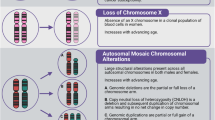
Abstract
The extent to which the biology of oncogenesis and ageing are shaped by factors that distinguish human populations is unknown. Haematopoietic clones with acquired mutations become common with advancing age and can lead to blood cancers1,2,3,4,5,6,7,8,9,10. Here we describe shared and population-specific patterns of genomic mutations and clonal selection in haematopoietic cells on the basis of 33,250 autosomal mosaic chromosomal alterations that we detected in 179,417 Japanese participants in the BioBank Japan cohort and compared with analogous data from the UK Biobank. In this long-lived Japanese population, mosaic chromosomal alterations were detected in more than 35.0% (s.e.m., 1.4%) of individuals older than 90 years, which suggests that such clones trend towards inevitability with advancing age. Japanese and European individuals exhibited key differences in the genomic locations of mutations in their respective haematopoietic clones; these differences predicted the relative rates of chronic lymphocytic leukaemia (which is more common among European individuals) and T cell leukaemia (which is more common among Japanese individuals) in these populations. Three different mutational precursors of chronic lymphocytic leukaemia (including trisomy 12, loss of chromosomes 13q and 13q, and copy-neutral loss of heterozygosity) were between two and six times less common among Japanese individuals, which suggests that the Japanese and European populations differ in selective pressures on clones long before the development of clinically apparent chronic lymphocytic leukaemia. Japanese and British populations also exhibited very different rates of clones that arose from B and T cell lineages, which predicted the relative rates of B and T cell cancers in these populations. We identified six previously undescribed loci at which inherited variants predispose to mosaic chromosomal alterations that duplicate or remove the inherited risk alleles, including large-effect rare variants at NBN, MRE11 and CTU2 (odds ratio, 28–91). We suggest that selective pressures on clones are modulated by factors that are specific to human populations. Further genomic characterization of clonal selection and cancer in populations from around the world is therefore warranted.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
A table for mosaic events detected in the current study is available as Supplementary Data 1. The BBJ genotype is available from the Japanese Genotype-phenotype Archive (JGA; http://trace.ddbj.nig.ac.jp/jga/index_e.html) under accession code JGAD00000000123. Individual-level linkage of mosaic events can be provided by the BBJ project upon request (https://biobankjp.org/english/index.html).
Code availability
All computational codes are available upon request from the corresponding authors (although they are not immediately portable to other computing environments). A standalone software implementation (MoChA) of the algorithm used to call mCAs is available at https://github.com/freeseek/mocha.
References
Jacobs, K. B. et al. Detectable clonal mosaicism and its relationship to aging and cancer. Nat. Genet. 44, 651–658 (2012).
Laurie, C. C. et al. Detectable clonal mosaicism from birth to old age and its relationship to cancer. Nat. Genet. 44, 642–650 (2012)
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Machiela, M. J. et al. Characterization of large structural genetic mosaicism in human autosomes. Am. J. Hum. Genet. 96, 487–497 (2015).
Vattathil, S. & Scheet, P. Extensive hidden genomic mosaicism revealed in normal tissue. Am. J. Hum. Genet. 98, 571–578 (2016).
Young, A. L., Challen, G. A., Birmann, B. M. & Druley, T. E. Clonal haematopoiesis harbouring AML-associated mutations is ubiquitous in healthy adults. Nat. Commun. 7, 12484 (2016).
Forsberg, L. A., Gisselsson, D. & Dumanski, J. P. Mosaicism in health and disease — clones picking up speed. Nat. Rev. Genet. 18, 128–142 (2017).
Abelson, S. et al. Prediction of acute myeloid leukaemia risk in healthy individuals. Nature 559, 400–404 (2018).
Loh, P.-R. et al. Insights into clonal haematopoiesis from 8,342 mosaic chromosomal alterations. Nature 559, 350–355 (2018).
Zhou, W. et al. Mosaic loss of chromosome Y is associated with common variation near TCL1A. Nat. Genet. 48, 563–568 (2016).
Hinds, D. A. et al. Germ line variants predispose to both JAK2 V617F clonal hematopoiesis and myeloproliferative neoplasms. Blood 128, 1121–1128 (2016).
