Abstract
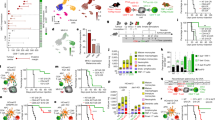
Conventional type 1 dendritic cells (cDC1)1 are thought to perform antigen cross-presentation, which is required to prime CD8+ T cells2,3, whereas cDC2 are specialized for priming CD4+ T cells4,5. CD4+ T cells are also considered to help CD8+ T cell responses through a variety of mechanisms6,7,8,9,10,11, including a process whereby CD4+ T cells ‘license’ cDC1 for CD8+ T cell priming12. However, this model has not been directly tested in vivo or in the setting of help-dependent tumour rejection. Here we generated an Xcr1Cre mouse strain to evaluate the cellular interactions that mediate tumour rejection in a model requiring CD4+ and CD8+ T cells. As expected, tumour rejection required cDC1 and CD8+ T cell priming required the expression of major histocompatibility class I molecules by cDC1. Unexpectedly, early priming of CD4+ T cells against tumour-derived antigens also required cDC1, and this was not simply because they transport antigens to lymph nodes for processing by cDC2, as selective deletion of major histocompatibility class II molecules in cDC1 also prevented early CD4+ T cell priming. Furthermore, deletion of either major histocompatibility class II or CD40 in cDC1 impaired tumour rejection, consistent with a role for cognate CD4+ T cell interactions and CD40 signalling in cDC1 licensing. Finally, CD40 signalling in cDC1 was critical not only for CD8+ T cell priming, but also for initial CD4+ T cell activation. Thus, in the setting of tumour-derived antigens, cDC1 function as an autonomous platform capable of antigen processing and priming for both CD4+ and CD8+ T cells and of the direct orchestration of their cross-talk that is required for optimal anti-tumour immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The microarray data generated during the course of this study have been deposited and are available at the Gene Expression Omnibus (GEO) database. The microarrays used in Fig. 4b and Extended Data Fig. 7b can be accessed with the accession number GSE152196. All other primary data and materials that support the findings of this study are available from the corresponding author upon request. Source data are provided with this paper.
References
Guilliams, M. et al. Dendritic cells, monocytes and macrophages: a unified nomenclature based on ontogeny. Nat. Rev. Immunol. 14, 571–578 (2014).
Hildner, K. et al. Batf3 deficiency reveals a critical role for CD8α+ dendritic cells in cytotoxic T cell immunity. Science 322, 1097–1100 (2008).
Cancel, J. C., Crozat, K., Dalod, M. & Mattiuz, R. Are conventional type 1 dendritic cells critical for protective antitumor immunity and how? Front. Immunol. 10, 9 (2019).
Dudziak, D. et al. Differential antigen processing by dendritic cell subsets in vivo. Science 315, 107–111 (2007).
Binnewies, M. et al. Unleashing type-2 dendritic cells to drive protective antitumor CD4+ T cell immunity. Cell 177, 556–571 (2019).
Bennett, S. R., Carbone, F. R., Karamalis, F., Miller, J. F. & Heath, W. R. Induction of a CD8+ cytotoxic T lymphocyte response by cross-priming requires cognate CD4+ T cell help. J. Exp. Med. 186, 65–70 (1997).
Bennett, S. R. et al. Help for cytotoxic-T-cell responses is mediated by CD40 signalling. Nature 393, 478–480 (1998).
Schoenberger, S. P., Toes, R. E., van der Voort, E. I., Offringa, R. & Melief, C. J. T-cell help for cytotoxic T lymphocytes is mediated by CD40-CD40L interactions. Nature 393, 480–483 (1998).
Bourgeois, C., Rocha, B. & Tanchot, C. A role for CD40 expression on CD8+ T cells in the generation of CD8+ T cell memory. Science 297, 2060–2063 (2002).
Janssen, E. M. et al. CD4+ T-cell help controls CD8+ T-cell memory via TRAIL-mediated activation-induced cell death. Nature 434, 88–93 (2005).
