Abstract
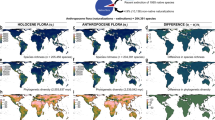
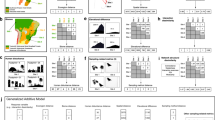
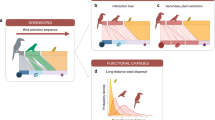
Introductions of species by humans are causing the homogenization of species composition across biogeographic barriers1,2,3. The ecological and evolutionary consequences of introduced species derive from their effects on networks of species interactions4,5, but we lack a quantitative understanding of the impacts of introduced species on ecological networks and their biogeographic patterns globally. Here we address this data gap by analysing mutualistic seed-dispersal interactions from 410 local networks, encompassing 24,455 unique pairwise interactions between 1,631 animal and 3,208 plant species. We show that species introductions reduce biogeographic compartmentalization of the global meta-network, in which nodes are species and links are interactions observed within any local network. This homogenizing effect extends across spatial scales, decreasing beta diversity among local networks and modularity within networks. The prevalence of introduced interactions is directly related to human environmental modifications and is accelerating, having increased sevenfold over the past 75 years. These dynamics alter the coevolutionary environments that mutualists experience6, and we find that introduced species disproportionately interact with other introduced species. These processes are likely to amplify biotic homogenization in future ecosystems7 and may reduce the resilience of ecosystems by allowing perturbations to propagate more quickly and exposing disparate ecosystems to similar drivers. Our results highlight the importance of managing the increasing homogenization of ecological complexity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data used in the analyses are available through the Dryad digital data repository (https://doi.org/10.5061/dryad.44j0zpcbx).
Code availability
All scripts that are needed to reproduce the analyses and figures are available through the Dryad digital data repository (https://doi.org/10.5061/dryad.44j0zpcbx).
References
Helmus, M. R., Mahler, D. L. & Losos, J. B. Island biogeography of the Anthropocene. Nature 513, 543–546 (2014).
Capinha, C., Essl, F., Seebens, H., Moser, D. & Pereira, H. M. Biogeography. The dispersal of alien species redefines biogeography in the Anthropocene. Science 348, 1248–1251 (2015).
Seebens, H. et al. No saturation in the accumulation of alien species worldwide. Nat. Commun. 8, 14435 (2017).
David, P. et al. Impacts of invasive species on food webs: a review of empirical data. Adv. Ecol. Res. 56, 1–60 (2017).
Traveset, A. & Richardson, D. M. Mutualistic interactions and biological invasions. Annu. Rev. Ecol. Evol. Syst. 45, 89–113 (2014).
Guimarães, P. R. Jr, Pires, M. M., Jordano, P., Bascompte, J. & Thompson, J. N. Indirect effects drive coevolution in mutualistic networks. Nature 550, 511–514 (2017).
McKinney, M. L. & Lockwood, J. L. Biotic homogenization: a few winners replacing many losers in the next mass extinction. Trends Ecol. Evol. 14, 450–453 (1999).
Wallace, A. R. The Geographical Distribution of Animals and Plants (Harper & Brothers, 1876).
Holt, B. G. et al. An update of Wallace’s zoogeographic regions of the world. Science 339, 74–78 (2013).
Vizentin-Bugoni, J. et al. Structure, spatial dynamics, and stability of novel seed dispersal mutualistic networks in Hawai’i. Science 364, 78–82 (2019).
Jordano, P. in Seeds: The Ecology of Regeneration in Plant Communities 2nd edn (ed. Fenner, M.) 18–61 (CABI, 2000).
Nathan, R. & Muller-Landau, H. C. Spatial patterns of seed dispersal, their determinants and consequences for recruitment. Trends Ecol. Evol. 15, 278–285 (2000).
Herrera, C. M. in Plant–Animal Interactions: An Evolutionary Approach (eds Herrera, C. M. & Pellmyr, O.) 185–208 (Blackwell, 2002).
