Abstract
Plants grow within a complex web of species that interact with each other and with the plant1,2,3,4,5,6,7,8,9,10. These interactions are governed by a wide repertoire of chemical signals, and the resulting chemical landscape of the rhizosphere can strongly affect root health and development7,8,9,11,12,13,14,15,16,17,18. Here, to understand how interactions between microorganisms influence root growth in Arabidopsis, we established a model system for interactions between plants, microorganisms and the environment. We inoculated seedlings with a 185-member bacterial synthetic community, manipulated the abiotic environment and measured bacterial colonization of the plant. This enabled us to classify the synthetic community into four modules of co-occurring strains. We deconstructed the synthetic community on the basis of these modules, and identified interactions between microorganisms that determine root phenotype. These interactions primarily involve a single bacterial genus (Variovorax), which completely reverses the severe inhibition of root growth that is induced by a wide diversity of bacterial strains as well as by the entire 185-member community. We demonstrate that Variovorax manipulates plant hormone levels to balance the effects of our ecologically realistic synthetic root community on root growth. We identify an auxin-degradation operon that is conserved in all available genomes of Variovorax and is necessary and sufficient for the reversion of root growth inhibition. Therefore, metabolic signal interference shapes bacteria–plant communication networks and is essential for maintaining the stereotypic developmental programme of the root. Optimizing the feedbacks that shape chemical interaction networks in the rhizosphere provides a promising ecological strategy for developing more resilient and productive crops.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The 16S rRNA amplicon sequencing data associated with this study have been deposited in the NCBI Sequence Read Archive under the project accession PRJNA543313. The raw transcriptomic data have been deposited in the Gene Expression Omnibus under the accession GSE131158. We deposited all scripts and additional data structures required to reproduce the results of this study in the following GitHub repository: https://github.com/isaisg/variovoraxRGI. Source data are provided with this paper.
References
Castrillo, G. et al. Root microbiota drive direct integration of phosphate stress and immunity. Nature 543, 513–518 (2017).
Durán, P. et al. Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175, 973–983.e14 (2018).
Herrera Paredes, S. et al. Design of synthetic bacterial communities for predictable plant phenotypes. PLoS Biol. 16, e2003962 (2018).
Fitzpatrick, C. R. et al. Assembly and ecological function of the root microbiome across angiosperm plant species. Proc. Natl Acad. Sci. USA 115, E1157–E1165 (2018).
Finkel, O. M. et al. The effects of soil phosphorus content on plant microbiota are driven by the plant phosphate starvation response. PLoS Biol. 17, e3000534 (2019).
Thiergart, T. et al. Root microbiota assembly and adaptive differentiation among European Arabidopsis populations. Nat. Ecol. Evol. 4, 122–131 (2020).
Hogenhout, S. A., Van der Hoorn, R. A. L., Terauchi, R. & Kamoun, S. Emerging concepts in effector biology of plant-associated organisms. Mol. Plant Microbe Interact. 22, 115–122 (2009).
Mylona, P., Pawlowski, K. & Bisseling, T. Symbiotic nitrogen fixation. Plant Cell 7, 869–885 (1995).
Ludwig-Müller, J. Bacteria and fungi controlling plant growth by manipulating auxin: balance between development and defense. J. Plant Physiol. 172, 4–12 (2015).
Carlström, C. I. et al. Synthetic microbiota reveal priority effects and keystone strains in the Arabidopsis phyllosphere. Nat. Ecol. Evol. 3, 1445–1454 (2019).
Faure, D., Vereecke, D. & Leveau, J. H. J. Molecular communication in the rhizosphere. Plant Soil 321, 279–303 (2009).
Leadbetter, J. R. & Greenberg, E. P. Metabolism of acyl-homoserine lactone quorum-sensing signals by Variovorax paradoxus. J. Bacteriol. 182, 6921–6926 (2000).