Wright, D. J. et al. Genetic variants associated with mosaic Y chromosome loss highlight cell cycle genes and overlap with cancer susceptibility. Nat. Genet. 49, 674–679 (2017).
Nagai, A. et al. Overview of the BioBank Japan project: study design and profile. J. Epidemiol. 27, S2–S8 (2017).
Loh, P.-R., Genovese, G. & McCarroll, S. A. Monogenic and polygenic inheritance become instruments for clonal selection. Nature https://doi.org/10.1038/s41586-020-2430-6 (2020).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Iwanaga, M., Watanabe, T. & Yamaguchi, K. Adult T-cell leukemia: a review of epidemiological evidence. Front. Microbiol. 3, 322 (2012).
Tamura, K. et al. Chronic lymphocytic leukemia (CLL) is rare, but the proportion of T-CLL is high in Japan. Eur. J. Haematol. 67, 152–157 (2001).
Li, Y., Wang, Y., Wang, Z., Yi, D. & Ma, S. Racial differences in three major NHL subtypes: descriptive epidemiology. Cancer Epidemiol. 39, 8–13 (2015).
Landau, D. A. et al. Mutations driving CLL and their evolution in progression and relapse. Nature 526, 525–530 (2015).
Puente, X. S. et al. Non-coding recurrent mutations in chronic lymphocytic leukaemia. Nature 526, 519–524 (2015).
Iwai, M. et al. Expression and methylation status of the FHIT gene in acute myeloid leukemia and myelodysplastic syndrome. Leukemia 19, 1367–1375 (2005).
Schmitz, R. et al. TNFAIP3 (A20) is a tumor suppressor gene in Hodgkin lymphoma and primary mediastinal B cell lymphoma. J. Exp. Med. 206, 981–989 (2009).
Liu, Y. et al. The genomic landscape of pediatric and young adult T-lineage acute lymphoblastic leukemia. Nat. Genet. 49, 1211–1218 (2017).
Zink, F. et al. Clonal hematopoiesis, with and without candidate driver mutations, is common in the elderly. Blood 130, 742–752 (2017).
Knudson, A. G. Jr. Mutation and cancer: statistical study of retinoblastoma. Proc. Natl Acad. Sci. USA 68, 820–823 (1971).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Okada, Y. et al. Deep whole-genome sequencing reveals recent selection signatures linked to evolution and disease risk of Japanese. Nat. Commun. 9, 1631 (2018).
Kilpivaara, O. et al. A germline JAK2 SNP is associated with predisposition to the development of JAK2 V617F-positive myeloproliferative neoplasms. Nat. Genet. 41, 455–459 (2009).
Jones, A. V. et al. JAK2 haplotype is a major risk factor for the development of myeloproliferative neoplasms. Nat. Genet. 41, 446–449 (2009).
Olcaydu, D. et al. A common JAK2 haplotype confers susceptibility to myeloproliferative neoplasms. Nat. Genet. 41, 450–454 (2009).
Lee, J. H. & Paull, T. T. ATM activation by DNA double-strand breaks through the Mre11–Rad50–Nbs1 complex. Science 308, 551–554 (2005).
Dewez, M. et al. The conserved Wobble uridine tRNA thiolase Ctu1–Ctu2 is required to maintain genome integrity. Proc. Natl Acad. Sci. USA 105, 5459–5464 (2008).
Adzhubei, I., Jordan, D. M. & Sunyaev, S. R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. 76, 7.20.1–7.20.41 (2013).
Sim, N. L. et al. SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 40, W452–W457 (2012).
DeAntoni, A., Sala, V. & Musacchio, A. Explaining the oligomerization properties of the spindle assembly checkpoint protein Mad2. Phil. Trans. R. Soc. Lond. B 360, 637–648 (2005).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Jaiswal, S. et al. Clonal hematopoiesis and risk of atherosclerotic cardiovascular disease. N. Engl. J. Med. 377, 111–121 (2017).