Williams, M. A., Tyznik, A. J. & Bevan, M. J. Interleukin-2 signals during priming are required for secondary expansion of CD8+ memory T cells. Nature 441, 890–893 (2006).
Borst, J., Ahrends, T., Bąbała, N., Melief, C. J. M. & Kastenmüller, W. CD4+ T cell help in cancer immunology and immunotherapy. Nat. Rev. Immunol. 18, 635–647 (2018).
Ridge, J. P., Di Rosa, F. & Matzinger, P. A conditioned dendritic cell can be a temporal bridge between a CD4+ T-helper and a T-killer cell. Nature 393, 474–478 (1998).
Smith, C. M. et al. Cognate CD4+ T cell licensing of dendritic cells in CD8+ T cell immunity. Nat. Immunol. 5, 1143–1148 (2004).
Eickhoff, S. et al. Robust anti-viral immunity requires multiple distinct T Cell-dendritic cell interactions. Cell 162, 1322–1337 (2015).
Hor, J. L. et al. Spatiotemporally distinct interactions with dendritic cell subsets facilitates CD4+ and CD8+ T cell activation to localized viral infection. Immunity 43, 554–565 (2015).
Lehmann, C. H. K. et al. DC subset-specific induction of T cell responses upon antigen uptake via Fcγ receptors in vivo. J. Exp. Med. 214, 1509–1528 (2017).
Bajaña, S., Roach, K., Turner, S., Paul, J. & Kovats, S. IRF4 promotes cutaneous dendritic cell migration to lymph nodes during homeostasis and inflammation. J. Immunol. 189, 3368–3377 (2012).
Schlitzer, A. et al. IRF4 transcription factor-dependent CD11b+ dendritic cells in human and mouse control mucosal IL-17 cytokine responses. Immunity 38, 970–983 (2013).
Williams, J. W. et al. Transcription factor IRF4 drives dendritic cells to promote Th2 differentiation. Nat. Commun. 4, 2990 (2013).
Valdez, Y. et al. Major histocompatibility complex class II presentation of cell-associated antigen is mediated by CD8α+ dendritic cells in vivo. J. Exp. Med. 195, 683–694 (2002).
Theisen, D. J. et al. WDFY4 is required for cross-presentation in response to viral and tumor antigens. Science 362, 694–699 (2018).
Sun, J. C. & Bevan, M. J. Defective CD8 T cell memory following acute infection without CD4 T cell help. Science 300, 339–342 (2003).
Sun, J. C., Williams, M. A. & Bevan, M. J. CD4+ T cells are required for the maintenance, not programming, of memory CD8+ T cells after acute infection. Nat. Immunol. 5, 927–933 (2004).
Matsushita, H. et al. Cancer exome analysis reveals a T-cell-dependent mechanism of cancer immunoediting. Nature 482, 400–404 (2012).
Alspach, E. et al. MHC-II neoantigens shape tumour immunity and response to immunotherapy. Nature 574, 696–701 (2019).
Bos, R. & Sherman, L. A. CD4+ T-cell help in the tumor milieu is required for recruitment and cytolytic function of CD8+ T lymphocytes. Cancer Res. 70, 8368–8377 (2010).
Ferris, S. T. et al. A minor subset of Batf3-dependent antigen-presenting cells in islets of Langerhans is essential for the development of autoimmune diabetes. Immunity 41, 657–669 (2014).
Theisen, D. J. et al. Batf3-dependent genes control tumor rejection induced by dendritic cells independently of cross-presentation. Cancer Immunol. Res. 7, 29–39 (2019).
Durai, V. et al. Cryptic activation of an Irf8 enhancer governs cDC1 fate specification. Nat. Immunol. 20, 1161–1173 (2019).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Bern, M. D. et al. Inducible down-regulation of MHC class I results in natural killer cell tolerance. J. Exp. Med. 216, 99–116 (2019).
Hashimoto, K., Joshi, S. K. & Koni, P. A. A conditional null allele of the major histocompatibility IA-beta chain gene. Genesis 32, 152–153 (2002).
