Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), a novel coronavirus with high nucleotide identity to SARS-CoV and to SARS-related coronaviruses that have been detected in horseshoe bats, has spread across the world and had a global effect on healthcare systems and economies1,2. A suitable small animal model is needed to support the development of vaccines and therapies. Here we report the pathogenesis and transmissibility of SARS-CoV-2 in golden (Syrian) hamsters (Mesocricetus auratus). Immunohistochemistry assay demonstrated the presence of viral antigens in nasal mucosa, bronchial epithelial cells and areas of lung consolidation on days 2 and 5 after inoculation with SARS-CoV-2, followed by rapid viral clearance and pneumocyte hyperplasia at 7 days after inoculation. We also found viral antigens in epithelial cells of the duodenum, and detected viral RNA in faeces. Notably, SARS-CoV-2 was transmitted efficiently from inoculated hamsters to naive hamsters by direct contact and via aerosols. Transmission via fomites in soiled cages was not as efficient. Although viral RNA was continuously detected in the nasal washes of inoculated hamsters for 14 days, the communicable period was short and correlated with the detection of infectious virus but not viral RNA. Inoculated and naturally infected hamsters showed apparent weight loss on days 6–7 post-inoculation or post-contact; all hamsters returned to their original weight within 14 days and developed neutralizing antibodies. Our results suggest that features associated with SARS-CoV-2 infection in golden hamsters resemble those found in humans with mild SARS-CoV-2 infections.
Similar content being viewed by others
Main
SARS-CoV-2 was first detected from a cluster of patients with pneumonia in Wuhan (Hubei province, China) in December 2019. Although 55% of the initial cases were linked to one seafood wholesale market at which wild animals were also sold3, multiple viral (sustained human-to-human transmissibility by symptomatic and pre-symptomatic individuals4) and ecological (extensive domestic and international travel during Chinese Lunar New Year) factors have contributed to the rapid global spread of the virus. The clinical spectrum of patients with coronavirus disease 2019 (COVID-19) is wide; 19% of 44,415 confirmed patients in China progressed to severe and critical illness5, with an estimated 1.4% case fatality risk in symptomatic cases6. There is no approved vaccine or treatment against SARS-CoV-2, and the available interventions—including country lockdowns and social distancing—have severely disrupted the global supply chain and economy.
A suitable animal model is essential for understanding the pathogenesis of this disease and for evaluating vaccine and therapeutic candidates. Previous animal studies on SARS-CoV have suggested the importance of the interaction between the viral spike protein and the host angiotensin-converting enzyme 2 (ACE2) receptor7,8,9,10, as well as age and innate immune status of the animals11,12,13,14 in pathogenesis. As with SARS-CoV, the spike protein of SARS-CoV-2 also uses ACE2 as the entry receptor1. ACE2 is expressed on the surface of alveolar epithelial cells and enterocytes of the small intestine and can be detected in endothelial cells and arterial smooth muscle cells15. SARS-CoV-2 showed good binding to human ACE2 but limited binding to mouse ACE21, which has limited the use of inbred mice for research. Macaques and transgenic ICR mice that express the human ACE2 receptor have been shown to be susceptible to infection with SARS-CoV-216,17,18; however, there is limited availability of these animal models. Cynomolgus macaques and rhesus macaques challenged with SARS-CoV-2 showed pneumonia with limited17 and moderate18 clinical signs, respectively. The transgenic mice challenged with SARS-CoV-2 showed pneumonia, moderate weight loss and no apparent histological changes in nonrespiratory tissues16. Previously generated transgenic mice that express the human ACE2 receptor have been reported to support the replication of SARS-CoV in the epithelial cells of the airway, but were associated with neurological-related mortality owing to high ACE2 expression in the brain7,8,9,10.