Leveau, J. H. J. & Lindow, S. E. Utilization of the plant hormone indole-3-acetic acid for growth by Pseudomonas putida strain 1290. Appl. Environ. Microbiol. 71, 2365–2371 (2005).
Zúñiga, A. et al. Quorum sensing and indole-3-acetic acid degradation play a role in colonization and plant growth promotion of Arabidopsis thaliana by Burkholderia phytofirmans PsJN. Mol. Plant Microbe Interact. 26, 546–553 (2013).
Sun, S.-L. et al. The plant growth-promoting rhizobacterium Variovorax boronicumulans CGMCC 4969 regulates the level of indole-3-acetic acid synthesized from indole-3-acetonitrile. Appl. Environ. Microbiol. 84, e00298-18 (2018).
Gilbert, S. et al. Bacterial production of indole related compounds reveals their role in association between duckweeds and endophytes. Front. Chem. 6, 265 (2018).
Donoso, R. et al. Biochemical and genetic bases of indole-3-acetic acid (auxin phytohormone) degradation by the plant-growth-promoting rhizobacterium Paraburkholderia phytofirmans PsJN. Appl. Environ. Microbiol. 83, e01991-16 (2016).
Leveau, J. H. J. & Gerards, S. Discovery of a bacterial gene cluster for catabolism of the plant hormone indole 3-acetic acid. FEMS Microbiol. Ecol. 65, 238–250 (2008).
Brumos, J. et al. Local auxin biosynthesis is a key regulator of plant development. Dev. Cell 47, 306–318.e5 (2018).
Levy, A. et al. Genomic features of bacterial adaptation to plants. Nat. Genet. 50, 138–150 (2018).
Kremer, J. M. et al. FlowPot axenic plant growth system for microbiota research. Preprint at https://www.biorxiv.org/content/10.1101/254953v1 (2018).
Clauss, M. J. & Aarssen, L. W. Phenotypic plasticity of size–fecundity relationships in Arabidopsis thaliana. J. Ecol. 82, 447 (1994).
Abreu, M. E. & Munné-Bosch, S. Salicylic acid deficiency in NahG transgenic lines and sid2 mutants increases seed yield in the annual plant Arabidopsis thaliana. J. Exp. Bot. 60, 1261–1271 (2009).
Bai, Y. et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528, 364–369 (2015).
Klepikova, A. V., Kasianov, A. S., Gerasimov, E. S., Logacheva, M. D. & Penin, A. A. A high resolution map of the Arabidopsis thaliana developmental transcriptome based on RNA-seq profiling. Plant J. 88, 1058–1070 (2016).
Uchida, N. et al. Chemical hijacking of auxin signaling with an engineered auxin–TIR1 pair. Nat. Chem. Biol. 14, 299–305 (2018).
Takase, T. et al. ydk1-D, an auxin-responsive GH3 mutant that is involved in hypocotyl and root elongation. Plant J. 37, 471–483 (2004).
Chen, L., Dodd, I. C., Theobald, J. C., Belimov, A. A. & Davies, W. J. The rhizobacterium Variovorax paradoxus 5C-2, containing ACC deaminase, promotes growth and development of Arabidopsis thaliana via an ethylene-dependent pathway. J. Exp. Bot. 64, 1565–1573 (2013).
Cary, A. J., Liu, W. & Howell, S. H. Cytokinin action is coupled to ethylene in its effects on the inhibition of root and hypocotyl elongation in Arabidopsis thaliana seedlings. Plant Physiol. 107, 1075–1082 (1995).
Gómez-Gómez, L., Felix, G. & Boller, T. A single locus determines sensitivity to bacterial flagellin in Arabidopsis thaliana. Plant J. 18, 277–284 (1999).
Robles, L., Stepanova, A. & Alonso, J. Molecular mechanisms of ethylene–auxin interaction. Mol. Plant 6, 1734–1737 (2013).
Nagpal, P. et al. AXR2 encodes a member of the Aux/IAA protein family. Plant Physiol. 123, 563–574 (2000).
