Abstract
Members of the conserved Argonaute protein family use small RNA guides to locate their mRNA targets and regulate gene expression and suppress mobile genetic elements in eukaryotes1,2. Argonautes are also present in many bacterial and archaeal species3,4,5. Unlike eukaryotic proteins, several prokaryotic Argonaute proteins use small DNA guides to cleave DNA, a process known as DNA interference6,7,8,9,10. However, the natural functions and targets of DNA interference are poorly understood, and the mechanisms of DNA guide generation and target discrimination remain unknown. Here we analyse the activity of a bacterial Argonaute nuclease from Clostridium butyricum (CbAgo) in vivo. We show that CbAgo targets multicopy genetic elements and suppresses the propagation of plasmids and infection by phages. CbAgo induces DNA interference between homologous sequences and triggers DNA degradation at double-strand breaks in the target DNA. The loading of CbAgo with locus-specific small DNA guides depends on both its intrinsic endonuclease activity and the cellular double-strand break repair machinery. A similar interaction was reported for the acquisition of new spacers during CRISPR adaptation, and prokaryotic genomes that encode Ago nucleases are enriched in CRISPR–Cas systems. These results identify molecular mechanisms that generate guides for DNA interference and suggest that the recognition of foreign nucleic acids by prokaryotic defence systems involves common principles.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated during this study are included in the published Article and the Extended Data and are available from the Gene Expression Omnibus (GEO) database with the accession number GSE148596.
Code availability
The code used for data analysis is available at the GitHub repository at https://github.com/AntKuzmenko/CbAgo_DNAi.git.
References
Gebert, D. & Rosenkranz, D. RNA-based regulation of transposon expression. Wiley Interdiscip. Rev. RNA 6, 687–708 (2015).
Meister, G. Argonaute proteins: functional insights and emerging roles. Nat. Rev. Genet. 14, 447–459 (2013).
Makarova, K. S., Wolf, Y. I., van der Oost, J. & Koonin, E. V. Prokaryotic homologs of Argonaute proteins are predicted to function as key components of a novel system of defense against mobile genetic elements. Biol. Direct 4, 29 (2009).
Ryazansky, S., Kulbachinskiy, A. & Aravin, A. A. The expanded universe of prokaryotic Argonaute proteins. MBio 9, e01935–e01918 (2018).
Swarts, D. C. et al. The evolutionary journey of Argonaute proteins. Nat. Struct. Mol. Biol. 21, 743–753 (2014).
Hegge, J. W. et al. DNA-guided DNA cleavage at moderate temperatures by Clostridium butyricum Argonaute. Nucleic Acids Res. 47, 5809–5821 (2019).
Kuzmenko, A., Yudin, D., Ryazansky, S., Kulbachinskiy, A. & Aravin, A. A. Programmable DNA cleavage by Ago nucleases from mesophilic bacteria Clostridium butyricum and Limnothrix rosea. Nucleic Acids Res. 47, 5822–5836 (2019).
Swarts, D. C. et al. Argonaute of the archaeon Pyrococcus furiosus is a DNA-guided nuclease that targets cognate DNA. Nucleic Acids Res. 43, 5120–5129 (2015).
Swarts, D. C. et al. DNA-guided DNA interference by a prokaryotic Argonaute. Nature 507, 258–261 (2014).
Zander, A. et al. Guide-independent DNA cleavage by archaeal Argonaute from Methanocaldococcus jannaschii. Nat. Microbiol. 2, 17034 (2017).
Willkomm, S., Makarova, K. S. & Grohmann, D. DNA silencing by prokaryotic Argonaute proteins adds a new layer of defense against invading nucleic acids. FEMS Microbiol. Rev. 42, 376–387 (2018).
Lisitskaya, L., Aravin, A. A. & Kulbachinskiy, A. DNA interference and beyond: structure and functions of prokaryotic Argonaute proteins. Nat. Commun. 9, 5165 (2018).
Kaya, E. et al. A bacterial Argonaute with noncanonical guide RNA specificity. Proc. Natl Acad. Sci. USA 113, 4057–4062 (2016).