Hirata, M. et al. Overview of BioBank Japan follow-up data in 32 diseases. J. Epidemiol. 27, S22–S28 (2017).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Staaf, J. et al. Normalization of Illumina Infinium whole-genome SNP data improves copy number estimates and allelic intensity ratios. BMC Bioinformatics 9, 409 (2008).
Loh, P.-R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443–1448 (2016).
Kanai, M. et al. Genetic analysis of quantitative traits in the Japanese population links cell types to complex human diseases. Nat. Genet. 50, 390–400 (2018).
Loh, P.-R., Palamara, P. F. & Price, A. L. Fast and accurate long-range phasing in a UK Biobank cohort. Nat. Genet. 48, 811–816 (2016).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Ma, C., Blackwell, T., Boehnke, M. & Scott, L. J. Recommended joint and meta-analysis strategies for case-control association testing of single low-count variants. Genet. Epidemiol. 37, 539–550 (2013).
Acknowledgements
We thank the staff of the BBJ for collecting and managing samples and clinical information. This study was funded by the BioBank Japan project, which was supported by the Ministry of Education, Culture, Sports, Sciences and Technology of the Japanese Government and AMED under grant numbers 17km0305002 and 18km0605001. This research was conducted using the UK Biobank Resource under application no. 19808. P.-R.L. was supported by NIH grant DP2 ES030554, a Burroughs Wellcome Fund Career Award at the Scientific Interfaces, the Next Generation Fund at the Broad Institute of MIT and Harvard, a Glenn Foundation for Medical Research and AFAR Grants for Junior Faculty award, and a Sloan Research Fellowship.
Author information
Authors and Affiliations
Contributions
C.T., P.-R.L. and Y.K. conceived the study design. P.-R.L. and Y.K. supervised the project. C.T. and P.-R.L. analysed the data. A.S. and K.Y. conducted functional analyses. M.A. and K.I. contributed to the construction of the Japanese reference panel for genotype imputation. Y. Momozawa, K.M., Y. Murakami and M.K. contributed to the generation of the BBJ data. C.T., S.A.M., P.-R.L. and Y.K. wrote the manuscript. All the authors critically reviewed the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Paul Scheet, George Vassiliou and John Witte for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Age and sex of the carriers of mosaic event types.
Mean age and sex of carriers of specific mCA types (defined by chromosome and copy number) with at least 100 carriers in the 179,417 participants. Marker sizes are proportional to mCA frequencies. Data are mean ± s.e.m. Numeric data are provided in Supplementary Table 7.
Extended Data Fig. 2 Comparable chromosomal coverage by heterozygous genotypes in the BBJ and UKB data.
Average numbers of heterozygous genotyped sites (averaged across individuals) in each 1-Mb region of the genome for the BBJ and UKB genotyping arrays. hets, heterozygous sites.
Extended Data Fig. 3 Similar breakpoint distributions of CN-LOH events in the BBJ and UKB data.
Relative frequencies of estimated CN-LOH breakpoint locations in the BBJ and UKB data. Breakpoints were smoothed over ±2 Mb to enable plotting of frequency curves, which were rescaled to 1.
Extended Data Fig. 4 Quantile–quantile plots of mosaic events with significant associations demonstrate that there is no inflation of association statistics.
Quantile–quantile plots of results for mosaic events with significant associations. Analysis results of Fisher’s exact test (two-sided, nominal P values) using 173,599 participants are shown. We defined the following hits as hit loci: 42–49 Mb at chromosome 1 (1p CN-LOH), 88–94 Mb at chromosome 8 (8q CN-LOH), 92–96 Mb at chromosome 11 (11q CN-LOH), 88–90 Mb at chromosome 16 (16q CN-LOH), 23–26 Mb and 100–103 Mb at chromosome 14 (cis association of 14q CN-LOH), 4–6 Mb at chromosome 9 (9p CN-LOH), 0–2 Mb at chromosome 5 (trans association of 14q CN-LOH) and 1–3 Mb at chromosome 7 (trans association of chromosome 15 gain).