Hall, A. E., Findell, J. L., Schaller, G. E., Sisler, E. C. & Bleecker, A. B. Ethylene perception by the ERS1 protein in Arabidopsis. Plant Physiol. 123, 1449–1458 (2000).
Gould, S. J. & Vrba, E. S. Exaptation—a missing term in the science of form. Paleobiology 8, 4–15 (1982).
Bardoel, B. W. et al. Pseudomonas evades immune recognition of flagellin in both mammals and plants. PLoS Pathog. 7, e1002206 (2011).
Lundberg, D. S., Yourstone, S., Mieczkowski, P., Jones, C. D. & Dangl, J. L. Practical innovations for high-throughput amplicon sequencing. Nat. Methods 10, 999–1002 (2013).
Yourstone, S. M., Lundberg, D. S., Dangl, J. L. & Jones, C. D. MT-Toolbox: improved amplicon sequencing using molecule tags. BMC Bioinformatics 15, 284 (2014).
Joshi, N. & Fass, J. Sickle: a sliding-window, adaptive, quality-based trimming tool for FastQ files (version 1.33), https://github.com/najoshi/sickle (2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Oksanen, J. et al. Vegan: Community Ecology Package, R package version 2.5-6, https://CRAN.R-project.org/package=vegan (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Lenth, R. emmeans: Estimated Marginal Means, aka Least-Squares Means, R package version 1.4.7, https://CRAN.R-project.org/package=emmeans (2020).
Logemann, J., Schell, J. & Willmitzer, L. Improved method for the isolation of RNA from plant tissues. Anal. Biochem. 163, 16–20 (1987).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Wheeler, T. J. & Eddy, S. R. nhmmer: DNA homology search with profile HMMs. Bioinformatics 29, 2487–2489 (2013).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Gordon, S. A. & Weber, R. P. Colorimetric estimation of indoleacetic acid. Plant Physiol. 26, 192–195 (1951).
Friml, J. et al. Efflux-dependent auxin gradients establish the apical–basal axis of Arabidopsis. Nature 426, 147–153 (2003).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Ebenau-Jehle, C. et al. Anaerobic metabolism of indoleacetate. J. Bacteriol. 194, 2894–2903 (2012).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Kovach, M. E. et al. Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166, 175–176 (1995).
Figurski, D. H. & Helinski, D. R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc. Natl Acad. Sci. USA 76, 1648–1652 (1979).
Hamad, M. A., Zajdowicz, S. L., Holmes, R. K. & Voskuil, M. I. An allelic exchange system for compliant genetic manipulation of the select agents Burkholderia pseudomallei and Burkholderia mallei. Gene 430, 123–131 (2009).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Winter, D. et al. An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2, e718 (2007).
Acknowledgements
We thank S. Barth, J. Shen, M. Priegel, D. Chudasma, D. Panda, I. Castillo, N. Del Risco, C. Lindberg and R. Pérez-Torres for technical assistance; D. Pelletier and P. Schulze-Lefert for strains; the Dangl laboratory microbiome group, A. Stepanova, J. Alonso, J. Brumos, J. Kieber, J. Reed and I. Greenhut for useful discussions; and D. Lundberg and A. Bishopp for critical comments on the manuscript. This work was supported by NSF grant IOS-1917270 and by Office of Science (BER), US Department of Energy, Grant DE-SC0014395 to J.L.D. J.L.D. is an Investigator of the Howard Hughes Medical Institute, supported by the HHMI. O.M.F. was supported by NIH NRSA Fellowship F32-GM117758. C.R.F. was supported by an NSERC postdoctoral fellowship (532852-2019).