Archambault, A. S. et al. Cutting edge: conditional MHC class II expression reveals a limited role for B cell antigen presentation in primary and secondary CD4 T cell responses. J. Immunol. 191, 545–550 (2013).
Laidlaw, B. J., Craft, J. E. & Kaech, S. M. The multifaceted role of CD4+ T cells in CD8+ T cell memory. Nat. Rev. Immunol. 16, 102–111 (2016).
Ardouin, L. et al. Broad and largely concordant molecular changes characterize tolerogenic and immunogenic dendritic cell maturation in thymus and periphery. Immunity 45, 305–318 (2016).
Pinschewer, D. D. et al. FTY720 immunosuppression impairs effector T cell peripheral homing without affecting induction, expansion, and memory. J. Immunol. 164, 5761–5770 (2000).
Ahrends, T. et al. CD4+ T cell help confers a cytotoxic T cell effector program including coinhibitory receptor downregulation and increased tissue invasiveness. Immunity 47, 848–861 (2017).
Kamphorst, A. O., Guermonprez, P., Dudziak, D. & Nussenzweig, M. C. Route of antigen uptake differentially impacts presentation by dendritic cells and activated monocytes. J. Immunol. 185, 3426–3435 (2010).
Huysamen, C., Willment, J. A., Dennehy, K. M. & Brown, G. D. CLEC9A is a novel activation C-type lectin-like receptor expressed on BDCA3+ dendritic cells and a subset of monocytes. J. Biol. Chem. 283, 16693–16701 (2008).
Zhang, J. G. et al. The dendritic cell receptor Clec9A binds damaged cells via exposed actin filaments. Immunity 36, 646–657 (2012).
Lesley, R., Kelly, L. M., Xu, Y. & Cyster, J. G. Naive CD4 T cells constitutively express CD40L and augment autoreactive B cell survival. Proc. Natl Acad. Sci. USA 103, 10717–10722 (2006).
Pasqual, G. et al. Monitoring T cell-dendritic cell interactions in vivo by intercellular enzymatic labelling. Nature 553, 496–500 (2018).
Satpathy, A. T. et al. Zbtb46 expression distinguishes classical dendritic cells and their committed progenitors from other immune lineages. J. Exp. Med. 209, 1135–1152 (2012).
Kohyama, M. et al. Role for Spi-C in the development of red pulp macrophages and splenic iron homeostasis. Nature 457, 318–321 (2009).
Wu, X. et al. Mafb lineage tracing to distinguish macrophages from other immune lineages reveals dual identity of Langerhans cells. J. Exp. Med. 213, 2553–2565 (2016).
Lybarger, L., Wang, X., Harris, M. R., Virgin, H. W., IV & Hansen, T. H. Virus subversion of the MHC class I peptide-loading complex. Immunity 18, 121–130 (2003).
Sedy, J. R. et al. B and T lymphocyte attenuator regulates T cell activation through interaction with herpesvirus entry mediator. Nat. Immunol. 6, 90–98 (2005).
Toebes, M. et al. Design and use of conditional MHC class I ligands. Nat. Med. 12, 246–251 (2006).
Andersen, R. S. et al. Parallel detection of antigen-specific T cell responses by combinatorial encoding of MHC multimers. Nat. Protocols 7, 891–902 (2012).