The golden hamster is a widely used experimental animal model and has previously been reported to support replication of SARS-CoV19,20—but not Middle Eastern respiratory syndrome coronavirus (MERS-CoV)21, which uses the dipeptidyl peptidase 4 (DPP4) protein as the main receptor for viral entry. A previous study of the SARS-CoV Urbani strain in 5-week-old golden hamsters showed robust viral replication, with peak viral titres detected in the lungs at 2 days post-inoculation (dpi) followed by rapid viral clearance by 7 dpi, but without weight loss or evidence of disease in the inoculated hamsters20. A follow-up study that reported the testing of different strains of SARS-CoV in golden hamsters found differences in virulence between these strains; lethality was reported in hamsters challenged with the Frk-1 strain, which differs from the nonlethal Urbani strain by an L1148F substitution in the S2 domain19. Hamsters are permissive for infection by other respiratory viruses—including human metapneumovirus22, human parainfluenza virus 323 and influenza A virus—and may support influenza transmission by contact or airborne routes24,25. Alignment of the ACE2 proteins of human, macaque, mouse and hamster suggest that the spike protein of SARS-CoV-2 may interact more efficiently with hamster ACE2 than mouse ACE2 (Extended Data Fig. 1). Here we evaluate the pathogenesis and contact transmissibility of SARS-CoV-2 in 4–5-week-old male golden hamsters.
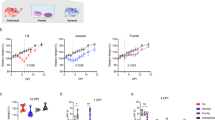
Hamsters were infected intranasally with 8 × 104 50% tissue culture infective dose (TCID50) of SARS-CoV-2 (BetaCoV/Hong Kong/VM20001061/2020; GISAID identifier EPI_ISL_412028), isolated in Vero E6 cells from the nasopharynx aspirate and throat swab of a patient from Hong Kong with a confirmed case of COVID-19. At 2, 5 and 7 dpi, the nasal turbinate, brain, lungs, heart, duodenum, liver, spleen and kidney were collected to monitor viral replication and histopathological changes. Peak viral load in the lungs was detected at 2 dpi and decreased at 5 dpi; no infectious virus was detected at 7 dpi despite of the continued detection of high copies of viral RNA (Fig. 1a). Infectious viral load was significantly different between 2 and 7 dpi (P = 0.019, Dunn’s multiple comparisons test), but the RNA copy number was not (P = 0.076). No infectious virus was detected in the kidney, although low copies of viral RNA were detected at 2 and 5 dpi (Fig. 1b).
a, Infectious viral load (log10(TCID50 per ml)) and viral RNA (log10(RNA copies per ml)) detected in the lungs of hamsters challenged with SARS-CoV-2 (n = 3) at 2, 5 and 7 dpi. b, Infectious viral load and viral RNA detected in the kidney of hamsters challenged with SARS-CoV-2 (n = 3) at 2, 5 and 7 dpi. Individual data points and mean ± s.d. are shown; the detection limit (1.789 log10(TCID50 per ml)) is shown by the dotted line. c, Haematoxylin and eosin (H&E) staining of the lungs of hamsters challenged with SARS-CoV-2 at 2 dpi. d, Detection of SARS-CoV-2 N protein at bronchial epithelial cells (indicated by an arrow) by immunohistochemistry at 2 dpi. e, H&E staining of the lungs at 5 dpi. f, Detection of N protein in pneumocytes with lung consolidation (indicated by an arrow) at 5 dpi. g, H&E staining of the lungs at 7 dpi. h, The lack of detection of N protein in the lungs at 7 dpi. i, H&E staining of nasal turbinate of hamsters challenged with SARS-CoV-2 at 2 dpi. j, Detection of N protein in nasal epithelial cells (arrow on the right) and cells morphologically resembling olfactory neurons (arrow on the left) at 2 dpi. k, H&E staining of duodenum of hamsters challenged with SARS-CoV-2 at 2 dpi. l, Detection of N protein in the duodenum epithelial cells at 2 dpi. The experiment was performed once with 9 hamsters challenged with 8 × 104 TCID50 of SARS-CoV-2, and tissues were collected from 3 hamsters for histopathology examination and immunohistochemistry at each time point. H&E staining and immunohistochemistry performed using tissues from three hamsters showed comparable results; representative images are shown. Scale bars, 200 μm (i, j), 500 μm (c–h, k, l).