Sheng, G. et al. Structure-based cleavage mechanism of Thermus thermophilus Argonaute DNA guide strand-mediated DNA target cleavage. Proc. Natl Acad. Sci. USA 111, 652–657 (2014).
Willkomm, S. et al. Structural and mechanistic insights into an archaeal DNA-guided Argonaute protein. Nat. Microbiol. 2, 17035 (2017).
Olina, A. et al. Genome-wide DNA sampling by Ago nuclease from the cyanobacterium Synechococcus elongatus. RNA Biol. 17, 677–688 (2020).
Swarts, D. C. et al. Autonomous generation and loading of DNA uides by bacterial Argonaute. Mol. Cell 65, 985–998.e6 (2017).
Olovnikov, I., Chan, K., Sachidanandam, R., Newman, D. K. & Aravin, A. A. Bacterial argonaute samples the transcriptome to identify foreign DNA. Mol. Cell 51, 594–605 (2013).
Duggin, I. G. & Bell, S. D. Termination structures in the Escherichia coli chromosome replication fork trap. J. Mol. Biol. 387, 532–539 (2009).
Dillingham, M. S. & Kowalczykowski, S. C. RecBCD enzyme and the repair of double-stranded DNA breaks. Microbiol. Mol. Biol. Rev. 72, 642–671 (2008).
Smith, G. R. How RecBCD enzyme and Chi promote DNA break repair and recombination: a molecular biologist’s view. Microbiol. Mol. Biol. Rev. 76, 217–228 (2012).
Wigley, D. B. Bacterial DNA repair: recent insights into the mechanism of RecBCD, AddAB and AdnAB. Nat. Rev. Microbiol. 11, 9–13 (2013).
Chaudhury, A. M. & Smith, G. R. Escherichia coli recBC deletion mutants. J. Bacteriol. 160, 788–791 (1984).
Sinha, A. K. et al. Division-induced DNA double strand breaks in the chromosome terminus region of Escherichia coli lacking RecBCD DNA repair enzyme. PLoS Genet. 13, e1006895 (2017).
Sinha, A. K. et al. Broken replication forks trigger heritable DNA breaks in the terminus of a circular chromosome. PLoS Genet. 14, e1007256 (2018).
Capaldo, F. N. & Barbour, S. D. DNA content, synthesis and integrity in dividing and non-dividing cells of rec- strains of Escherichia coli K12. J. Mol. Biol. 91, 53–66 (1975).
Marraffini, L. A. & Sontheimer, E. J. Self versus non-self discrimination during CRISPR RNA-directed immunity. Nature 463, 568–571 (2010).
Westra, E. R. et al. Type I-E CRISPR-cas systems discriminate target from non-target DNA through base pairing-independent PAM recognition. PLoS Genet. 9, e1003742 (2013).
White, M. A., Azeroglu, B., Lopez-Vernaza, M. A., Hasan, A. M. M. & Leach, D. R. F. RecBCD coordinates repair of two ends at a DNA double-strand break, preventing aberrant chromosome amplification. Nucleic Acids Res. 46, 6670–6682 (2018).
Eykelenboom, J. K., Blackwood, J. K., Okely, E. & Leach, D. R. SbcCD causes a double-strand break at a DNA palindrome in the Escherichia coli chromosome. Mol. Cell 29, 644–651 (2008).
White, M. A., Darmon, E., Lopez-Vernaza, M. A. & Leach, D. R. F. DNA double strand break repair in Escherichia coli perturbs cell division and chromosome dynamics. PLoS Genet. 16, e1008473 (2020).
Modell, J. W., Jiang, W. & Marraffini, L. A. CRISPR-Cas systems exploit viral DNA injection to establish and maintain adaptive immunity. Nature 544, 101–104 (2017).
Levy, A. et al. CRISPR adaptation biases explain preference for acquisition of foreign DNA. Nature 520, 505–510 (2015).
Ivančić-Baće, I., Cass, S. D., Wearne, S. J. & Bolt, E. L. Different genome stability proteins underpin primed and naïve adaptation in E. coli CRISPR-Cas immunity. Nucleic Acids Res. 43, 10821–10830 (2015).