Acknowledgements
K.M.M. was supported by the Howard Hughes Medical Institute and the US National Institutes of Health (R01AI150297); R.D.S. by grants from the National Institutes of Health (R01CA190700) and The Parker Institute for Cancer Immunotherapy, and a Stand Up To Cancer–Lustgarden Foundation Pancreatic Cancer Foundation Convergence Team Translational Research Grant; W.M.Y. by a grant from the National Institutes of Health (R01AI129545); M.D.B. and V.D. by fellowship grants from the National Institutes of Health (F30DK112466 and F30DK108498, respectively); S.T.F. by a postdoctoral training grant from the National Institutes of Health (T32CA95473); G.F.W., D.J.T. and J.T.D. by the National Institutes of Health (R01NS106289, T32 AI007163 and T32CA009621, respectively); and P.B. by the US National Science Foundation (DGE-1143954). We thank the Genome Technology Access Center, Department of Genetics, Washington University School of Medicine in St Louis, for help with genomic analysis. The Center is supported by Cancer Center Support Grant P30 CA91842 from the US National Cancer Institute and by Institute of Clinical and Translational Sciences/Clinical and Translational Science Award UL1 TR000448 from the US National Center for Research Resources. Aspects of studies including tetramer production were performed with assistance by the Immunomonitoring Laboratory, which is supported by the Andrew M. and Jane M. Bursky Center for Human Immunology and Immunotherapy Programs and the Alvin J. Siteman Comprehensive Cancer Center that, in turn, is supported by the National Cancer Institute of the National Institutes of Health Cancer Center Support Grant (P30CA91842) and the Washington University Rheumatic Diseases Research Resource-based Center Grant (P30AR073752).
Author information
Authors and Affiliations
Contributions
S.T.F., R.W. and V.D. conceived and designed the experiments, collected the data, and performed and interpreted the analyses. S.T.F. and R.W. wrote the manuscript. J.T.D., P.B., D.J.T., T.L., L.L. and C.G.B. collected and analysed data. J.P.W. and D.J.T. helped generate the 1956-mOVA fibrosarcoma cell line. M.D.B. provided the β2mfl/fl mouse and interpreted the analyses. G.F.W., W.M.Y., W.E.G., T.L.M. and R.D.S. provided assistance with experimental design. K.M.M. conceived experiments, interpreted data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.D.S. is a cofounder, scientific advisory board member, stockholder and royalty recipient of Jounce Therapeutics and Neon Therapeutics and is a scientific advisory board member for A2 Biotherapeutics, BioLegend, Codiak Biosciences, Constellation Pharmaceuticals, NGM Biopharmaceuticals and Sensei Biotherapeutics.
Additional information
Peer review information Nature thanks Cornelis Melief, Sebastian Amigorena and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 cDC1 are required to prime CD4+ T cells during the tumour immune response.
a, 1956-EV and 1956-mOVA were stained with antibodies against (left) Thy1.1 and (right) OVA. b, Tumour growth curves of B6 wild-type (WT) mice injected with 106 (left)1956-EV or (right)1956-mOVA. c, Tumour growth curves of (left) B6 WT or (right) Irf8 +32–/– mice injected with 106 1956-mOVA. d, B6 WT or Irf8 +32–/– mice were subcutaneously injected with 106 1956-EV or 106 1956-mOVA. Spleens were isolated and stained for presence of SIINFEKL-Kb-tetramer+ CD8+ T cells. (Left) Representative flow plots of percentage of tetramer+ CD8+ T cells. (Right) Graph of tetramer+ CD8+ T cells as a percentage of all T cells. Data are pooled biologically independent samples from two independent experiments (n = 3 for WT EV, n = 4 for WT 1956-mOVA, n = 5 for Irf8 +32–/– 1956-mOVA).
Extended Data Fig. 2 cDC1 are required to prime CD4+ T cells during the immune response to B16F10 melanoma.
a, B16F10-EV and B16F10-mOVA were stained with antibodies against (left) Thy1.1 and (right) OVA. b, B6 WT or Irf8 +32–/– mice were subcutaneously injected with 106 B16F10-EV or 106 B16F10-mOVA. (Left) Representative flow plots of OT-II T cells 3 days after transfer. (Right) Graph of per cent proliferated OT-II transferred. Data are pooled biologically independent samples from two independent experiments (n = 4 for WT B16F10-EV, n = 6 for all other groups).*P = 0.04 (unpaired, two-tailed Mann–Whitney test).
