Abstract
Complex organisms can rapidly induce select genes in response to diverse environmental cues. This regulation occurs in the context of large genomes condensed by histone proteins into chromatin. The sensing of pathogens by macrophages engages conserved signalling pathways and transcription factors to coordinate the induction of inflammatory genes1,2,3. Enriched integration of histone H3.3, the ancestral histone H3 variant, is a general feature of dynamically regulated chromatin and transcription4,5,6,7. However, how chromatin is regulated at induced genes, and what features of H3.3 might enable rapid and high-level transcription, are unknown. The amino terminus of H3.3 contains a unique serine residue (Ser31) that is absent in ‘canonical’ H3.1 and H3.2. Here we show that this residue, H3.3S31, is phosphorylated (H3.3S31ph) in a stimulation-dependent manner along rapidly induced genes in mouse macrophages. This selective mark of stimulation-responsive genes directly engages the histone methyltransferase SETD2, a component of the active transcription machinery, and ‘ejects’ the elongation corepressor ZMYND118,9. We propose that features of H3.3 at stimulation-induced genes, including H3.3S31ph, provide preferential access to the transcription apparatus. Our results indicate dedicated mechanisms that enable rapid transcription involving the histone variant H3.3, its phosphorylation, and both the recruitment and the ejection of chromatin regulators.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Source data for immunoblots are provided in Supplementary Fig. 1. All ChIP–seq and RNA-seq data have been deposited in the NCBI Gene Expression Omnibus (GEO) under accession GSE125159. Coordinates and structure factors are available from the Protein Data Bank (PDB) with accession code 6J9J. There are no restrictions on data availability.
References
Medzhitov, R. & Horng, T. Transcriptional control of the inflammatory response. Nat. Rev. Immunol. 9, 692–703 (2009).
Smale, S. T., Tarakhovsky, A. & Natoli, G. Chromatin contributions to the regulation of innate immunity. Annu. Rev. Immunol. 32, 489–511 (2014).
Akira, S., Uematsu, S. & Takeuchi, O. Pathogen recognition and innate immunity. Cell 124, 783–801 (2006).
Filipescu, D., Szenker, E. & Almouzni, G. Developmental roles of histone H3 variants and their chaperones. Trends Genet. 29, 630–640 (2013).
Henikoff, S. & Smith, M. M. Histone variants and epigenetics. Cold Spring Harb. Perspect. Biol. 7, a019364 (2015).
Kamada, R. et al. Interferon stimulation creates chromatin marks and establishes transcriptional memory. Proc. Natl Acad. Sci. USA 115, E9162–E9171 (2018).
Shi, L., Wen, H. & Shi, X. The histone variant H3.3 in transcriptional regulation and human disease. J. Mol. Biol. 429, 1934–1945 (2017).
Wen, H. et al. ZMYND11 links histone H3.3K36me3 to transcription elongation and tumour suppression. Nature 508, 263–268 (2014).
Guo, R. et al. BS69/ZMYND11 reads and connects histone H3.3 lysine 36 trimethylation-decorated chromatin to regulated pre-mRNA processing. Mol. Cell 56, 298–310 (2014).
Duarte, F. M. et al. Transcription factors GAF and HSF act at distinct regulatory steps to modulate stress-induced gene activation. Genes Dev. 30, 1731–1746 (2016).
Healy, S., Khan, P., He, S. & Davie, J. R. Histone H3 phosphorylation, immediate-early gene expression, and the nucleosomal response: a historical perspective. Biochem. Cell Biol. 90, 39–54 (2012).
Rossetto, D., Avvakumov, N. & Côté, J. Histone phosphorylation: a chromatin modification involved in diverse nuclear events. Epigenetics 7, 1098–1108 (2012).
Sawicka, A. et al. H3S28 phosphorylation is a hallmark of the transcriptional response to cellular stress. Genome Res. 24, 1808–1820 (2014).
Zippo, A. et al. Histone crosstalk between H3S10ph and H4K16ac generates a histone code that mediates transcription elongation. Cell 138, 1122–1136 (2009).
Lau, P. N. I. & Cheung, P. Histone code pathway involving H3 S28 phosphorylation and K27 acetylation activates transcription and antagonizes polycomb silencing. Proc. Natl Acad. Sci. USA 108, 2801–2806 (2011).