Jordano, P. Patterns of mutualistic interactions in pollination and seed dispersal: connectance, dependence asymmetries, and coevolution. Am. Nat. 129, 657–677 (2002).
Tiffney, B. H. Vertebrate dispersal of seed plants through time. Annu. Rev. Ecol. Evol. Syst. 35, 1–29 (2004).
McConkey, K. R. et al. Seed dispersal in changing landscapes. Biol. Conserv. 146, 1–13 (2012).
Terborgh, J. et al. Tree recruitment in an empty forest. Ecology 89, 1757–1768 (2008).
Galetti, M. et al. Functional extinction of birds drives rapid evolutionary changes in seed size. Science 340, 1086–1090 (2013).
Rogers, H. S. et al. Effects of an invasive predator cascade to plants via mutualism disruption. Nat. Commun. 8, 14557 (2017).
Aslan, C. E., Zavaleta, E. S., Tershy, B. & Croll, D. Mutualism disruption threatens global plant biodiversity: a systematic review. PLoS One 8, e66993 (2013).
Schleuning, M. et al. Ecological networks are more sensitive to plant than to animal extinction under climate change. Nat. Commun. 7, 13965 (2016).
Thompson, J. The Geographic Mosaic of Coevolution (Univ. Chicago Press, 2005).
Watts, D. J. & Strogatz, S. H. Collective dynamics of ‘small-world’ networks. Nature 393, 440–442 (1998).
Medeiros, L. P., Garcia, G., Thompson, J. N. & Guimarães, P. R. Jr. The geographic mosaic of coevolution in mutualistic networks. Proc. Natl Acad. Sci. USA 115, 12017–12022 (2018).
Poisot, T., Canard, E., Mouillot, D., Mouquet, N. & Gravel, D. The dissimilarity of species interaction networks. Ecol. Lett. 15, 1353–1361 (2012).
Wang, S. & Loreau, M. Biodiversity and ecosystem stability across scales in metacommunities. Ecol. Lett. 19, 510–518 (2016).
Simberloff, D. & Von Holle, B. Positive interactions of nonindigenous species: invasional meltdown? Biol. Invasions 1, 21–32 (1999).
Tylianakis, J. M., Laliberté, E., Nielsen, A. & Bascompte, J. Conservation of species interaction networks. Biol. Conserv. (2010).
Kennedy, C. M., Oakleaf, J. R., Theobald, D. M., Baruch-Mordo, S. & Kiesecker, J. Managing the middle: a shift in conservation priorities based on the global human modification gradient. Glob. Chang. Biol. 25, 811–826 (2019).
Bastin, J.-F. et al. The global tree restoration potential. Science 365, 76–79 (2019).
Rezende, E. L., Lavabre, J. E., Guimarães, P. R., Jordano, P. & Bascompte, J. Non-random coextinctions in phylogenetically structured mutualistic networks. Nature 448, 925–928 (2007).
Mello, M. A. R. et al. The missing part of seed dispersal networks: structure and robustness of bat-fruit interactions. PLoS One 6, e17395 (2011).
Schleuning, M. et al. Ecological, historical and evolutionary determinants of modularity in weighted seed-dispersal networks. Ecol. Lett. 17, 454–463 (2014).
Chen, S. C. & Moles, A. T. A mammoth mouthful? A test of the idea that larger animals ingest larger seeds. Glob. Ecol. Biogeogr. 24, 1269–1280 (2015).
Dalsgaard, B. et al. Opposed latitudinal patterns of network-derived and dietary specialization in avian plant–frugivore interaction systems. Ecography 40, 1395–1401 (2016).
de Assis Bomfim, J., Guimarães, P. R., Peres, C. A., Carvalho, G. & Cazetta, E. Local extinctions of obligate frugivores and patch size reduction disrupt the structure of seed dispersal networks. Ecography 41, 1899–1909 (2018).
Castaño, J. H., Carranza, J. A. & Pérez-Torres, J. Diet and trophic structure in assemblages of montane frugivorous phyllostomid bats. Acta Oecol. 91, 81–90 (2018).
