Abstract
Mitochondria require nicotinamide adenine dinucleotide (NAD+) to carry out the fundamental processes that fuel respiration and mediate cellular energy transduction. Mitochondrial NAD+ transporters have been identified in yeast and plants1,2, but their existence in mammals remains controversial3,4,5. Here we demonstrate that mammalian mitochondria can take up intact NAD+, and identify SLC25A51 (also known as MCART1)—an essential6,7 mitochondrial protein of previously unknown function—as a mammalian mitochondrial NAD+ transporter. Loss of SLC25A51 decreases mitochondrial—but not whole-cell—NAD+ content, impairs mitochondrial respiration, and blocks the uptake of NAD+ into isolated mitochondria. Conversely, overexpression of SLC25A51 or SLC25A52 (a nearly identical paralogue of SLC25A51) increases mitochondrial NAD+ levels and restores NAD+ uptake into yeast mitochondria lacking endogenous NAD+ transporters. Together, these findings identify SLC25A51 as a mammalian transporter capable of importing NAD+ into mitochondria.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its supplementary information files. Source data are provided with this paper.
Code availability
No custom codes were used during this study. Mathematical calculations are described in the materials and methods section or by cited works.
References
Palmieri, F. et al. Molecular identification and functional characterization of Arabidopsis thaliana mitochondrial and chloroplastic NAD+ carrier proteins. J. Biol. Chem. 284, 31249–31259 (2009).
Todisco, S., Agrimi, G., Castegna, A. & Palmieri, F. Identification of the mitochondrial NAD+ transporter in Saccharomyces cerevisiae. J. Biol. Chem. 281, 1524–1531 (2006).
Berger, F., Ramírez-Hernández, M. H. & Ziegler, M. The new life of a centenarian: signalling functions of NAD(P). Trends Biochem. Sci. 29, 111–118 (2004).
Yang, H. et al. Nutrient-sensitive mitochondrial NAD+ levels dictate cell survival. Cell 130, 1095–1107 (2007).
VanLinden, M. R. et al. Subcellular distribution of NAD+ between cytosol and mitochondria determines the metabolic profile of human cells. J. Biol. Chem. 290, 27644–27659 (2015).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Bertomeu, T. et al. A high-resolution genome-wide CRISPR/Cas9 viability screen reveals structural features and contextual diversity of the human cell-essential proteome. Mol. Cell. Biol. 38, e00302-17 (2017).
Yoshino, J., Baur, J. A. & Imai, S. I. NAD+ intermediates: the biology and therapeutic potential of NMN and NR. Cell Metab. 27, 513–528 (2018).
Cambronne, X. A. et al. Biosensor reveals multiple sources for mitochondrial NAD+. Science 352, 1474–1477 (2016).
Pittelli, M. et al. Inhibition of nicotinamide phosphoribosyltransferase: cellular bioenergetics reveals a mitochondrial insensitive NAD pool. J. Biol. Chem. 285, 34106–34114 (2010).
Sims, C. A. et al. Nicotinamide mononucleotide preserves mitochondrial function and increases survival in hemorrhagic shock. JCI Insight 3, 120182 (2018).
Titus, S. A. & Moran, R. G. Retrovirally mediated complementation of the glyB phenotype. Cloning of a human gene encoding the carrier for entry of folates into mitochondria. J. Biol. Chem. 275, 36811–36817 (2000).
Spaan, A. N. et al. Identification of the human mitochondrial FAD transporter and its potential role in multiple acyl-CoA dehydrogenase deficiency. Mol. Genet. Metab. 86, 441–447 (2005).
Berger, F., Lau, C., Dahlmann, M. & Ziegler, M. Subcellular compartmentation and differential catalytic properties of the three human nicotinamide mononucleotide adenylyltransferase isoforms. J. Biol. Chem. 280, 36334–36341 (2005).
Davila, A. et al. Nicotinamide adenine dinucleotide is transported into mammalian mitochondria. eLife 7, e33246 (2018).