Bobay, L. M., Touchon, M. & Rocha, E. P. Manipulating or superseding host recombination functions: a dilemma that shapes phage evolvability. PLoS Genet. 9, e1003825 (2013).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Ofir, G. et al. DISARM is a widespread bacterial defence system with broad anti-phage activities. Nat. Microbiol. 3, 90–98 (2018).
Koonin, E. V. Evolution of RNA- and DNA-guided antivirus defense systems in prokaryotes and eukaryotes: common ancestry vs convergence. Biol. Direct 12, 5 (2017).
El Karoui, M., Biaudet, V., Schbath, S. & Gruss, A. Characteristics of Chi distribution on different bacterial genomes. Res. Microbiol. 150, 579–587 (1999).
Jolly, S. M. et al. A DNA-guided Argonaute protein functions in DNA replication in Thermus thermophilus. Cell https://doi.org/10.1016/j.cell.2020.07.036 (2020)
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Bohn, C., Collier, J. & Bouloc, P. Dispensable PDZ domain of Escherichia coli YaeL essential protease. Mol. Microbiol. 52, 427–435 (2004).
He, F. E. coli genomic DNA extraction. Bio-101 e97, https://doi.org/10.21769/BioProtoc.97 (2011).
Thomason, L. C., Costantino, N. & Court, D. L. E. coli genome manipulation by P1 transduction. Curr. Protocols Mol. Biol. Ch. 1, Unit 1.17 (2007).
Bernheim, A., Bikard, D., Touchon, M. & Rocha, E. P. C. A matter of background: DNA repair pathways as a possible cause for the sparse distribution of CRISPR-Cas systems in bacteria. Phil. Trans. R. Soc. Lond. B 374, 20180088 (2019).
Couvin, D. et al. CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucleic Acids Res. 46 (W1), W246–W251 (2018).
Acknowledgements
We thank M. A. White for strains with engineered I-SceI sites, G. Smith for RecBCD antibodies, S. Lysenkov for the help with statistical analysis and A. Olina for the help with the phage experiments. D.Y. would like to thank M. Kolesnik for discussions. The project was in part supported by the Ministry of Science and Higher Education of the Russian Federation (14.W03.31.0007), Russian Science Foundation (19-14-00359, analysis of DSB formation), Russian Foundation for Basic Research (18-29-07086). A.A.A. is supported by an HHMI Faculty Scholar Award. D.L. is supported by grant MR/M019160/1 from the MRC (UK).
Author information
Authors and Affiliations
Contributions
A. Kuzmenko, A.A.A. and A. Kulbachinskiy conceptualized the study. A. Kuzmenko, D.Y. and D.E. constructed strains. A. Kuzmenko, A.O., D.Y. and D.E. prepared smDNA libraries. A. Kuzmenko, A.O. and D.E. prepared genomic DNA libraries. A. Kuzmenko, A.O. and D.Y. analysed sequencing data, M.N. and S.R. helped with data analysis. S.R. performed phylogenetic analysis. D.L. conceptualized experiments with engineered DSBs. A. Kuzmenko, A. Kudinova, O.M., M.P., A.O. and D.E. performed experiments on plasmid elimination and phage infection. All authors interpreted the results. A. Kuzmenko and A.O. prepared the figures. A. Kulbachinskiy and A.A.A. wrote the manuscript with contribution from other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Small DNAs associated with CbAgo.