Josefowicz, S. Z. et al. Chromatin kinases act on transcription factors and histone tails in regulation of inducible transcription. Mol. Cell 64, 347–361 (2016).
Waterborg, J. H. Evolution of histone H3: emergence of variants and conservation of post-translational modification sites. Biochem. Cell Biol. 90, 79–95 (2012).
Duronio, R. J. & Marzluff, W. F. Coordinating cell cycle-regulated histone gene expression through assembly and function of the histone locus body. RNA Biol. 14, 726–738 (2017).
Zink, L. M. & Hake, S. B. Histone variants: nuclear function and disease. Curr. Opin. Genet. Dev. 37, 82–89 (2016).
Gomes, A. P. et al. Dynamic incorporation of histone H3 variants into chromatin is essential for acquisition of aggressive traits and metastatic colonization. Cancer Cell 36, 402–417 (2019).
Sitbon, D., Boyarchuk, E. & Almouzni, G. Histone variant H3.3 residue S31 is essential for Xenopus gastrulation regardless of the deposition pathway. Nat. Commun. 11, 1256 (2020)
Martire, S. et al. Phosphorylation of histone H3.3 at serine 31 promotes p300 activity and enhancer acetylation. Nat. Genet. 51, 941–946 (2019).
Hake, S. B. et al. Serine 31 phosphorylation of histone variant H3.3 is specific to regions bordering centromeres in metaphase chromosomes. Proc. Natl Acad. Sci. USA 102, 6344–6349 (2005).
Thorne, J. L., Ouboussad, L. & Lefevre, P. F. Heterochromatin protein 1 gamma and IκB kinase alpha interdependence during tumour necrosis factor gene transcription elongation in activated macrophages. Nucleic Acids Res. 40, 7676–7689 (2012).
Chang, F. T. M. et al. CHK1-driven histone H3.3 serine 31 phosphorylation is important for chromatin maintenance and cell survival in human ALT cancer cells. Nucleic Acids Res. 43, 2603–2614 (2015).
Li, M., Dong, Q. & Zhu, B. Aurora kinase B phosphorylates histone H3.3 at serine 31 during mitosis in mammalian cells. J. Mol. Biol. 429, 2042–2045 (2017).
Hinchcliffe, E. H. et al. Chromosome missegregation during anaphase triggers p53 cell cycle arrest through histone H3.3 Ser31 phosphorylation. Nat. Cell Biol. 18, 668–675 (2016).
Madabhushi, R. et al. Activity-induced DNA breaks govern the expression of neuronal early-response genes. Cell 161, 1592–1605 (2015).
Huang, W.-C. & Hung, M.-C. Beyond NF-κB activation: nuclear functions of IκB kinase α. J. Biomed. Sci. 20, 3 (2013).
Wagner, E. J. & Carpenter, P. B. Understanding the language of Lys36 methylation at histone H3. Nat. Rev. Mol. Cell Biol. 13, 115–126 (2012).
Baubec, T. et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520, 243–247 (2015).
Neri, F. et al. Intragenic DNA methylation prevents spurious transcription initiation. Nature 543, 72–77 (2017).
Fierz, B. & Muir, T. W. Chromatin as an expansive canvas for chemical biology. Nat. Chem. Biol. 8, 417–427 (2012).
Zheng, W. et al. Sinefungin derivatives as inhibitors and structure probes of protein lysine methyltransferase SETD2. J. Am. Chem. Soc. 134, 18004–18014 (2012).
Yang, S. et al. Molecular basis for oncohistone H3 recognition by SETD2 methyltransferase. Genes Dev. 30, 1611–1616 (2016).
Kruidenier, L. et al. A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nature 488, 404–408 (2012).
De Santa, F. et al. Jmjd3 contributes to the control of gene expression in LPS-activated macrophages. EMBO J. 28, 3341–3352 (2009).
Jones, S. E., Olsen, L. & Gajhede, M. Structural basis of histone demethylase KDM6B histone 3 lysine 27 specificity. Biochemistry 57, 585–592 (2018).
Andrews, F. H., Gatchalian, J., Krajewski, K., Strahl, B. D. & Kutateladze, T. G. Regulation of methyllysine readers through phosphorylation. ACS Chem. Biol. 11, 547–553 (2016).