Author information
Authors and Affiliations
Contributions
J.L.D. supervised the project. O.M.F., I.S.-G., G.C. and J.L.D. conceptualized the project. O.M.F., I.S.-G., G.C., J.M.C. and J.L.D. designed experiments. O.M.F., I.S.-G., G.C. and T.F.L. designed and performed most in planta agar-based experiments. C.R.F., O.M.F. and I.S.-G. designed and performed soil experiments. O.M.F., I.S.-G., G.C. and T.F.L. processed and sequenced bacterial DNA samples. J.M.C. and E.D.W. designed and performed bacterial gain- and loss-of function experiments and bacterial RNA-seq experiments. O.M.F., T.F.L. and P.J.P.L.T. processed and sequenced plant RNA-seq samples. O.M.F. and G.C. performed plant morphometric measurements. I.S.-G., O.M.F., G.C. and J.M.C analysed data. I.S.-G. performed bioinformatics analyses, including bacterial 16S rRNA, plant and bacterial RNA-seq analysis and bacterial comparative genomics, with input from O.M.F., G.C., J.M.C. and J.L.D. I.S.-G. performed statistical analysis, with input from O.M.F., G.C., C.D.J. and J.L.D. I.S.-G. curated the data used in the manuscript with help from O.M.F. O.M.F., I.S.-G. and G.C. wrote the manuscript with input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
J.L.D. is a co-founder of, and shareholder in, AgBiome LLC, a corporation with the goal of using plant-associated microorganisms to improve plant productivity.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Synthetic community resembles the taxonomic make-up of natural communities.
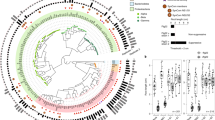
a, Phylogenetic tree of 185 genome-sequenced isolates obtained from surface-sterilized Arabidopsis roots, included in the synthetic community. The composition of this synthetic community captures the diversity of Actinobacteria, Proteobacteria and Bacteroidetes; the three major root-enriched phyla1,4,5,6,20; and Firmicutes, which are abundant in plant-associated culture collections20. Tree tips are coloured according to phylum. The outer ring shows the distribution of the 12 bacterial orders present in the synthetic community. b, Comparison of proportions of Proteobacteria, Bacteroidetes and Actinobacteria in synthetic-community (SynCom)-inoculated roots to root microbiota derived from plants grown in natural soil1. Firmicutes, which are not plant-enriched, were reduced to <0.1% of the relative abundance (Fig. 1a). The left panel (wild soil) shows the proportion of ASVs enriched (q value < 0.1) in the plant root in comparison to soil in a microbiota profiling study from the same soil from which the synthetic community strains were isolated. ASVs are coloured according to phylum and proteobacterial ASVs are coloured by class. The right panel (synthetic community) represents the relative abundance profiles of bacterial isolates across the initial inoculum, planted agar, and root and shoot in plants inoculated with the full synthetic community. c, Comparison of synthetic community composition in agar versus the soil-based microcosms. Left, relative abundance in the substrate. Middle, relative abundance in root. Right, enrichment in root versus substrate. Each dot represents a single unique sequence. Pearson correlation line, 95% confidence intervals, r value and P value are shown for each comparison. n = 24 (soil system) and 8 (agar system) biological replicates. d, Canonical analysis of principal coordinates (CAP) showing the influence of the fraction (planted agar, root or shoot) on the assembly of the bacterial synthetic community across the four gradients used in this Article (phosphate, salinity, pH and temperature). Different colours differentiate between the fractions, and different shapes differentiate between experiments. Ellipses denote the 95% confidence interval of each fraction. Fraction (substrate, root or shoot) explains most (22%) of the variance across all abiotic variables. n = 94 (substrate), 90 (root) and 95 (shoot) biological replicates across 8 independent experiments. e, Abiotic conditions displayed reproducible effects on α-diversity. Each panel represents bacterial α-diversity across the different abiotic gradients (phosphate, salinity, pH and temperature) and fractions (substrate, root and shoot) used in this Article. Bacterial α-diversity was estimated using Shannon diversity index. Letters represent the results of the post hoc test of an ANOVA model testing the interaction between fraction and abiotic condition. f, Canonical analysis of principal coordinates scatter plots showing the effect on the composition of the synthetic community of each of the four abiotic gradients (phosphate, salinity, pH and temperature) within the substrate, root and shoot fractions. PERMANOVA R2 values are shown within each plot. e, f, Phosphate, n = 6, 5, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 5 and 6; salinity n = 6, 6, 6, 6, 6, 6, 6, 4, 5, 6, 4 and 5; pH, n = 6, 6, 6, 5, 5, 6, 6, 6 and 6; temperature n = 6, 6, 6, 6, 6, 6, 6, 6 and 6. All samples are biologically independent and represent two independent experiments.