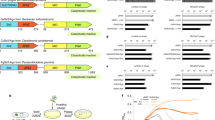
a, Analysis of small nucleic acids isolated from wild-type CbAgo and dCbAgo. The samples were treated with alkaline phosphatase, 32P-labelled with polynucleotide kinase and treated with DNase I (D), RNase A (R) or left without further treatment (−). CbAgo is associated with small DNAs, as confirmed by their sensitivity to DNase treatment and resistance to RNase. The DNA marker (M) lengths are indicated. For gel source data, see Supplementary Fig. 1. b, Length distribution of smDNAs associated with CbAgo in the wild-type, recBrecD, recC and recA strains. For the recC strain, there is a small increase in the smDNA length, suggesting that their processing might be different in this strain. c, d, Analysis of nucleotide biases for chromosomal (wild-type CbAgo and dCbAgo), plasmid (pNonChi) and phage M13 smDNAs associated with CbAgo. c, Nucleotide frequencies at different guide positions. d, AT/GC-content along the guide length and in surrounding genomic sequences. Guide positions starting from the 5′ end are indicated below the plots. For genomic DNA, the AT-bias around the first position is seen for both active CbAgo and dCbAgo. The AT-bias in the downstream region (positions 14–18) is seen for active CbAgo but not for dCbAgo. For each replicon, the average GC content of smDNAs corresponds to the GC content of this replicon (shown in percentage in each panel), indicating that the efficiency of smDNA processing does not strongly depend on the GC content. e, Model of processing of smDNAs by CbAgo from double-stranded DNA precursors. Binding of the guide 5′ end in the MID-pocket of CbAgo may be facilitated by melting of the DNA duplex in the upstream guide region (left). Guide DNA loading is completed after CbAgo-dependent cleavage of the complementary DNA strand and its dissociation, depending on the AT content of the downstream guide–target duplex (right).
Extended Data Fig. 2 Whole-genome mapping of smDNAs associated with CbAgo in strains with various genetic backgrounds.
a–h, For each strain, the distribution of smDNAs along the chromosome is shown in RPKM. Left, total smDNA counts. Right, strand distribution of smDNAs for each strain (plus DNA strand, green; minus DNA strand, red). Positions of the araC, lacI and ter sites are shown above the plots. smDNA coverage is shown in RPKM. The identities of the strains and plasmids, with plasmid or chromosomal localizations of the CbAgo gene, are indicated (Supplementary Tables 2 and 3). a, Wild-type BL21(DE3) with plasmid-encoded CbAgo (pBAD containing the araC gene). b, As in a, in BL21(DE3) with knockout of Tus. c, MG1655Z1 with genomic CbAgo. d, As in a with pET28b containing lacI. e, Plasmid-encoded catalytically dead dCbAgo in BL21(DE3). f, Knockout of RecB/RecD in BL21(DE3) with plasmid-encoded CbAgo. g, Knockout of RecC in BL21(DE3) with plasmid-encoded CbAgo. h, Knockout of RecA in BL21(DE3) with plasmid-encoded CbAgo. The observed enrichment of smDNAs around the ori region in the recC and recA strains may possibly reflect the higher DNA content and/or a higher likelihood of DSB formation in this region in these strains. i, Targeting of specific genomic regions depends on the catalytic activity of CbAgo. The ratio of smDNAs between wild-type CbAgo and dCbAgo (obtained for BL21(DE3) containing corresponding pBAD_CbAgo plasmids) is shown in the logarithmic scale. Normalized densities of smDNA reads (RPKM) were calculated for each CbAgo variant and plotted as a wild-type/dCb ratio. The regions with the ratio of >1 correspond to the sites of active smDNA processing by CbAgo. CbAgo targets the araC locus, ter region and multicopy sequences: rDNA operons (indicated with arrows above the plot) and IS elements. Positions of IS1 (29 copies) and IS3 (12 copies) in the BL21(DE3) genome are shown with dotted lines below the plot.
Extended Data Fig. 3 Asymmetry in smDNA distribution at specific genomic loci.
a, Zoomed-in peaks of smDNAs around the araC and lacI genes in strains containing plasmids with corresponding genes. b, Examples of smDNA distributions around rRNA operons, rrsD and rrsC, in wild-type E. coli and strains with knockouts of recBrecD and recC. The reads from the plus and minus genomic strands are shown in green and red, respectively. Positions of Chi sites in surrounding genomic regions are indicated (forward for the plus strand and reverse for the minus strand); the closest Chi sites in the corresponding strands are shown with dotted lines. c, Metaplot of the number of smDNAs calculated in the 500-bp windows around Chi sites in each genomic strand (red, plus-strand smDNAs for plus-strand Chi sites; green, minus-strand smDNAs for minus-strand Chi sites) in the 2–3-Mb genomic region. Position around Chi is shown in kilobases. d, Strand-specific asymmetry in smDNA distribution for various strains (ratio of RPKM values for the plus and minus genomic strands). A similar bias is observed for the wild-type and recBrecD, recC and recA mutant strains expressing CbAgo but not in wild-type cells expressing catalytically inactive dCbAgo.