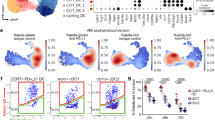
Extended Data Fig. 3 Validation of mCherry expression and lineage tracing of Xcr1Cre mouse.
a, Schematic diagrams of the mouse Xcr1 WT allele, the targeting vector (IRES-mCherry-T2a-hCRE with FRT flanked pGK-Neo cassettes), and the targeted allele. Filled and open boxes denote coding and noncoding exons of Xcr1, respectively. b, Southern blot analysis of Xcr1+/+ and Xcr1Cre/+. Genomic DNAs were isolated from mice tails, digested with SalI, electrophoresed, and hybridized with the 5′-radiolabelled probe indicated in a. Southern blot gave a 10.6 and a 6.9 kbp band for WT and targeted allele, respectively. For Southern blot source data, see Supplementary Fig. 1. c, Southern blot analysis of Xcr1+/+ and Xcr1Cre/+. Genomic DNAs were isolated from mice tails, digested with SalI, electrophoresed, and hybridized with the 3′ radiolabelled probe indicated in a. Southern blot gave a 10.6 and a 8.2 kbp band for WT and targeted allele, respectively. For Southern blot source data, see Supplementary Fig. 1. d, Gating strategy to delineate splenic cell populations. e, FACs histograms of mCherry expression from subpopulations in d isolated from Xcr1Cre/+ and Xcr1+/+ mice. f, Graph of mCherry expression in antigen-presenting cell populations gated from d isolated from Xcr1Cre/+ and Xcr1+/+ mice. Data are pooled biologically independent samples from two independent experiments (n = 5 in all groups). g, FACs histograms for YFP expression from subpopulations in d isolated from Xcr1Cre/+ R26LSLYFP/+ and Xcr1+/+ R26LSLYFP/+ mice. h, Graph of YFP expression in splenic cell populations (Mo, monocytes; N, neutrophils; RPM, red pulp macrophages). Data are pooled biologically independent samples from two independent experiments (n = 6 for cDC1, cDC2, plasmacytoid dendritic cells (pDC) and B cells from Xcr1+/+ R26LSLYFP/+ mice, n = 7 for cDC1, cDC2, pDC and B cells from Xcr1Cre/+ R26LSLYFP/+ mice, and n = 4 for all other groups). i, (Top) Gating strategy to delineate SDLN cell populations. (Bottom) FACs histograms for YFP expression in cDC1 and cDC2 isolated from Xcr1Cre/+ R26LSLYFP/+ and Xcr1+/+ R26LSLYFP/+ mice. j, Graph of YFP expression from subpopulations in i isolated from Xcr1Cre/+ R26LSLYFP/+ and Xcr1+/+ R26LSLYFP/+ mice. Data are pooled biologically independent samples from two independent experiments (n = 3 in all groups).
Extended Data Fig. 4 Proliferation of OT-I in Xcr1Cre/+ β2mfl/fl mice receiving soluble or cell-associated OVA.
a, Representative FACs analysis and histograms of CFSE dilution of proliferated OT-I on day 3 after transfer into (left) Xcr1+/+ β2mfl/fl and (right) Xcr1Cre/+ β2mfl/fl immunized with soluble OVA. b, Graph of per cent proliferation of transferred OT-I in mice immunized with soluble OVA. Data are pooled biologically independent samples from two independent experiments (n = 5 for all groups). c, Representative FACs analysis and histograms of CFSE dilution of proliferated OT-I on day 3 after transfer into (left) Xcr1+/+ β2mfl/fl and (right) Xcr1Cre/+ β2mfl/fl immunized with cell-associated OVA. d, Graph of per cent proliferation of transferred OT-I in mice immunized with cell-associated OVA. Data are pooled biologically independent samples from two independent experiments (n = 5 Xcr1+/+ β2mfl/fl –OVA, n = 6 for Xcr1Cre/+ β2mfl/fl +OVA, n = 7 for Xcr1+/+ β2mfl/fl +OVA, and n = 8 for Xcr1Cre/+ β2mfl/fl +OVA).