Escribano-Avila, G., Lara-Romero, C., Heleno, R. & Traveset, A. in Ecological Networks in the Tropics (eds Dáttilo, W. & Rico-Gray, V.) 93–110 (Springer, 2018).
Laurindo, R. S., Novaes, R. L. M., Vizentin-Bugoni, J. & Gregorin, R. The effects of habitat loss on bat-fruit networks. Biodivers. Conserv. 28, 589–601 (2019).
Dugger, P. J. et al. Seed-dispersal networks are more specialized in the Neotropics than in the Afrotropics. Glob. Ecol. Biogeogr. 28, 248–261 (2019).
Cayuela, L., Granzow-de la Cerda, Í., Albuquerque, F. S. & Golicher, D. J. Taxonstand: an R package for species names standardisation in vegetation databases. Methods Ecol. Evol. 3, 1078–1083 (2012).
Boyle, B. et al. The taxonomic name resolution service: an online tool for automated standardization of plant names. BMC Bioinformatics 14, 16 (2013).
Trøjelsgaard, K., Heleno, R. & Traveset, A. Native and alien flower visitors differ in partner fidelity and network integration. Ecol. Lett. 22, 1264–1273 (2019).
Aslan, C. E. Implications of non-native species for mutualistic network resistance and resilience. PLoS One 14, e0217498 (2019).
Araujo, A. C. et al. Spatial distance and climate determine modularity in a cross-biomes plant–hummingbird interaction network in Brazil. J. Biogeogr. 45, 1846–1858 (2018).
Emer, C. et al. Seed-dispersal interactions in fragmented landscapes—a metanetwork approach. Ecol. Lett. 21, 484–493 (2018).
Nnakenyi, C. A., Traveset, A., Heleno, R., Minoarivelo, H. O. & Hui, C. Fine-tuning the nested structure of pollination networks by adaptive interaction switching, biogeography and sampling effect in the Galápagos Islands. Oikos 128, 1413–1423 (2019).
Dunne, J. A. in Ecological Networks: Linking Structure to Dynamics in Food Webs (eds Pascual, M. & Dunne, J. A.) 27–86 (Oxford Univ. Press, 2006).
Briatte, F. ggnetwork: geometries to plot networks with ‘ggplot2’. R package v.0.5.1. (R Foundation for Statistical Computing, 2016).
Becker, R. A., Wilks, A. R., Brownrigg, R., Minka, T. P. & Deckmyn, A. maps: draw geographical maps. R package v.3 (R Foundation for Statistical Computing, 2013).
Clauset, A., Newman, M. E. J. & Moore, C. Finding community structure in very large networks. Phys. Rev. E. 70, 066111 (2004).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal 1695 (2006).
Dinerstein, E. et al. An ecoregion-based approach to protecting half the terrestrial realm. Bioscience 67, 534–545 (2017).
Warton, D. I., Duursma, R. A., Falster, D. S. & Taskinen, S. smatr 3—an R package for estimation and inference about allometric lines. Methods Ecol. Evol. 3, 257–259 (2012).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Dormann, C. F., Gruber, B. & Fründ, J. Introducing the bipartite package: analysing ecological networks. Interaction 1, 0.2413793 (2008).
Bascompte, J., Jordano, P., Melián, C. J. & Olesen, J. M. The nested assembly of plant-animal mutualistic networks. Proc. Natl Acad. Sci. USA 100, 9383–9387 (2003).
Marquitti, F. M. D., Guimarães, P. R., Pires, M. M. & Bittencourt, L. F. MODULAR: software for the autonomous computation of modularity in large network sets. Ecography 37, 221–224 (2014).
Newman, M. E. J. & Girvan, M. Finding and evaluating community structure in networks. Phys. Rev. E 69, 026113 (2004).
Geraci, M. Linear quantile mixed models: the lqmm package for Laplace quantile regression. J. Stat. Softw. 57, 1–29 (2014).