Fletcher, R. S. et al. Nicotinamide riboside kinases display redundancy in mediating nicotinamide mononucleotide and nicotinamide riboside metabolism in skeletal muscle cells. Mol. Metab. 6, 819–832 (2017).
Nikiforov, A., Dölle, C., Niere, M. & Ziegler, M. Pathways and subcellular compartmentation of NAD biosynthesis in human cells: from entry of extracellular precursors to mitochondrial NAD generation. J. Biol. Chem. 286, 21767–21778 (2011).
Hikosaka, K. et al. Deficiency of nicotinamide mononucleotide adenylyltransferase 3 (nmnat3) causes hemolytic anemia by altering the glycolytic flow in mature erythrocytes. J. Biol. Chem. 289, 14796–14811 (2014).
Yamamoto, M. et al. Nmnat3 is dispensable in mitochondrial NAD level maintenance in vivo. PLoS ONE 11, e0147037 (2016).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Sallin, O. et al. Semisynthetic biosensors for mapping cellular concentrations of nicotinamide adenine dinucleotides. eLife 7, e32638 (2018).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Hart, T. et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell 163, 1515–1526 (2015).
Agerholm, M. et al. Perturbations of NAD+ salvage systems impact mitochondrial function and energy homeostasis in mouse myoblasts and intact skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 314, E377–E395 (2018).
Frederick, D. W. et al. Loss of NAD homeostasis leads to progressive and reversible degeneration of skeletal muscle. Cell Metab. 24, 269–282 (2016).
Rustin, P. et al. Fluxes of nicotinamide adenine dinucleotides through mitochondrial membranes in human cultured cells. J. Biol. Chem. 271, 14785–14790 (1996).
Agrimi, G. et al. Deletion or overexpression of mitochondrial NAD+ carriers in Saccharomyces cerevisiae alters cellular NAD and ATP contents and affects mitochondrial metabolism and the rate of glycolysis. Appl. Environ. Microbiol. 77, 2239–2246 (2011).
Floyd, B. J. et al. Mitochondrial protein interaction mapping identifies regulators of respiratory chain function. Mol. Cell 63, 621–632 (2016).
Dölle, C., Niere, M., Lohndal, E. & Ziegler, M. Visualization of subcellular NAD pools and intra-organellar protein localization by poly-ADP-ribose formation. Cell. Mol. Life Sci. 67, 433–443 (2010).
Trammell, S. A. & Brenner, C. Targeted, LCMS-based metabolomics for quantitative measurement of NAD+ metabolites. Comput. Struct. Biotechnol. J. 4, e201301012 (2013).
Zhang, Q., Piston, D. W. & Goodman, R. H. Regulation of corepressor function by nuclear NADH. Science 295, 1895–1897 (2002).
Zhao, Y. et al. SoNar, a highly responsive NAD+/NADH sensor, allows high-throughput metabolic screening of anti-tumor agents. Cell Metab. 21, 777–789 (2015).
Ho, B., Baryshnikova, A. & Brown, G. W. Unification of protein abundance datasets yields a quantitative Saccharomyces cerevisiae proteome. Cell Syst. 6, 192–205 (2018).
Malecki, M. J. et al. Leukemia-associated mutations within the NOTCH1 heterodimerization domain fall into at least two distinct mechanistic classes. Mol. Cell. Biol. 26, 4642–4651 (2006).
Eller, J. M. et al. Flow cytometry analysis of free intracellular NAD+ using a targeted biosensor. Curr. Protoc. Cytom. 88, e54 (2019).
Wang, L. et al. Peak annotation and verification engine for untargeted LC–MS metabolomics. Anal. Chem. 91, 1838–1846 (2019).
Adusumilli, R. & Mallick, P. Data conversion with ProteoWizard msConvert. Methods Mol. Biol. 1550, 339–368 (2017).
Su, X., Lu, W. & Rabinowitz, J. D. Metabolite spectral accuracy on orbitraps. Anal. Chem. 89, 5940–5948 (2017).
Chong, J., Wishart, D. S. & Xia, J. Using MetaboAnalyst 4.0 for comprehensive and integrative metabolomics data analysis. Curr. Protoc. Bioinformatics 68, e86 (2019).