Extended Data Fig. 2 RGI trait is distributed across bacterial phylogeny.
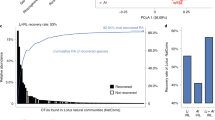
a, b, Primary root elongation of seedlings inoculated with single bacterial isolates (one box plot per isolate). Isolates are coloured by taxonomy. a, Isolates are grouped by module membership. The strips across the panels correspond to the interquartile range (IQR), as noted at the far right. The dotted line represents the cut-off used to classify isolates as root-growth inhibiting (cutoff RGI). b, Isolates are ordered according to the phylogenetic tree on the left, and coloured on the basis of their genome-based taxonomy. The vertical blue stripes across the panel correspond to the IQR of plants grown in sterile conditions. The vertical dotted line represents the 3-cm cut-off used to classify strains as RGI strains. The bar on the right denotes the genus classification of each isolate. c, Binarized image of representative seedlings grown axenically (no bacteria) or with 34 RGI strains individually.
Extended Data Fig. 3 Variovorax-mediated reversion of RGI.
a, Heat map coloured by average primary root elongation of seedlings inoculated with eighteen RGI-inducing strains (rows) alone (self) or in combination with Burkholderia CL11, Variovorax CL14 or Variovorax MF160 (columns). Statistically significant RGI reversions were determined via ANOVA and are outlined in black. b, Variovorax-mediated reversion of RGI is maintained in a second plant species. Primary root elongation of uninoculated tomato seedlings (no bacteria) or seedlings inoculated with the RGI-inducer Arthrobacter CL28 individually or along with Variovorax CL14 grown on vertical agar plates. Significance was determined via ANOVA while controlling for experiment; letters correspond to a Tukey post hoc test. n = 24, 25 and 23 biological replicates across 2 independent experiments.
Extended Data Fig. 4 Variovorax maintain stereotypic plant growth.
a, Representative plate images of plants grown with the full 185-member synthetic community (full SynCom) or the Variovorax drop-out community (−Vario SynCom) for 12 d. b, Bar graphs showing the isolate composition of synthetic communities composed by module A, module C, module D and a previously described1 34-member synthetic community (34-member). Isolates are ordered according to the phylogenetic tree on the left. The tips of the phylogenetic tree are coloured on the basis of the genome-based taxonomy of each isolate. Presence of an isolate across the different synthetic communities is denoted by a black filled rectangle (labelled ‘inoculated’). c, Total root network quantification of Arabidopsis seedlings grown with the full synthetic community (full), or with the full synthetic community excluding Variovorax (−Vario), across different abiotic conditions: JM agar control, low phosphate (JM agar 10 μM Pi), high salt (JM agar 150 mM NaCl), high pH (JM agar pH 8.2) and high temperature (JM agar 31 °C), as well as half Murashige and Skoog medium (MS agar control). Significance was determined within each condition via ANOVA while controlling for experiment. n = 110, 77, 42, 44, 41, 35, 48, 49, 44, 45, 33 and 26 biological replicates across 2 independent experiments. d, Shoot circumference of plants grown in pots with potting soil with the full synthetic community or with the full synthetic community excluding Variovorax. Significance was determined within each condition via ANOVA while controlling for experiment. n = 30 and 36 biological replicates across 2 independent experiments.