Sarma, K., Margueron, R., Ivanov, A., Pirrotta, V. & Reinberg, D. Ezh2 requires PHF1 to efficiently catalyze H3 lysine 27 trimethylation in vivo. Mol. Cell. Biol. 28, 2718–2731 (2008).
Bush, K. M. et al. Endogenous mammalian histone H3.3 exhibits chromatin-related functions during development. Epigenetics Chromatin 6, 7 (2013).
Jang, C.-W., Shibata, Y., Starmer, J., Yee, D. & Magnuson, T. Histone H3.3 maintains genome integrity during mammalian development. Genes Dev. 29, 1377–1392 (2015).
Wen, D. et al. Histone variant H3.3 is an essential maternal factor for oocyte reprogramming. Proc. Natl Acad. Sci. USA 111, 7325–7330 (2014).
Kong, Q. et al. Histone variant H3.3-mediated chromatin remodeling is essential for paternal genome activation in mouse preimplantation embryos. J. Biol. Chem. 293, 3829–3838 (2018).
Yuen, B. T. K., Bush, K. M., Barrilleaux, B. L., Cotterman, R. & Knoepfler, P. S. Histone H3.3 regulates dynamic chromatin states during spermatogenesis. Development 141, 3483–3494 (2014).
Bachu, M. et al. A versatile mouse model of epitope-tagged histone H3.3 to study epigenome dynamics. J. Biol. Chem. 294, 1904–1914 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Shen, L., Shao, N., Liu, X. & Nestler, E. ngs.plot: quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 15, 284 (2014).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Tong, A. J. et al. A stringent systems approach uncovers gene-specific mechanisms regulating inflammation. Cell 165, 165–179 (2016).
Bhatt, D. M. et al. Transcript dynamics of proinflammatory genes revealed by sequence analysis of subcellular RNA fractions. Cell 150, 279–290 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Shechter, D., Dormann, H. L., Allis, C. D. & Hake, S. B. Extraction, purification and analysis of histones. Nat. Protocols 2, 1445–1457 (2007).
Jack, A. P. M. et al. H3K56me3 is a novel, conserved heterochromatic mark that largely but not completely overlaps with H3K9me3 in both regulation and localization. PLoS One 8, e51765 (2013).
Wiedemann, S. M. et al. Identification and characterization of two novel primate-specific histone H3 variants, H3.X and H3.Y. J. Cell Biol. 190, 777–791 (2010).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protocols 8, 2281–2308 (2013).
Mossessova, E. & Lima, C. D. Ulp1-SUMO crystal structure and genetic analysis reveal conserved interactions and a regulatory element essential for cell growth in yeast. Mol. Cell 5, 865–876 (2000).
Ruthenburg, A. J. et al. Recognition of a mononucleosomal histone modification pattern by BPTF via multivalent interactions. Cell 145, 692–706 (2011).
Lowary, P. T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998).
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol. 375, 23–44 (2004).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Rothbart, S. B. et al. An interactive database for the assessment of histone antibody specificity. Mol. Cell 59, 502–511 (2015).
Dickson, B. M., Cornett, E. M., Ramjan, Z. & Rothbart, S. B. ArrayNinja: an open source platform for unified planning and analysis of microarray experiments. Methods Enzymol. 574, 53–77 (2016).
Cornett, E. M., Dickson, B. M. & Rothbart, S. B. Analysis of histone antibody specificity with peptide microarrays. J. Vis. Exp. 126, 2017 (2017).
Jung, H. R., Pasini, D., Helin, K. & Jensen, O. N. Quantitative mass spectrometry of histones H3.2 and H3.3 in Suz12-deficient mouse embryonic stem cells reveals distinct, dynamic post-translational modifications at Lys-27 and Lys-36. Mol. Cell. Proteomics 9, 838–850 (2010).
Fischle, W., Wang, Y. & Allis, C. D. Binary switches and modification cassettes in histone biology and beyond. Nature 425, 475–479 (2003).
Cao, R. et al. Role of hPHF1 in H3K27 methylation and Hox gene silencing. Mol. Cell. Biol. 28, 1862–1872 (2008).