Extended Data Fig. 4 Growth kinetics of E. coli strains depending on the expression of CbAgo.
a, b, Growth kinetics of E. coli BL21(DE3) and its mutant derivatives with or without CbAgo (containing pBAD_CbAgo or empty pBAD plasmids) at 30 °C in the rich LB (a) and minimal M9 (b) medium. Overnight cultures of cells were inoculated into fresh LB to OD600 of 0.01 in the presence of the inductor (0.05% l-arabinose) and cell density was measured at 15 min intervals in a microplate reader.
Extended Data Fig. 5 Whole-genome analysis of DNA content in the wild-type and tus− E. coli strains depending on the expression of CbAgo.
a–d, The experiment was performed with wild-type (a, b) or tus− (c, d) BL21(DE3) containing or lacking the pBAD_CbAgo plasmid. The cells were collected at the exponential phase (OD600 = 0.5) (a, c) or stationary phase (OD600 = 6) (b, d), followed by isolation of total DNA and sequencing. For each condition, genomic DNA coverage is shown for strains without and with CbAgo, and the ratio for the +CbAgo and −CbAgo strains is shown in a separate panel (black). The enlarged ter region and the araC locus are shown separately. Genomic DNA coverage is shown in RPKM. At the stationary phase, a peak in genomic DNA coverage was detected in the strains containing CbAgo, which exactly corresponded to the DE3 prophage in BL21(DE3). This may indicate formation of DSBs in this region, possibly as a result of partial prophage excision, leading to DNA repair and replication.
Extended Data Fig. 6 Targeting of engineered DSBs by CbAgo.
a, Top, smDNA abundance in the chromosomal area spanning the engineered DSBs (palindrome or I-SceI-dependent; I-SceImut, the mutated cleavage site) and ter sites, for the wild-type CbAgo or dCbAgo. In each strain, the numbers of smDNAs mapping to the region of DSB are shown in percentage of total smDNAs. The presence of the DSB shifts the ratio between the terA and terC peaks in favour of terA, probably as a result of impediment of the clockwise replisome, moving towards terC, by the DSB formation. Bottom, strand-specific distribution of smDNAs around engineered DSBs for strains with dCbAgo (palindrome and I-SceI DSBs) or with wild-type CbAgo and the I-SceImut site. The reads from the plus and minus DNA strands are shown in green and red, respectively. Most smDNAs are produced from the 3′ strand at each end of the DSB, and the boundaries of the smDNA peaks are defined by Chi sites. b, Genomic DNA coverage in the same region in palindrome-containing strains depending on the expression of active CbAgo or dCbAgo. c, The ratio of genomic DNA profiles for palindrome-containing strains with wild-type CbAgo and dCbAgo relative to the strain without CbAgo. Wild-type CbAgo but not dCbAgo triggers DNA loss around the DSB with overreplication of genomic DNA at the site of termination. d, Genomic DNA coverage at DSBs formed by the I-SceI meganuclease in E. coli strains with induced I-SceI but without expression of CbAgo (left) and with expression of both I-SceI and CbAgo (right). e, The ratio between genomic DNA profiles for the strains with and without expression of CbAgo. Genomic DNA coverage is shown in RPKM.
Extended Data Fig. 7 Targeting of plasmid and phage DNA by CbAgo.
a, smDNA coverage of plasmids (pNonChi, left; pBAD_CbAgo, right) in strains with plasmid-encoded CbAgo. The moving average of smDNA coverage in a 200-nucleotide window is shown for the plus and minus DNA strands (green and red, respectively). b, smDNA coverage of a pET28 plasmid in a strain with chromosomal CbAgo. c, Distribution of smDNAs along the M13 genome. smDNAs were isolated from CbAgo expressed in E. coli NEB Turbo strain during infection with M13. d, Coverage of plasmid DNA in whole-genome sequencing in the wild-type and tus− strains, depending on the expression of CbAgo. The values represent the moving average of genomic DNA coverage in a 200-nucleotide window (in RPM). e–g, Targeting of the F′ plasmid by CbAgo. Total smDNA coverage (e), coverage of the plus and minus DNA strands (f) and the plus-to-minus strand ratio (g) are shown along the F′ sequence. Positions of the three copies of IS3 element, the origin of replication (oriS), the core part of the F factor, and the chromosomal insertion (‘chr’) are indicated. For strand-specific smDNA distribution, positions of the nearest Chi sites in the corresponding strands are shown. Most reads map to the F episome core sequence lacking Chi sites, and the numbers of smDNAs drop considerably upon encountering the first Chi site. The distribution is also asymmetric relative to the origin of replication. Similarly to the chromosome (Extended Data Fig. 3d), the lagging DNA strand is targeted with a higher efficiency, which suggests a connection to replication.