Amberg, D. C., Burke, D. J. & Strathern, J. N. High-efficiency transformation of yeast. CSH Protoc 2006, pdb.prot4145 (2006).
Meisinger, C., Pfanner, N. & Truscott, K. N. Isolation of yeast mitochondria. Methods Mol. Biol. 313, 33–39 (2006).
Izawa, T. & Unger, A. K. Isolation of mitochondria from Saccharomyces cerevisiae. Methods Mol. Biol. 1567, 33–42 (2017).
Bricker, D. K. et al. A mitochondrial pyruvate carrier required for pyruvate uptake in yeast, Drosophila, and humans. Science 337, 96–100 (2012).
Acknowledgements
We thank all members of the Baur and Cambronne laboratories, V. Moiseenkova-Bell, A. Ellington, E. Marcotte, R. Goodman, I. Heiland, M. Whorton, E. Gouaux and J. Dixon for constructive discussions and suggestions; and M. Blair, Q. Chen, V. Annamalai, X. Yu, A. Slepian and CBRS UT Austin Proteomics Facility for technical support. This work was supported by grants from the National Institutes of Health (R01DK098656 to J.A.B., DP2GM126897 to X.A.C., TL1TR001880, T32AR53461 and F32HL145923 to T.S.L.) and the Norwegian Research Council (250395/F20 to M.Z.).
Author information
Authors and Affiliations
Contributions
T.S.L., X.A.C. and J.A.B. conceived and designed the overall study. T.S.L., J.M.E., M.-J.L., M.E.M., J.D.R., F.B.J., X.A.C. and J.A.B. contributed to the development of the hypotheses and experimental approaches. T.S.L., J.M.E., M.-J.L., M.N., F.R., M.R.M, C.P., M.R.B. and P.O. performed and analysed experiments. All authors contributed to the interpretation of experiments. T.S.L., X.A.C. and J.A.B. wrote the manuscript. J.M.E., M.-J.L., K.J. and M.Z. edited, and all authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.D.R. is a co-founder of Toran Therapeutics. The remaining authors declare no competing interests.
Additional information
Peer review information Nature thanks Ferdinando Palmieri and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 SLC25A51 is a mitochondrial protein that affects cellular NAD+ distribution, proliferation, and metabolome profiles.
a, b, qPCR quantification of SLC25A51 mRNA expression in HEK 293T (n = 3) (a) and HeLa cells (n = 3) (b) expressing shRNA targeting SLC25A51. c, d, NAD+ content of isolated mitochondria (n = 3) (c) and whole cell lysates (n = 3) (d) from HeLa cells with stable shRNA knockdown of SLC25A51 (KD) and non-targeting control (Ctrl). e, Western blot confirming shRNA targeting murine Slc25a51 reduces SLC25A51 protein expression in cells transfected with cDNA encoding SLC25A51-FLAG. f, Mitochondrial free NAD+ levels in mouse embryonic stem cells expressing shRNA against Slc25a51 and non-targeting shRNA (shFF2), as measured with the mitochondrial cpVenus NAD+ biosensor (n = 3). g, qPCR quantification of SLC25A51 mRNA expression in HeLa cells transfected with siRNA targeting SLC25A51 (n = 3). h, Western blot confirming protein expression of Flag-tagged mitochondrial carriers. Controls include stable expression of the NAD+ biosensor (sensor) and anti-Tubulin for loading. i, Immunofluorescent detection of SLC25A51 and SLC25A52 subcellular localization. Cells were transiently transfected with cDNA encoding Flag-HA-tagged SLC25A51 or SLC25A52 and probed with anti-Flag and the mitochondrial marker, anti-MTC02. Scale bar: 10 μM, 2 μM on inset. Inset represents zoomed view of Flag localization and mitochondria. j–l, Proliferation of HAP1 SLC25A51-KO (n = 8) (j), HEK 293T SLC25A51 shRNA-knockdown (n = 8) (k), HeLa SLC25A51 shRNA-knockdown cells (n = 8) (l) and their respective controls. Proliferation was measured by CyQuant, a fluorescent DNA dye, at 0h and 96h after plating and expressed as fold change. m–o, qPCR quantification of SLC25A52 mRNA expression in HAP1 SLC25A51-KO (m), HEK 293T SLC25A51 shRNA-knockdown (n) and HeLa SLC25A51 shRNA-knockdown cells (n = 3) (o). p, Western blot of whole cell protein lysates from HAP1 wild type (WT) and SLC25A51 knockout (KO) cells confirming SLC25A51 loss. Loading control is total protein measured by Revert 700 Total Protein. q, r, Heat map of top 30 mitochondrial (q) and whole cell metabolites (r) that differ between HAP1 wild type and SLC25A51-KO cells (n = 3). Data represented as mean ± SEM. P values were determined by unpaired, two-tailed Student’s t-test (for two groups) or one-way ANOVA with multiple comparisons analysis using Dunnett’s method (for groups of three or more). *P < 0.05, and ***P < 0.001 vs control or WT (exact P values are provided in the source data).