Extended Data Fig. 5 Reversion of RGI is prevalent across the Variovorax phylogeny.
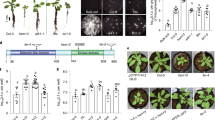
a, Phylogenetic tree of 69 publically available Variovorax genomes and 2 outgroup isolates, Acidovorax root 219 and Burkholderia CL11. The CL28 RGI reversion bar categorizes (positive, negative or untested) the ability of each isolate in the phylogeny to revert the RGI caused by Arthrobacter CL28. The ACC deaminase bar denotes the presence of the KEGG Orthology (KO) term KO1505 (1-aminocyclopropane-1-carboxylate deaminase) in each of the genomes. The heat map denotes the per cent identity of BLASTp hits in the genomes to the genes from the auxin-degrading iac operon in Paraburkholderia phytophirmans, described by ref. 17. Synteny is not necessarily conserved, as these BLAST hits may be spread throughout the genomes. b,All tested Variovorax isolates reverted RGI. Phylogenetic tree of 19 Variovorax genomes and 2 outgroup isolates (Acidovorax root 219 and Burkholderia CL11) that were tested for their ability to revert the RGI imposed by Arthrobacter CL28. The blue vertical stripe denotes the IQR of plants treated solely with Arthrobacter CL28. The dotted vertical line denotes the 3-cm cut-off used to classify a treatment as an RGI. Each box plot is coloured according to the genus classification of each isolate. Significance was determined via ANOVA while controlling for experiment, letters correspond to a Tukey post hoc test. n = 59, 9, 55, 71, 10, 10, 10, 9, 10, 10, 57, 10, 10, 48, 10, 10, 10, 9, 10, 10 and 9 biological replicates across 2 independent experiments.
Extended Data Fig. 6 Variovorax does not compete with or antagonize RGI strains.
To test whether Variovorax attenuates RGI by inhibiting the growth of RGI-inducing strains, we compared the bacterial relative abundance profiles in seedlings colonized with the full synthetic community to that of seedlings colonized with the Variovorax drop-out community. We found no changes in the abundances of RGI-inducing strains in response to the Variovorax drop-out (Fig. 2g). In addition, we measured in planta absolute abundance of Arthrobacter CL28 when inoculated alone or with two Variovorax representatives: Variovorax B4 and Variovorax CL14. Here we show log-transformed CFUs of Arthrobacter CL28 normalized to root weight. To selectively count Arthrobacter CL28, CFUs were counted on Luria Bertani (LB) agar plates containing 50 μg ml−1 of apramycin, on which neither Variovorax B4 nor Variovorax CL14 grow. Arthrobacter CL28 CFUs are not reduced in the presence of Variovorax. Notably, Variovorax account for only about 1.5% of the root community (Fig. 2h). These results rule out the possibility that Variovorax enforces stereotypic root growth by antagonizing or outcompeting RGI inducers. Significance was determined via ANOVA, letters correspond to a Tukey post hoc test. n = 12 biologically independent samples.
Extended Data Fig. 7 Auxin-responsive genes are induced in response to RGI strains.
