobiology
Chromosoma (2024)
-
The chromatin-associated RNAs in gene regulation and cancer
Molecular Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.