Extended Data Fig. 2 NAD+ and SLC25A51 affect oxidative phosphorylation.
a, Respiration of isolated mitochondria from HEK 293T cells treated with either vehicle (Veh) or the NAMPT inhibitor FK866 to deplete mitochondrial NAD+. Mitochondria were treated with pyruvate and malate (state 2), then ADP was added to induce state 3 respiration. 1 mM NAD+ was added to test the ability of exogenous NAD+ to rescue respiration in the setting of mitochondria NAD+ depletion (Trace is representative of n = 4 independent experiments). P values were determined by two-way ANOVA with multiple comparisons analysis using the Sidak method. b, Oxygen consumption rate (OCR) was measured in SLC25A51 shRNA knockdown (KD) and control (Ctrl) HeLa cells using a Seahorse XF96e. Basal OCR was measured before the addition of treatments and maximal respiration was measured after the sequential addition of oligomycin (Oligo, ATP synthase inhibitor) and FCCP (uncoupler). Rotenone (Rot) and Antimycin A (AA) were added as a control to completely block mitochondrial oxygen consumption (n = 6). c, Respiration of isolated mitochondria from SLC25A51 knockdown HEK 293T cells. Mitochondria were treated with pyruvate/malate (state 2), and then ADP was added to induce state 3 respiration. Oligomycin was added to block ATP synthase-mediated respiration (n = 3 independent experiments). d, Mitochondria were isolated from HEK 293T control, SLC25A51 shRNA knockdown cells, and controls treated with FK866 to deplete mitochondrial NAD+. Mitochondrial oxygen consumption rate was measured after treatment with pyruvate/malate (state 2), ADP (state 3), and 1 mM NAD+ (n = 4 independent experiments). e, f, Mean volume per mitochondrial unit (e) and number of distinct mitochondria per cell (f) quantified from confocal image reconstructions of mitochondrial voxels in SLC25A51 shRNA knockdown (n = 31 cells) and control (n = 32 cells) HeLa cells. Data represented as mean ± SEM. P values were determined by unpaired, two-tailed Student’s t-test. *P < 0.05, **P < 0.01, and ***P < 0.001 vs vehicle or control; ###P < 0.001 vs state 3 (exact P values are provided in the source data).
Extended Data Fig. 3 Intact NAD+, but not nicotinamide or nicotinamide mononucleotide contributes to the mitochondrial NAD+ pool.