a, Box plots showing the average standardized expression of genes significantly induced only under RGI conditions (RGI-induced), across the following treatments. Left (tripartite system), uninoculated seedlings (NB) or seedlings inoculated with Variovorax CL14, Arthrobacter CL28 or both Arthrobacter CL28 and Variovorax CL14 (CL14/CL28). Right (drop-out system), uninoculated seedlings (NB), Burkholderia drop-out synthetic community (−Burk SynCom), Variovorax drop-out synthetic community (−Vario SynCom), Variovorax and Burkholdria drop-out synthetic community (−Vario −Burk SynCom) or the full synthetic community (full SynCom). RGI-induced genes are defined as genes that are significantly overexpressed in RGI treatments. Left, genes overexpressed in Arthrobacter CL28-inoculated seedling versus NB and in Arthrobacter CL28-inoculated seedlings versus CL14/CL28. Right, genes overexpressed in −Vario versus NB and in −Vario versus full. n = 3 (left); 5 (right). b, Venn diagram showing the overlap of enriched genes between the tripartite and drop-out systems. The heat map shows the pairwise correlation in expression of these 18 genes across tissues on the basis of the Klepikova Atlas25. Seventeen of the 18 genes show high correlation across the atlas, with the exception of the auxin-conjugating gene GH3.2. A root apex diagram from the Arabidopsis eFP Browser (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi)68 is shown, illustrating the spatial distribution of transcripts from the 17 highly correlated root apex-associated genes in the Klepikova Atlas25. Significance of the overlap of enriched genes was determined via hypergeometric test. c, RGI-related genes share gene ontologies. Network of statistically significant gene ontology terms contained within the 18 overlapping RGI-induced genes (a, b). The network was computed using the emapplot function from the package clusterProfiler in R. A P value for terms across the gene ontology was computed using a hypergeometric test (only significant ontologies are shown). Point size (gene ontology term) denotes the number of genes mapped to that particular term. d, Standardized expression of 12 late-responsive auxin genes26 across the tripartite and drop-out systems. Each dot represents a gene. Identical genes are connected between bacterial treatments with a black line. Mean expression (95% confidence intervals) of the aggregated 12 genes in each treatment is highlighted in red and connected between bacterial treatments with a red line. Significance was calculated using a resampling approach, in which we compared the calculated difference between means across groups. After 10,000 resamplings, we calculated the P value by comparing the distribution of means calculated against the real observed differences between means.
Extended Data Fig. 8 Variovorax degrades auxin and quenches auxin perception by the plant.
a, Primary root elongation of seedlings grown with six hormonal or microbial-associated molecular pattern RGI treatments (panels) individually (self) or with either Burkholderia CL11 or each of four Variovorax isolates (CL14, MF160, B4 and YR216). Significance was determined via ANOVA within each panel; letters correspond to a Tukey post hoc test. n = 74, 46, 61, 48, 49, 49, 45, 44, 46, 43, 49, 40, 22, 19, 22, 19, 20, 25, 28, 30, 29, 29, 29, 29, 12, 12, 21, 20, 19, 18, 26, 30, 30, 29, 30, 30, 29, 30, 30, 26, 29 and 28 biological replicates across 2 independent experiments. b, Variovorax degrades auxin. Growth curves showing optical density at OD600 (top) and IAA concentrations (mg ml−1) (bottom) in Variovorax CL14 cultures grown in M9 medium with different carbon sources. Left, IAA (+ 0.5% ethanol as solvent); middle, succinate; right, succinate and IAA (+ 0.5% ethanol as solvent). c, Variovorax quenches induction of the auxin bioreporter DR5::GFP. Quantification of GFP intensity in DR5::GFP Arabidopsis seedlings grown with no bacteria, Arthrobacter CL28 and Arthrobacter CL28 + Variovorax CL14. GFP fluorescence was imaged 1, 3, 6, 9 and 13 d after inoculation, and quantified in the root elongation zone. Significance was determined via ANOVA within each time point, while controlling for experiment, and denoted with asterisks. n = 20 biological replicates across 2 independent experiments. d, Representative primary root images of DR5::GFP plants quantified in c, showing roots from 1, 3 and 6 d after inoculation. In addition to the bacterial treatments shown in c, an exogenous IAA control is shown (IAA, second column), as well as IAA-treated plants inoculated with Variovorax CL14, illustrating that IAA-induced fluorescence is quenched in the presence of Variovorax CL14 within 3 d.