Extended Data Fig. 8 Loss of plasmids after various number of passages in E. coli strains with or without CbAgo.
Cells expressing genome-encoded CbAgo (Cb), its catalytically dead mutant (dCb) or without Ago (‘without’) were transformed with one of the six different plasmids from different incompatibility groups. The percentage of plasmid-free cells was measured after indicated number of passages (mean and s.d. from 2–6 biological replicates). CbAgo, but not dCbAgo, facilitates plasmid elimination regardless of the plasmid type.
Extended Data Fig. 9 Effects of CbAgo and dCbAgo on P1 infection.
a, Bacterial culture growth during P1 infection with different MOI in strain with or without dCbAgo. Data are mean and s.d. from three independent experiments. b, Titres of P1 at MOI 1 and 5 at different times after infection in strains without CbAgo or with expression CbAgo or dCbAgo. Data are mean and s.d. from three–four independent measurements. *P < 0.05, **P < 0.01, ***P < 0.001, Scheffe’s test for multiple comparison of mean values after normalization of data by log-transformation.
Extended Data Fig. 10 Co-occurrence of pAgo proteins, DSB repair systems and CRISPR–Cas in prokaryotic genomes.
a, Circular phylogenetic tree of pAgos from prokaryotic strains with fully assembled genomes based on the multiple alignment of the MID-PIWI domains. Three major phylogenetic groups of pAgos are indicated4: long-A pAgos usually contain all characteristic domains of the Ago family (N, PAZ, MID and PIWI) and have a predicted nuclease site; long-B pAgos also contain all domains but are inactive; and short pAgos contain only MID and PIWI domains and are inactive. The pAgo proteins were annotated as follows, from the inner to the outer circles: the superkingdom to which the corresponding pAgo belongs; the type of the PIWI domain, depending on the presence of the catalytic tetrad DEDX; the type of the DSB repair system encoded in the corresponding genome; the class of CRISPR–Cas system; the type and subtype of CRISPR–Cas system. CbAgo, T. thermophilus Ago (TtAgo) and Marinitoga piezophila Ago (MpAgo) are highlighted in red. The scale bar represents the evolutionary rate calculated under the JTT+CAT evolutionary model. b, The distribution of various subtypes of type I and type III CRISPR–Cas system in the fully assembled genomes encoding pAgos. The number of genomes for each pAgo group is indicated.
Supplementary information
Supplementary Figure 1
The file contains raw images for all data obtained by gel electrophoresis in the indicated figures.
Supplementary Table
The file contains Supplementary Tables 1-5.
Rights and permissions
About this article
Cite this article
Kuzmenko, A., Oguienko, A., Esyunina, D. et al. DNA targeting and interference by a bacterial Argonaute nuclease. Nature 587, 632–637 (2020). https://doi.org/10.1038/s41586-020-2605-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2605-1
This article is cited by
-
Inhibitors of bacterial immune systems: discovery, mechanisms and applications
Nature Reviews Genetics (2024)
-
DNA-targeting short Argonautes complex with effector proteins for collateral nuclease activity and bacterial population immunity
Nature Microbiology (2024)
-
Structural basis of antiphage immunity generated by a prokaryotic Argonaute-associated SPARSA system
Nature Communications (2024)
-
Nucleic-acid-triggered NADase activation of a short prokaryotic Argonaute
Nature (2024)
-
Origins and diversification of animal innate immune responses against viral infections
Nature Ecology & Evolution (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.