Acknowledgements
This work was supported by the following funding sources: R00GM113019 (S.Z.J.), R01AI148416 (S.Z.J.), AAI Intersect Award (S.Z.J.), R01GM040922 (C.D.A.), R01GM115882 (K.-J.A.), R01AI118891 (B.A.G.), R01CA196539 (B.A.G.), CIPSM (S.B.H.), TRR81/Project A15 (S.B.H.), R35GM124736 (S.B.R.), NIH training grant 5T32AI134632 (A.W.D.), Lymphoma Research Foundation fellowship (A.M.P.), National Natural Science Foundation of China (91753203 and 31725014) and the National Key R&D Program of China (2016YFA0500700) (H.L.). We thank the staff members at beamline BL17U of the Shanghai Synchrotron Radiation Facility and the China National Center for Protein Sciences Beijing for providing facility support; J. Zinder for contributing the SETD2-pETduet-smt3 construct; C. Lu and S. Sidoli (laboratory of B.A.G.) for H3.3 peptide analysis; members of Weill Cornell Applied Bioinformatics Core, D. Betel, P. Zumbo, F. Dundar and L. Skrabanek for suggestions and assistance with bioinformatics; A. Soshnev for help with figures; J. Sun, N. Adams and E. Santosa for assistance with isolation of primary NK cells; and J. Cubillos-Ruiz and P. Giovanelli for BMDCs. We thank R. Niec, B. Sleckman, J. Tyler, J. Blenis, S. Rafii, J. Lis, S. Smale and G. Almouzni for discussions and input and M. Keogh (Epicypher) for developing the H3.3S31ph dNuc reagent.
Author information
Authors and Affiliations
Contributions
L.E.R., C.D., A.W.D., J.Q.C., A.M.P., A.R. and D.J.A. all contributed equally to this study as ‘co-second authors’. A.A. and S.Z.J. conceived and initiated the study in the laboratory of C.D.A. and completed the study in the laboratory of S.Z.J., with L.E.R., C.D., A.W.D., J.Q.C., D.J.A,, A.M.P., A.R. and T.K. performing biochemical, cellular and epigenomic experiments and analysing data supervised by S.Z.J. A.A. and S.Z.J. also performed biochemical, cellular and epigenomic experiments and analysed data. H.L., S.Y. and Y.Y. conceived and performed structural, binding and modelling studies. T.P. assisted A.A. with nucleosome assembly and enzymatic assays. S.T. and E.K. performed neuron experiments. M.W., K.-J.A., S.Y. and A.A. performed nucleosome assembly and HMTase assays. J.H. and S.B.R. performed histone peptide array antibody testing. T.A. and S.B.H. developed and tested the H3.3 antibody. S.L. performed mass spectrometry studies supervised by B.A.G. S.Z.J. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Sankar Ghosh and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Determination of anti-H3.3S31ph antibody specificity.
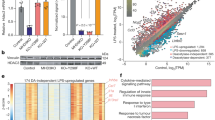
a, Alignment showing H3.1, H3.2 and H3.3 and the differing amino acids in the core and tail. b, Quantitative mass spectrometry analysis of H3.3S31ph (left) and total H3.3 protein (right), in resting (0 min) and bacterial LPS-stimulated (60 min) mouse BMDMs. c, Immunoblot with acid-extracted histones from asynchronous growing (A) or nocodazole-arrested mitotic (M) HeLa cells using bleeds from three rabbits immunized with H3.3S31ph peptides. Bleeds from rabbits 1 and 2 show a signal of the molecular mass of histone H3 only with mitotic samples. d, Peptide competition experiment to determine antibody-specificity. Asynchronous or mitotic histones were separated by SDS–PAGE gels and blotted onto PVDF membranes. H3.3S31ph antibody from rabbit 1 was pre-incubated with diverse peptides or without any peptide, as indicated, before adding it to the PVDF membrane. Staining with anti-H3 antibody shows equal loading. e, Left, immunoblot with acid-extracted histones from asynchronous or mitotic histones separated by 2D Triton-acid urea gels that allow a separation of histone variants due to charge and amino acid differences. The bleed from rabbit 1 shows a signal of the size of H3.3. Right, Coomassie blue staining of the gel and staining of the membrane with anti-H3 served as loading control. f, Deconvolved immunofluorescence microscopy images of asynchronously growing HeLa cells co-stained with DAPI (DNA, blue), anti-H3.3S31ph (rabbit 1, green) and anti-H3S10ph (marker of mitotic cells, red). Merged picture is shown on the right. Note that only mitotic cells, as apparent from stronger DAPI-staining and apparent H3S10ph signal, are H3.3S31ph positive. g, Deconvolved image of chromosome spread from mitotic HeLa cells co-stained with DAPI (blue) and anti-H3.3S31ph (rabbit 1, green). Notice the stronger staining of H3.3S31ph at peri-centromeric regions, as has been shown previously. Original magnifications, ×20 (f) and ×100 (g). h, Cell cycle analysis of BMDM by FACS using DAPI and H3S10ph, with mitotic index gate shown, indicating post-mitotic nature of BMDMs.