Extended Data Fig. 9 Detection of CL28-responsive Variovorax-unique operons.
a, Presence–absence matrix denoting the distribution of 12 Variovorax-unique hotspots containing at least 10 genes across the 185 members of the synthetic community. Hotspots are defined using the Variovorax CL14 genome as a reference. Phylogeny of the 185 members of the synthetic community is shown to the left of the matrix. We determined the presence of an orthogroup based on a hidden Markov model profile scanning of each core Variovorax (genus) orthogroup across the 185 genomes in the synthetic community. b, A map of the auxin-degrading hotspot 33. Genes are annotated with the last two digits of their IMG gene identifier (26436136XX) and their functional assignments are shown below the map, including per cent identity of any to genes from a known auxin degradation locus. Genes are coloured by the log2-transformed fold change in their transcript abundance in Variovorax CL14 cocultured with Arthrobacter CL28 versus Variovorax CL14 monoculture, as measured by RNA-seq (shown in c). The overlap of this region with vectors 1 and 2 and the region knocked out in Variovorax CL14 ΔHS33 are shown below the map. Vector 1 extends beyond this region. c, Results of RNA-seq on Variovorax CL14 transcripts. Variovorax CL14 was cocultured with Arthrobacter CL28 versus Variovorax CL14 monoculture. Only Variovorax-unique genomic hotspots are presented, aligned with the genes in a. Genes are coloured by the log2-transformed fold change in their transcript abundance in Variovorax CL14 cocultured with Arthrobacter CL28 vs Variovorax CL14 monoculture. Note uniform upregulation of genes in cluster 33. d, log-transformed CFUs of Variovorax CL14 or with the Acidovorax gain-of-function strain Acidovorax root219::V2 normalized to root weight. Each of these two strains was co-inoculated with Arthrobacter CL28 onto 7-d-old seedlings and collected after 12 d of growth. CFU counts were determined on LB plates containing 100 μg ml−1 ampicillin, for which Arthrobacter CL28 is susceptible and Variovorax CL14 and Acidovorax root219::V2 are naturally resistant. n = 3 biologically independent samples. Significance was determined using a two-sided Student’s t-test. P = 2.57 × 10−5. Line represents median. e, log-transformed CFUs of Variovorax CL14 or of the loss-of-function strain Variovorax CL14 ΔHS33 normalized to root weight. Each of these two strains was inoculated individually onto 7-d-old seedlings and collected after 12 d of growth. CFU counts were determined on LB plates. n = 5 biologically independent samples. Significance was determined using a two-sided Student’s t-test. P = 0.049 (mutant is slightly higher). Red lines represent median.
Extended Data Fig. 10 Variovorax are highly prevalent across naturally occurring Arabidopsis microbiomes and across 30 plant species.
a, Correlation plot of data reanalysed from ref. 6, comparing bacterial ASV prevalence to log-transformed relative abundance in A. thaliana rhizoplane samples taken across 3 years in 17 sites in Europe. b, Correlation plot of data reanalysed from ref. 4, comparing bacterial ASV prevalence to log-transformed relative abundance across 30 phylogenetically diverse plant species grown in a common garden experiment.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-6.
Source data
Rights and permissions
About this article
Cite this article
Finkel, O.M., Salas-González, I., Castrillo, G. et al. A single bacterial genus maintains root growth in a complex microbiome. Nature 587, 103–108 (2020). https://doi.org/10.1038/s41586-020-2778-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2778-7
This article is cited by
-
Leaf microbiome dysbiosis triggered by T2SS-dependent enzyme secretion from opportunistic Xanthomonas pathogens
Nature Microbiology (2024)
-
Endophytic Pseudomonas fluorescens promotes changes in the phenotype and secondary metabolite profile of Houttuynia cordata Thunb.
Scientific Reports (2024)
-
Grazing intensity changes root traits and resource utilization strategies of Stipa breviflora in a desert steppe
Plant and Soil (2024)
-
Rubber-based agroforestry systems modify the soil fungal composition and function in Southwest China
Soil Ecology Letters (2024)
-
Diversity of symbiotic cyanobacteria in cycad coralloid roots using a short-read rbcL-X amplicon
Symbiosis (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.