a, Mitochondrial NAD+ content was measured in isolated mitochondria from HeLa control (Ctrl) cells, control cells treated with FK866 (Ctrl+FK), and SLC25A51 shRNA-knockdown (KD) cells. NAD+ content of isolated mitochondria was determined before (untreated) and after a 40-min incubation with 1 mM NAD+ (n = 3 independent experiments). b, NAD+ levels in HEK 293T mitochondria incubated with 1 mM nicotinamide (NAM), 1 mM nicotinamide mononucleotide (NMN), or 1 mM NAD+ (n = 3 independent experiments). c, NAD+ uptake in NAD+-depleted mitochondria isolated from HEK 293T cells incubated with NAD+ ± 2 mM NAM or 2mM NMN (n = 4 independent experiments). d, Fractional labelling of mitochondrial NAD+ in HAP1 cells treated with isotopically double labelled NaR (n = 3 biological independent replicates). Data represented as mean ± SEM. P values were determined by unpaired, two-tailed Student’s t-test (for two groups) or one-way ANOVA with multiple comparisons analysis using Dunnett’s or Tukey’s method (for groups of three or more). *P < 0.05 and ***P < 0.001 vs untreated, vehicle, and wild-type M+0; #P < 0.05 vs wild-type M+1.
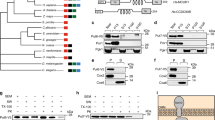
Extended Data Fig. 4 Generation and validation of yeast strains for testing mitochondrial NAD+ transport.
a, PCR genotyping to confirm double knockout gene deletion in BY4727 S. cerevisiae via antibiotic-resistance cassette replacement at the NDT1 and NDT2 loci. b, c, Deletion of the mitochondrial NAD+ carriers NDT1 and NDT2 in DKO strain phenocopied previously described growth defects on non-fermentative media (YP, 3% glycerol media)2, which was rescued by plasmid expression of NDT1. d, Western blot confirmed enrichment of mitochondrial markers (MTC02 and COXIV) and absence of cytoplasmic proteins (actin) or ER (SC2) in isolated mitochondria from yeast. e, RT-PCR confirmed ectopic expression from pRS415-SLC25A51 and pRS415-SLC25A52 in DKO strains.
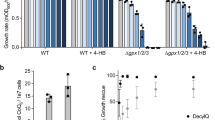
Extended Data Fig. 5 Kinetics and selectivity of NAD+ transport by human SLC25A51 expressed in yeast mitochondria.
a–c,, Co-incubation with excess unlabelled NAD+ (n = 5 independent experiments for 1 mM NAD+) (a), supraphysiological levels of NMN (100 μM, n = 4 independent experiments; 500 μM, n = 5 independent experiments) (b), NADH (n = 3 independent experiments) with 3H-NAD+ to measure uptake competition in mitochondria from DKO yeast expressing SLC25A51 (c). d, Proportional relationship between integrated peak intensities from mass spectrometry of mitochondrial samples compared to a known meta data set of absolute protein abundances; used to quantitate SLC25A51 abundance in yeast samples. e, Uptake measured with indicated NAD+ concentrations; calculated from specific activity (n = 3 independent experiments, mean ± SEM). f, Lineweaver-Burk plot based on a nonlinear fit with datapoints overlaid (n = 3 independent experiments). P values were determined by two-way ANOVA with multiple comparisons analysis using Sidak’s method. *P < 0.05 and **P < 0.01 vs 100 μM cold NAD+.
Supplementary information
Supplementary Figures
This file contains Supplementary Figure 1: the uncropped scan images of western blot membranes and DNA gels, and Supplementary Figure 2: the gating strategy for flow cytometry analysis.
Rights and permissions
About this article
Cite this article
Luongo, T.S., Eller, J.M., Lu, MJ. et al. SLC25A51 is a mammalian mitochondrial NAD+ transporter. Nature 588, 174–179 (2020). https://doi.org/10.1038/s41586-020-2741-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2741-7
This article is cited by
-
TREM2 macrophage promotes cardiac repair in myocardial infarction by reprogramming metabolism via SLC25A53
Cell Death & Differentiation (2024)
-
Visualizing subcellular changes in the NAD(H) pool size versus redox state using fluorescence lifetime imaging microscopy of NADH
Communications Biology (2024)
-
Mitochondrial proteome research: the road ahead
Nature Reviews Molecular Cell Biology (2024)
-
A CRISPRi/a screening platform to study cellular nutrient transport in diverse microenvironments
Nature Cell Biology (2024)
-
Long non-coding RNA SLC25A21-AS1 inhibits the development of epithelial ovarian cancer by specifically inducing PTBP3 degradation
Biomarker Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.