Extended Data Fig. 2 Histone peptide array-based specificity profiles for antibodies against H3.3S31ph, H3K36me2 and H3K36me3.
Scatter plots showing signal intensities obtained from each of the indicated antibodies on two separate peptide-arrays. Shown in certain graphs is the relative abundance of the indicated unmodified or modified histone species compared to any species of H3.3S31ph detected by quantitative mass spectrometry in bacterial LPS-stimulated (60 min) mouse BMDMs. Both antibodies recognizing H3K36me and the H3.3S31ph antibody highlighted in green were used.
Extended Data Fig. 3 H3.3S31ph is deposited in the gene body of response genes but not constitutively expressed genes.
a–c, Additional examples of H3 PTMs including H3.3S31ph (as in Fig. 2) at the LPS-induced genes Bcl2a1b, Ccl4, Cxcl2, Dusp1 and Myc (a), the constitutively expressed (housekeeping) genes Gapdh, Rps2, Tbp, Tubb5 and Tubb6 (b), and across the gene-dense chromosome 11 chemokine locus (>1 Mb) containing LPS-induced genes Slfn2, Ccl9, Ccl6, Ccl3 and Ccl4 (c).
Extended Data Fig. 4 H3.3S31ph requires transcription elongation.
a, Comparative pie charts showing percentage of ChIP–seq reads for H3.3S31ph (top) and H3S28ph (bottom) in the indicated genomic regions. We called H3.3S31ph peaks and find that they associated with only 961 genes with 83.72% of peaks falling within gene bodies. By comparison, most H3S28ph peaks fall in promoters and intergenic regions. Given the selective gene body localization of H3.3S31ph, we ranked all annotated genes in the genome by H3.3S31ph ChIP signal density (TSS–TES) in resting and stimulated macrophages. This analysis shows that many more genes acquire high-density H3.3S31ph after stimulation compared with resting cells (Fig. 1e), which is consistent with our mass spectrometry analysis and other global analyses of H3.3S31ph levels. In addition, several of the top-ranked genes (note, by density, not fold change) are prominent LPS-induced genes, including Tnfaip3, Tnf, Il1a and Plk2 (Fig. 1e), all among the top-ranked peaks (Supplementary Table 2). b, We defined a threshold for the top 1% of genes (genome wide) by H3.3S31ph TSS-TES density in stimulated macrophages (167 genes) and find considerable overlap with genes annotated to H3.3S31ph peaks (961 genes). c, Comparison of Gene Ontology (GO) analysis for the top 1% of genes and ‘peak’ genes reveals extensive similarities of these two independent analyses with three of the top five GO categories shared and reflecting the stimulation-induced nature of genes featuring H3.3S31ph: ‘response to stress’, ‘immune system process’, and ‘cellular response to chemical stimulus’ (Supplementary Table 3). d, The H3.3S31ph chromatin state was compared with other ‘active’ chromatin states including H3K27ac, H3K36me3 and H3S28ph as they relate to stimulation-induced gene expression. Analysis reveals that genes with H3.3S31ph are highly enriched for stimulation-induced genes by RNA-seq, especially primary response genes, NF-κB targets and MAPK-dependent genes53,54 (Extended Data Fig. 5). e, H3.3S31ph ChIP–seq read densities of LPS-induced genes in LPS-stimulated BMDMs (60 min) after pre-treatment with FVP, CPT and ETO compared with DMSO treatment (for comparison with top 1% H3.3S31ph target genes in Fig. 2b). f, ChIP–seq average profiles of H3.3S31ph in LPS-stimulated BMDMs after pre-treatment with DMSO, FVP, CPT or ETO for 60 min. g, Box plots showing H3.3S31ph ChIP–seq read densities in LPS-stimulated BMDMs (60 min) after pre-treatment with DMSO (left) and FVP (right). Comparisons are made between NF-κB target genes and LPS-induced genes, the top 1% of H3.3S31ph targets, primary and secondary response (PRG and SRG, respectively) genes, MAPK- and IRF3-dependent genes, and different sub-groups of PRG (classes A–D) and of SRG (classes E, F)54. h, ChIP–seq tracks of H3.3S31ph in LPS-stimulated macrophages (60 min) after pre-treatment with FVP, CPT and ETO (related to Fig. 2a).