Acknowledgements
We thank the many researchers that collected the field data that were used in this analysis. E.C.F. was supported by the National Socio-Environmental Synthesis Center (SESYNC) under funding received from the National Science Foundation DBI-1639145. J.C.S. considers this work a contribution to his VILLUM Investigator project “Biodiversity Dynamics in a Changing World” funded by VILLUM FONDEN (grant 16549).
Author information
Authors and Affiliations
Contributions
E.C.F. conceived the study with J.C.S., assembled and analysed data, and wrote the first draft. Both authors designed analyses and revised the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Mauro Galetti, Thilo Gross and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Biogeographic compartmentalization measured via meta-network modularity.
Modularity is calculated given species’ module membership based on the regions in which they are native. High modularity values indicate high compartmentalization along biogeographic regions, with distributions of bootstrapped values shown. Blue distribution shows modularity of the native-only meta-network. Red distribution shows modularity of the observed meta-network including introduced interactions. Red dashed line indicates distribution of modularity values for observed meta-networks sampled to have an equivalent number of interactions as in the native-only meta-network. Because this gives similar modularity values as in the observed meta-network, the higher modularity of the native-only meta-network is not explained by a lower number of interactions. Grey dashed line indicates distribution of modularity values for meta-networks simulated if species were homogenized randomly within biomes; this reflects a null expectation for biogeographic compartmentalization given a lack of dispersal barriers.
Extended Data Fig. 2 Beta diversity in species and interactions among local networks.
Relationship between pairwise distances and similarity of species in the community (1 – βS) and interactions (1 – βWN). Points represent 14,849 pairs of local networks and lines represent model fits from generalized additive models. Top row points in blue (a, b) are calculated using only native interactions and bottom row points in red (c, d) are calculated including introduced interactions.
Extended Data Fig. 3 Interaction breadth of species that have established as introduced species versus species that do not appear as introduced species.
For focal plant and animal species, the normalized degree—number of observed partners divided by the number of potential partners—was greater for species that appear as introduced than for species that do not. In a, b, introduced species are those that have been reported as introduced anywhere according to publicly available databases (χ2a = 59.7, d.f. = 1, Pa = 1.1 × 10−14, χ2p = 14.5, d.f. = 1, Pp = 1.4 × 10−4). In c, d, introduced species are those that are recorded in our database within a local network where they are introduced (χ2a = 235.3, d.f. = 1, Pa < 2.2 × 10−16, χ2p = 40.4, d.f. = 1, Pp = 2.1 × 10−10).
Extended Data Fig. 4 Introduced interactions reduce modularity of local networks.
Model estimates from mixed effects quantile regression, using random intercepts by study location. Points (jittered along x axis) show null model-corrected modularity calculated for 395 local networks as the difference between observed modularity and the average modularity of five corresponding networks generated under a null model. Coefficient estimates (±s.e.m.) at the 5th, 50th and 95th percentiles are, for the intercept, β0,5 = –0.050 (±0.005), β0,50 = 0.012 (±0.004), β0,95 = 0.130 (±0.019), and, for the effect of the proportion of introduced interactions, β1,5 = –0.009 (±0.020), β1,50 = –0.029 (±0.016), β1,95 = –0.079 (±0.037).
Supplementary information
Rights and permissions
About this article
Cite this article
Fricke, E.C., Svenning, JC. Accelerating homogenization of the global plant–frugivore meta-network. Nature 585, 74–78 (2020). https://doi.org/10.1038/s41586-020-2640-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2640-y
This article is cited by
-
Non-native ants are breaking down biogeographic boundaries and homogenizing community assemblages
Nature Communications (2024)
-
A Review on the State of the Art in Frugivory and Seed Dispersal on Islands and the Implications of Global Change
The Botanical Review (2024)
-
Disentangling the relationships among abundance, invasiveness and invasibility in trait space
npj Biodiversity (2023)
-
Common seed dispersers contribute most to the persistence of a fleshy-fruited tree
Communications Biology (2023)
-
Indirect effects shape species fitness in coevolved mutualistic networks
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.