Extended Data Fig. 5 Proliferation of OT-II in Xcr1Cre /+ MHCIIfl/fl and Xcr1Cre /+ MHCIILSL/– mice immunized with soluble and cell-associated OVA.
a, Representative FACS analysis of splenic CD4+ T cell percentage in WT B6, Xcr1+/+ MHCIILSL/–, and Xcr1Cre/+ MHCIILSL/– mice at steady state. b, (Left) Representative FACS analysis of splenic Treg percentage in Xcr1+/+ MHCIIfl/fl and Xcr1Cre/+ MHCIIfl/fl at steady state. (Right) Graph of splenic Treg percentage as a percentage of all CD4+ T cells. Data are pooled biologically independent samples from two independent experiments (n = 5 for Xcr1+/+ MHCIIfl/fl, n = 4 for Xcr1Cre/+ MHCIIfl/fl). c, Representative FACs analysis and histograms of CFSE dilution of proliferated OT-II on day 3 after transfer into Xcr1+/+ MHCIIfl/fl, Xcr1Cre/+ MHCIIfl/fl, and Xcr1Cre/+ MHCIILSL/– immunized with soluble OVA. d, Representative FACs analysis and histograms of CFSE dilution of proliferated OT-II on day 3 after transfer into Xcr1+/+ MHCIIfl/fl, Xcr1Cre/+ MHCIIfl/fl, and Xcr1Cre/+ MHCIILSL/– immunized with cell-associated OVA.
Extended Data Fig. 6 Analysis of cDC1 in conditionally deleted mice.
a, Graph of splenic cDC1 percentage in Xcr1+/+ β2mfl/fl and Xcr1Cre/+ β2mfl/fl. Data are pooled biologically independent samples from two independent experiments (n = 5 for all groups). P = NS (unpaired, two-tailed Mann–Whitney test). b, Graph of splenic cDC1 percentage in Xcr1+/+ MHCIIfl/fl and Xcr1Cre/+ MHCIIfl/fl. Data are pooled biologically independent samples from two independent experiments (n = 5 for all groups). P = NS (unpaired, two-tailed Mann–Whitney test). c, Graph of absolute numbers of transferred OT-I in soluble OVA treated Xcr1+/+ β2mfl/fl and Xcr1Cre/+ β2mfl/fl mice. Data are pooled biologically independent samples from two independent experiments (n = 5 for all groups). d, Graph of absolute numbers of transferred OT-II in soluble OVA treated Xcr1+/+ MHCIIfl/fl and Xcr1Cre/+ MHCIIfl/fl mice. Data are pooled biologically independent samples from two independent experiments (n = 5 for all groups). e, Graph of per cent proliferated OT-I in cell-associated treated Xcr1+/+ MHCIIfl/fl and Xcr1Cre/+ MHCIIfl/fl mice. Data are pooled biologically independent samples from two independent experiments (n = 6 for Xcr1Cre/+ MHCIIfl/fl OVA– and OVA+ and n = 5 for all other groups). f, Graph of per cent proliferation of OT-I after 72 h coculture with ex vivo migratory cDC2 or cDC1 collected from tumour-draining lymph nodes of Xcr1+/+ MHCIIfl/fl or Xcr1Cre/+ MHCIIfl/fl mice injected 6 days earlier with 106 1956-mOVA cells. Cells were cultured at a ratio of 10:1 naive OT-I:cDC. Data are pooled independent samples from two independent experiments (n = 4 for all groups). g, Graph of absolute number of proliferated OT-I per well after 72 h coculture with ex vivo migratory cDC2 or cDC1 collected from tumour-draining lymph nodes of Xcr1+/+ MHCIIfl/fl, Xcr1Cre/+ MHCIIfl/fl mice six days after injection with 106 1956-mOVA. Cells were cultured at 10:1 ratio of naive OT-I:cDC. Data are pooled independent samples from two independent experiments (n = 4 for all groups).
Extended Data Fig. 7 CD40 deficiency does not affect cDC1 development.