Extended Data Fig. 5 IKKα activity drives H3.3S31ph and is enriched at NF-κB target genes.
a, H3.3S31ph ChIP–seq read density fold change (0 to 60 min) for different gene sets54. b, H3.3S31ph ChIP–seq read density fold change (0 to 60 min) for genes sets defined by transcription kinetics (A, fastest/transient; F, slowest). Enrichment statistics for NF-κB-associated motifs for CpG-rich (CpG) and CpG-low (LCG) genes from each category are shown. c, Average gene H3.3S31ph ChIP–seq profiles of LPS-stimulated BMDMs (60 min) in the indicated gene categories54. d, RT–qPCR tracks for Chk1, Tnf and Cxcl2 (left), and western blot for H3.3S31ph (right) in LPS-stimulated mouse BMDMs after transduction with shRNA scrambled control (shSC) and three shRNAs targeting Chk1 (shCHK1). e, RT–qPCR tracks for Chk1, Tnf and Cxcl2 (left), and western blot for H3.3S31ph (right) in LPS-stimulated mouse BMDMs following transfection with siRNA non-target control (siNT) and siRNA against Chk1 (siCHK1) f, ChIP–seq tracks of H3.3S31ph in LPS-stimulated mouse BMDMs after pre-treatment with DMSO and 5 mM CHK1 inhibitor (Chk1i). g, RT–qPCR for Tnf (left) and Cxcl1 (right) in resting and LPS-stimulated macrophages (30 or 60 min) after pre-treatment with DMSO and the IKK inhibitors IKK-16 (1.5 μM) and ACHP (10 μM). h, Western blot analysis of H3.3S31ph in resting and LPS-stimulated BMDMs (30 or 60 min)) transduced with shRNA scrambled control (shSCR) and shRNA targeting Ikka. i, ChIP–seq tracks of IKKα and H3.3S31ph in resting and LPS-stimulated BMDMs (60 min) for Tnfaip3, Ccl4, Cxcl2 and Il1a. H3.3S31ph ChIPs were obtained after pre-treatment of BMDMs with DMSO and IKK-16 (1.5 μM) and after transduction with shRNA scrambled control (shSC) and shRNA against IKKα. *P < 0.05; **P < 0.005; ***P < 0.0005; ****P < 0.0001, Student’s t-test.
Extended Data Fig. 6 H3.3S31ph and other H3 PTMs at response genes and after stimulation.
a, ChIP–seq density fold change comparing the set of all genes to RNA-seq-defined LPS-stimulated genes for H3.3, K27ac, K36me2, K36me3, S28ph and S31ph. b, Average gene profiles shown here are H3K27ac, H3.3S31ph, H3K36me2, H3K36me3, H3S28ph and H3.3) comparing RNA-seq-defined LPS-induced genes before and after stimulation. c, Cumulative distribution function plots for H3K27ac H3K36me2, H3K36me3, H3S28ph, H3.3 and H3.3S31ph reveal selective role of H3.3S31ph compared with the ubiquitous role of H3K36me3. ***P < 0.0001, Student’s t-test.