Histopathological examination detected an increase in inflammatory cells and consolidation in 5–10% of the lungs at 2 dpi (Fig. 1c, d) and 15–35% of the lungs at 5 dpi (Fig. 1e, f). We observed mononuclear cell infiltrate in areas in which viral antigen was detected at 2 and 5 dpi. Immunohistochemistry for the nucleocapsid (N) protein of SARS-CoV-2 demonstrated viral antigen in the bronchial epithelial cells at 2 dpi (Fig. 1d), with progression to pneumocytes at 5 dpi (Fig. 1f). At 7 dpi, there was an increased consolidation in 30–60% of the lungs (Fig. 1g); however, no viral antigen was detected at this time point (Fig. 1h) and type-2 pneumocyte hyperplasia was prominent (Extended Data Fig. 2a). CD3-positive T lymphocytes were detected in the peribronchial region at 5 dpi, which may facilitate the rapid clearance of the infected cells (Extended Data Fig. 2b). There was moderate inflammatory-cell infiltration in the nasal turbinate (Fig. 1i), and viral antigen was detected in the nasal epithelial cells (Fig. 1j) and in olfactory sensory neurons at the nasal mucosa (Fig. 1j). Infection in the olfactory neurons was further confirmed in cells that express both SARS-CoV-2 N protein and neuron-specific tubulin-β III (Extended Data Fig. 2c). Compared to mock infection (Extended Data Fig. 2d, e), infection with SARS-CoV-2 leads to a reduction in the number of olfactory neurons at the nasal mucosal at 2 dpi (Extended Data Fig. 2f), prominent nasal epithelial attenuation at 7 dpi (Extended Data Fig. 2g), followed by tissue repair at 14 dpi (Extended Data Fig. 2h). Though no inflammation was present (Fig. 1k), viral antigen was detected from the epithelial cells of duodenum at 2 dpi (Fig. 1l). This resembles the detection of replication of SARS-CoV in the epithelial cells of the terminal ileum and colon of patients with SARS-CoV without observing apparent architectural disruption and inflammatory infiltrate26. No apparent histopathological change was observed from the brain, heart, liver and kidney at 5 dpi (Extended Data Fig. 2i–l).
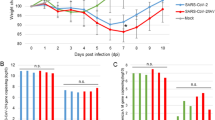
To assess the transmission potential of the SARS-CoV-2 in hamsters, we intranasally inoculated three donor hamsters with 8 × 104 TCID50 of the virus. At 24 hours after inoculation, each donor was transferred to a new cage and cohoused with one naive hamster. Weight changes and clinical signs were monitored daily and nasal washes were collected every other day from donors and contacts, for 14 days. In donors, the peak infectious viral load in nasal washes was detected soon after inoculation and was followed by a rapid decline, although viral RNA was detected continuously for 14 days (Fig. 2a). Hamsters inoculated with SARS-CoV-2 showed their maximal mean weight loss (mean ± s.d., −11.97 ± 4.51%, n = 6) at 6 dpi (Fig. 2b). Transmission from donor to cohoused contact hamsters was efficient, and SARS-CoV-2 was detected from the cohoused hamsters at 1 day post-contact (dpc), and the peak viral load in nasal washes was detected at 3 dpc (Fig. 2c). The total viral load shed in the nasal washes was approximated by calculating the area under the curve for each hamster. The amount of virus shed in the nasal washes of contact hamsters was comparable to that of the donor hamsters (P = 0.1, two-tailed Mann–Whitney test). Contact hamsters showed their maximal mean weight loss (mean ± s.d., −10.68 ± 3.42%, n = 3) at 6 dpc; all hamsters had returned to their original weight by 11 dpc (Fig. 2d). Neutralizing antibodies were detected using 90% plaque reduction neutralization (PRNT90) assay from donor hamsters at 14 dpi (titres at 1:640 for all) and from contact hamsters on 13 dpc (titres at 1:160, 1:320 and 1:160). As viral RNA was continuously detected in the nasal washes of the donor for 14 days (whereas infectious virus titres decreased rapidly), we repeated the experiment and cohoused naive contact with donor hamsters at 6 dpi. A low quantity of viral RNA was detected in the nasal washes in one contact hamster at 3 and 7 dpc, without detection of infectious virus in the nasal washes (Fig. 2e); none of the contact hamsters showed weight loss (Fig. 2f). A PRNT90 assay detected no neutralizing antibody (<1:10) from the contact hamsters at 12 dpc. The results suggest that the donor hamsters inoculated with SARS-CoV-2 have a short communicable period, of less than six days. Onward transmissibility from donor to cohoused contact hamsters was correlated with the detection of infectious virus, but not viral RNA, in the donor nasal washes.