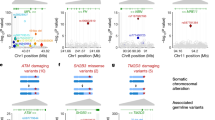
Extended Data Fig. 5 Local plots for cis and trans associations.
a–i, Associations of inherited variants with 8q CN-LOH (a), 11q CN-LOH (b), 16q CN-LOH (c), chromosome 15 gain (d), 1p CN-LOH (e), 9p CN-LOH (f) and 14q CN-LOH (g–i) are shown for regions containing the NBN, MRE11, CTU2, MAD1L1, MPL, JAK2, NEDD8–TINF2, DLK1 and TERT loci, respectively. a–c, e, Loci are rare cis associations. f–h, Loci are common cis associations. d, i, Loci are trans associations. a–d, g–i, Loci are in previously unreported regions. Purple points indicate lead variants. Other variants are colour-coded according to the linkage disequilibrium r2 with lead variants. The TCL1A variant that significantly associated with 14q CN-LOH allelic imbalance is not shown here because it did not significantly associate with 14q CN-LOH risk. Analysis results of Fisher’s exact test (two-sided, nominal P values) using 173,599 participants are shown.
Extended Data Fig. 6 Action of CN-LOH events on rare and common inherited variants.
Schematics show the patterns of selection or elimination of inherited variants by CN-LOH events. Asterisks indicate risk alleles. For the TCL1A locus, which did not significantly associate with the presence of 14q CN-LOH, we depict TCL1A as a gene for which CN-LOH mutations select an allele.
Extended Data Fig. 7 Examples of multiple overlapping CN-LOH clones in a single chromosome.
We identified 185 individuals who carried multiple CN-LOH clones on a single chromosome. a, Multiple clones were observed in at least one individual for all chromosomes except chromosomes 18, 20 and 22. The plots show phased BAF deviations (y axis) as a function of chromosome position (x axis) for the individual with the largest clone per chromosome (among all individuals with multiple CN-LOH clones on that chromosome). Coloured horizontal lines of different colours indicate distinct BAF deviations corresponding to overlapping CN-LOH events. b, The number of participants carrying multiple CN-LOH clones on a single chromosome is shown for each chromosomal arm.
Extended Data Fig. 8 Mortality risk conferred by mosaic chromosomal alterations.
a, Risk of mortality from various causes conferred by presence of an mCA at >1% cell fraction. Leukaemia, malignant lymphoma and multiple myeloma are subdivisions of blood cancer. Cardiovascular mortality includes deaths from coronary artery disease and ischaemic stroke. b, Risk of leukaemia mortality conferred by specific mCAs (grouped by chromosomal location and copy-number change) reaching Bonferroni-corrected significance. c, Risk of leukaemia mortality conferred by mosaic status stratified by mosaic cell fraction. d, Risk of leukaemia mortality conferred by mosaic status stratified by mosaic cell fraction and number of mosaic events detected (one versus two or more). All analyses were restricted to individuals with no previous cancer diagnosis and were corrected for age, sex, smoking status and genotyping array (Methods). Data are hazard ratio or odds ratio and 95% confidence intervals. Numeric data are provided in Supplementary Tables 24–27. Results using 86,546 participants are indicated. Cox proportional hazard models (two-sided) were used for a, b and d. A Cochran–Mantel–Haenszel test was used for c.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-8, which include descriptions of detailed methods and data interpretation, Supplementary References and Supplementary Tables 1-28.
Supplementary Data
This file contains a table of anonymized individual-level mosaic events in detail.
Rights and permissions
About this article
Cite this article
Terao, C., Suzuki, A., Momozawa, Y. et al. Chromosomal alterations among age-related haematopoietic clones in Japan. Nature 584, 130–135 (2020). https://doi.org/10.1038/s41586-020-2426-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2426-2
This article is cited by
-
Clonal haematopoiesis and dysregulation of the immune system
Nature Reviews Immunology (2023)
-
A pan-tissue survey of mosaic chromosomal alterations in 948 individuals
Nature Genetics (2023)
-
Lymphoid clonal hematopoiesis: implications for malignancy, immunity, and treatment
Blood Cancer Journal (2023)
-
Mutation rates and fitness consequences of mosaic chromosomal alterations in blood
Nature Genetics (2023)
-
Clonal Hematopoiesis: Connecting Aging and Inflammation in Atherosclerosis
Current Atherosclerosis Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.