Extended Data Fig. 7 The H3K36me3 methyltransferase SETD2 is stimulated by H3.3S31ph.
a, Quantitative measurements of three independent experiments (integrated fluorescence intensity) of SETD2 SET-domain HMT assays on unmodified (H3.3un) and H3.3S31ph semi-synthetic dNucs. Error bars in represent the range of three independent experiments. b, Sequence alignment of SETD2 in different species, highlighting the conserved (except in S. cerevisiae) residues K1600 and K1673. c, Sequence alignment of different H3K36 methyltransferases, highlighting the specificity of residues K1600 and K1673 for SETD2. d, Western blot analysis for H3K36me3 in HMT assays with SETD2 SET-domain mutants K1600E and K1673E on H3.3wt and H3.3S31E rNucs. With reduced mutant enzyme activity, enzyme concentration was increased to best visualize the ratio of H3.3wt-to-H3.3S31E activity. e–g, HMT activity assays (SAH accumulation) performed with wild-type (e, f) and K1600A/1673A (K/A) K1600E/K1673E (K/E) double mutant (g). h, Validation of siRNA knockdown for Setd2 using two RT–PCR primers. NT, non-targeting control. i, Western blot for H3K36me3 as a surrogate of SETD2 activity. j, RT–qPCR for the LPS-induced genes Tnf, Cxcl2, Plk2 and Tbp (constitutively expressed control) at 0, 30 or 60 min after LPS stimulation of BMDMs transfected (48 h before) with siRNAs against Setd2 or non-targeting controls. *P < 0.05; **P < 0.01; ***P < 0.001; #P = 0.07, 0.06, for Tnf, Plk2, respectively; Student’s t-test.
Extended Data Fig. 8 Broad potential for H3.3S31ph to regulate histone reader activity.
a, Modelling-based predictions (KDM6B38) and modelled visualizations of established interaction features (PHF139; ZMYND118) of reader interactions dually modified H3.3S31phK36me3. The structure of ZMYND11 PWWP domain bound to H3.3K36me3 peptide and modelling of ZMYND11 PWWP domain with the H3.3S31ph peptide are adapted from ref. 8. S31ph diameter is beyond the PWWP pocket capacity and the steric clashes between H3.3S31ph and PWWP are shown as red plates. H3.3 peptide (yellow sticks), ZMYND11-PWWP (light grey), and residues of PWWP having steric clashes with H3.3S31 are shown as blue sticks (E251, F262 and N266). Another example of accommodation of H3.3S31ph may be the H3K27me3 demethylase KDM6B. Repressive H3K27me3 is abundant on H3.372 and has been shown to have a role in inflammatory gene expression36,37, albeit at later kinetics than studied here. Modelling H3.3S31ph with an existing structure for JMJD338 reveals favourable interactions within an arginine-rich pocket of JMJD3 (R1246 and R1272) surrounding the location of H3.3S31ph (much like SETD2). In contrast to these predictions of augmented recruitment, we considered that the location of H3.3S31ph might act to eject ‘readers’ of proximal H3K36me3, extending the concept of the histone ‘methyl/phos switch’73 into the gene body and co-transcriptional regulation of chromatin. In this context, it has been shown that H3.3S31ph reduces the binding affinity of PHF1 Tudor domain for H3K36me3 by sevenfold39. Modelling H3.3S31phK36me3 with PHF1 Tudor reveals surrounding PHF1 acidic residues (E66, D67 and D68) and 2.5 Å proximity (E66). Given its important role in PRC2 activity40,74, PHF1 ejectionfrom H3K36me3 by H3.3S31ph could have implications for transitions between H3K36me3 and H3K27me3 chromatin. b, Complete ITC data representation including μcal s−1 and time (matching Fig. 4b).
Extended Data Fig. 9 ZMYND11 regulation of LPS-induced genes.
a, b, Slope graph (a) and rank plot (b) depicting differences in ZMYND11 ChIP densities between resting and LPS-stimulated (60 min) BMDMs. c, Cumulative distribution of read density of different sets of genes in S31ph ChIP data. S31 top 1% genes overall have more density so it is shifted to right compared to all genes. Genes extracted from top peaks of IKK, ZMYND11 and S31ph ChIP tends to have more enriched read density distribution over when we look at entire genes. Gene numbers are as follows: 44 IKK genes, 176 op 1% genes and 851 S31ph peak genes. d, ChIP–seq read density tracks of ZMYND11, H3.3S31ph and H3K36me3 in LPS-stimulated BMDMs (0 and 60 min) for Btg2, Junb, Marcks, Zfp36 (LPS-induced genes) and Tubb6 (constitutively expressed gene). e, Left, RT–qPCR for Zmynd11, Junb and Nfkbia after LPS stimulation of BMDMs transfected (48 h before) with siRNA non-target (NT) control and siRNA against Zmynd11. Right, western blot analysis of ZMYND11 transduced with shRNA scrambled control, and shRNA targeting Zmynd11. f, Box plots depicting differences in ZMYND11 ChIP densities between resting and LPS-stimulated (60 min) BMDMs. **P < 0.005; ***P < 0.0005; ****P < 0.0001; Student’s t-test.