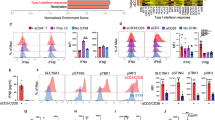
a, SDLN flow cytometry gating for cDC1 expression of CD40. Migratory cDC1 (CD11cint MHCIIhi; Red) were overlaid for expression with resident cDC1 (CD11chi MHCIIint; Blue). (Top) Xcr1+/+ CD40fl/fl and (bottom) Xcr1Cre/+ CD40fl/fl SDLN and splenic antigen-presenting cells stained for CD40 expression. b, Gene-expression data from Xcr1+/+ CD40fl/fl and Xcr1Cre/+ CD40fl/fl cDC1 from spleens and SDLN. Green lines indicate 2-fold changes. c, Graph of per cent proliferation of OT-I after 72 h coculture with ex vivo migratory cDC2 or cDC1 collected from tumour-draining lymph nodes of Xcr1+/+ CD40fl/fl or Xcr1Cre/+ CD40fl/fl mice injected 6 days earlier with 106 1956-mOVA cells. Cells were cultured at a ratio of 10:1 naive OT-I:cDC. Data are pooled independent samples from two independent experiments (n = 3 for Xcr1+/+ CD40fl/fl cDC2 and n = 4 for all other groups). d, Graph of per cent proliferation of OT-I in tumour-draining lymph node of tumour-bearing mice 3 days after transfer. Xcr1+/+ CD40fl/fl and Xcr1Cre/+ CD40fl/fl mice were injected with 106 1956-EV or 106 1956-mOVA. Data are pooled independent samples from two independent experiments (n = 5 for Xcr1Cre/+ CD40fl/fl1956-mOVA, n = 3 for all other groups). e, Graph of per cent proliferation of transferred OT-II in soluble OVA treated Xcr1+/+ CD40fl/fl and Xcr1Cre/+ CD40fl/fl mice. Data are pooled biologically independent samples from two independent experiments (n = 2 for 0 mg Xcr1+/+ CD40fl/fl and Xcr1Cre/+ CD40fl/fl and n = 4 for all other groups). f, Graph of per cent proliferation of OT-II in tumour-draining lymph node of tumour-bearing mice 3 days after transfer. Xcr1+/+ CD40fl/fl and Xcr1Cre/+ CD40fl/fl mice were injected with 106 1956-EV or 106 1956-mOVA. Data are pooled biologically independent samples from three independent experiments (n = 4 for 1956-EV, n = 6 for 1956-mOVA Xcr1+/+ CD40fl/fl, and n = 9 for 1956-mOVA Xcr1Cre/+ CD40fl/fl). g, Gating strategy to delineate day 6 1956-mOVA tumour immune cell antigen-presenting cell populations. h, FACs histogram of CD40 expression on gated antigen-presenting cell populations from e.
Extended Data Fig. 8 T cells are required at tumour site to induce memory.
a, Schematic of FTY720 injection during primary tumour response. b, Schematic of FTY720 injection during secondary tumour response. c, Peripheral blood CD4+ and CD8+ T cell percentage in control and FTY720 treated mice. Data represent mean ± s.d. pooled biologically independent samples from two independent experiments (n = 2 control, n = 5 FTY720). d, Tumour growth curves of mice injected with FTY720 during the primary or secondary 1956 tumour implantation. Data represent mean ± s.d. pooled biologically independent samples from two independent experiments (n = 4 for FTY720 1º and n = 5 for all other groups). e, Individual mouse tumour growth curves of control or FTY720 injected mice. f, (Left) Tumour growth curves of Xcr1+/+ MHCIIfl/fl and Xcr1Cre/+ MHCIIfl/fl mice during primary and secondary 1956 tumour implantation. Individual mouse tumour growth curves of (middle) Xcr1+/+ MHCIIfl/fl or (right) Xcr1Cre/+ MHCIIfl/fl mice during primary and secondary 1956 tumour implantation. Data represent mean ± s.d. pooled biologically independent samples from two independent experiments (n = 7 for Xcr1+/+ MHCIIfl/fl and n = 6 for Xcr1Cre/+ MHCIIfl/fl). g, (Left) Tumour growth curves of Xcr1+/+ CD40fl/fl and Xcr1Cre/+ CD40fl/fl mice during primary and secondary 1956 tumour implantation. Individual mouse tumour growth curves of (middle) Xcr1+/+ CD40fl/fl or (right) Xcr1Cre/+ CD40fl/fl mice during primary and secondary 1956 tumour implantation. Data represent mean ± s.d. pooled biologically independent samples from two independent experiments (n = 4 for Xcr1+/+ CD40fl/fl and n = 7 for Xcr1Cre/+ CD40fl/fl).