Extended Data Fig. 10 Characterization of H3.3 mutant RAW264.7 cell lines.
a, Western blot for H3.3 comparing wild-type, vector control (VC), HYPO and DKO RAW264.7 cell lines, membrane was stained with direct blue (DB) for equal loading. b, c, RNA-seq scatter (volcano) plot analysis, log2-transformed fold change and −log10(FDR) of DKO compared to wild-type (b), and HYPO compared to wild-type (c) RAW264.7 cells at 120 min. d, Ratio of RNA-seq fold change (log2-transformed) for HYPO or DKO compared with wild-type cells after LPS stimulation (60 and 120 min) for all LPS-induced genes (top) and for the intersection of top H3.3S31ph genes and LPS-induced genes (bottom). ***P < 0.0001, lower-tailed one-sample t-test (distribution below zero). e, Heat map of fold change (log2-transformed) for top H3.3S31ph genes among LPS-induced genes (left, 60 min; right, 120 min) with control constitutively expressed genes below. RNA-seq analysis was performed with two biological replicates per condition. f, Time course plots of mean RNA-seq expression (RPKM) from two experiments at time points 0, 60 and 120 min after LPS-stimulation for experiments performed in wild-type, HYPO and DKO RAW264.7cell lines at LPS-induced genes and at top H3.3S31ph genes among LPS-induced genes. g, Time course plots of mean RNA-seq expression (RPKM) from two experiments at time points 0, 60 and 120 min after LPS-stimulation for experiments performed in wild-type, HYPO and DKO RAW247.6 cell lines at LPS-induced genes Myc, Ccl9, Slfn2, Tnfaip3, Ccl4, Plk2 and constitutively expressed genes Tubb5 and Tbp. h, RT–qPCR for the viral expression of H3.3 transgene, wild-type, S31A or S31E in RAW macrophages. i, RT–qPCR for Ccl4 expression in a time course of stimulated DKO RAW264.7 cells rescued with wild-type H3.3 or S31A or S31E mutants.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 and Supplementary Tables S1-S4. Supplementary Figure 1: Uncropped western blots. Format: eighteen image display items. Table S1: Histone post translational modification mass spectrometry data in resting and stimulated BMDM. See methods for details. Format: spreadsheet and graph. Table S2: List of top ranked H3.3S31ph peaks, BMDM, 60 min LPS. Genes were ranked by H3.3S31ph ChIPseq tag density from transcription start site to transcription end site. Format: gene list in rank order. Table S3: SETD2:H3.3S31ph structure characteristics. Data collection and refinement statistics. Format: text table. Table S4: Intersection of H3.3S31ph top 1% genes and H3.3S31ph peak-annotated genes. Format: gene list in alphabetical order.
Rights and permissions
About this article
Cite this article
Armache, A., Yang, S., Martínez de Paz, A. et al. Histone H3.3 phosphorylation amplifies stimulation-induced transcription. Nature 583, 852–857 (2020). https://doi.org/10.1038/s41586-020-2533-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2533-0
This article is cited by
-
HIRA vs. DAXX: the two axes shaping the histone H3.3 landscape
Experimental & Molecular Medicine (2024)
-
H3.3 contributes to chromatin accessibility and transcription factor binding at promoter-proximal regulatory elements in embryonic stem cells
Genome Biology (2023)
-
(B)On(e)-cohistones and the epigenetic alterations at the root of bone cancer
Cell Death & Differentiation (2023)
-
Targeting HECTD3-IKKα axis inhibits inflammation-related metastasis
Signal Transduction and Targeted Therapy (2022)
-
SMYD5 catalyzes histone H3 lysine 36 trimethylation at promoters
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.