Extended Data Fig. 9 OT-II CD4+ T cells fail to localize to the tumour in Xcr1Cre/+ MHCIIfl/fl mice.
a, Graph of per cent accumulation of transferred OT-I in tumours of Xcr1+/+ β2mfl/fl, Xcr1Cre/+ β2mfl/fl injected with 106 1956-EV or 1956-mOVA on day 0. OT-I cells were transferred intravenously on day 2 and assessed as a percentage of total CD45+ cells on (left) day 5 and (right) day 7. Data represent pooled biologically independent samples from two independent experiments (n = 1 for Xcr1+/+ β2mfl/fl tdLN and tumour 1956-EV day 5, n = 4 for Xcr1+/+ β2mfl/fl 1956-mOVA tdLN and tumour Day 7 n = 2 for all other groups). b, Graph of per cent accumulation of transferred OT-I in tumours of Xcr1+/+ MHCIIfl/fl, Xcr1Cre/+ MHCIIfl/fl injected with 106 1956-EV or 1956-mOVA on day 0. OT-I cells were transferred intravenously on day 2 and assessed as a percentage of total CD45+ cells on (left) day 5 and (right) day 7. Data represent pooled biologically independent samples from two independent experiments (n = 4 for Xcr1+/+ MHCIIfl/fl tdLN and tumour 1956-mOVA day 7, n = 3 for Xcr1+/+ MHCIIfl/fl tdLN and tumour 1956-mOVA day 5 and for Xcr1+/+ MHCIIfl/fl tdLN and tumour 1956-EV Day 7, n = 2 for all other samples). c, Graph of per cent accumulation of transferred OT-II in tumours of Xcr1+/+ MHCIIfl/fl, Xcr1Cre/+ MHCIIfl/fl injected with 106 1956-EV or 1956-mOVA on day 0. OT-II cells were transferred intravenously on day 2 and assessed as a percentage of total CD45+ cells on (left) day 5 and (right) day 7. Data represent pooled biologically independent samples from two independent experiments (n = 6 for Xcr1+/+ MHCIIfl/fl tdLN and tumour 1956-mOVA day 7, n = 4 for Xcr1+/+ MHCIIfl/fl tdLN and tumour 1956-EV Day 7, n = 3 for Xcr1+/+ MHCIIfl/fl tdLN and tumour 1956-mOVA day 5, n = 2 for all other samples).
Supplementary information
Supplementary Figure
This file contains full scan southern blots for Extended Figures 3b-c.
Supplementary Table
This file contains Supplementary Table 1.
Source data
Rights and permissions
About this article
Cite this article
Ferris, S.T., Durai, V., Wu, R. et al. cDC1 prime and are licensed by CD4+ T cells to induce anti-tumour immunity. Nature 584, 624–629 (2020). https://doi.org/10.1038/s41586-020-2611-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2611-3
This article is cited by
-
Immune modulation in malignant pleural effusion: from microenvironment to therapeutic implications
Cancer Cell International (2024)
-
Dendritic cell-targeted therapy expands CD8 T cell responses to bona-fide neoantigens in lung tumors
Nature Communications (2024)
-
Dendritic cells as orchestrators of anticancer immunity and immunotherapy
Nature Reviews Clinical Oncology (2024)
-
A hepatic network of dendritic cells mediates CD4 T cell help outside lymphoid organs
Nature Communications (2024)
-
Lymphatic vessels in the age of cancer immunotherapy
Nature Reviews Cancer (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.