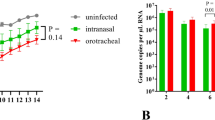
a, Infectious viral load (log10(TCID50 per ml), shown as bars) and viral RNA copy numbers (log10(RNA copies per ml), shown as dots) detected in the nasal washes of donor hamsters (n = 3) inoculated with 8 × 104 TCID50 of SARS-CoV-2. Colour-matched bars and dots represent results from the same hamster. b, Changes in body weight (per cent weight change compared to day 0) of hamsters inoculated with SARS-CoV-2 (n = 9, including 3 donors and 9 hamsters challenged with SARS-CoV-2, described in Fig. 1); individual data points and mean ± s.d. are shown. c, Transmission of SARS-CoV-2 to naive hamsters (n = 3), each of which was cohoused with one inoculated donor at 1 dpi; infectious viral load and viral RNA copy numbers detected in the nasal washes of contact hamsters are shown. d, Changes in body weight (per cent weight change compared to the day of exposure) of contact hamsters (n = 3) cohoused with inoculated donor at 1 dpi. e, Transmission of SARS-CoV-2 to naive hamsters (n = 3), each of which was cohoused with one donor at 6 dpi; infectious viral load and viral RNA copy numbers detected in the nasal washes of contact hamsters are shown. f, Changes in body weight (per cent weight change compared to the day of exposure) of contact hamsters (n = 3) cohoused with inoculated donors at 6 dpi. Direct-contact transmission experiments of cohoused donors and naive contacts at 1 dpi and 6 dpi (of the donor), respectively, were each performed once, each with three pairs of donor:direct contact at 1:1 ratio.
The transmission from donor to cohoused contact hamster may have been mediated by multiple transmission routes. To investigate the transmissibility of SARS-CoV-2 among hamsters via aerosols (a collection of liquid or solid particles suspended in air and may include large droplets and fine droplet nuclei27), donor hamsters and naive aerosol-contact hamsters were placed in two adjacent wire cages for 8 h at 1 dpi (of the donor) (Extended Data Fig. 3). The experiment was performed in three pairs of donor:aerosol-contact hamsters at a 1:1 ratio. The hamsters were single-housed after exposure, and were monitored daily for 14 days. Donor hamsters shed infectious virus in the nasal washes for 6 days, whereas viral RNA could be continuously detected for 14 days (Fig. 3a). Viral RNA was detected in the faecal samples of the donors at 2, 4 and 6 dpi, without detection of infectious virus (Fig. 3b). All donor hamsters showed comparable weight loss (Fig. 3c), as observed previously (Fig. 2b). Transmission via aerosols was efficient, as shown by the fact that infectious virus was detected in the nasal washes from all exposed contact hamsters at 1 dpc, with peak viral loads detected at 3 dpc (Fig. 3d). Viral RNA was continuously detected from the faecal samples of the infected aerosol-contact hamsters for 14 days, although no infectious virus was isolated (Fig. 3e). The aerosol-contact hamsters showed their maximal weight loss (mean ± s.d., −7.72 ± 5,42%, n = 3) at 7 dpc (Fig. 3f). The amount of virus shed in the nasal washes (approximated by area under the curve) of aerosol-contact hamsters was comparable to that of the donor hamsters (P = 0.4, two-tailed Mann–Whitney test). Neutralizing antibodies were detected using PRNT90 assay from the donor hamsters at 16 dpi (titres at 1:320, 1:640 and 1:640) and the contact hamsters at 15 dpc (titres at 1:640 for all). To evaluate the transmission potential of SARS-CoV-2 via fomites, three naive fomite-contact hamsters were each introduced to a soiled cage that had housed one donor between 0 and 2 dpi. The fomite-contact hamsters were single-housed in the soiled cages for 48 hours, and each was transferred to a new cage at 2 dpc (equivalent to 4 dpi of the donor hamster). Viral RNA was detected from different surfaces sampled from the soiled cages used for housing the fomite-contact hamsters, with a low titre of infectious virus detected in the bedding (at 2 dpi), cage side surface (at 4 dpi) and water bottle nozzle (at 4 dpi) (Extended Data Table 1). One out of three fomite-contact hamsters shed infectious virus in the nasal washes starting from 1 dpc, with the peak viral load detected at 3 dpc (Fig. 3g). Viral RNA, but not infectious virus, was detected from the faecal samples (Fig. 3h). The maximal weight loss was 8.79% at 7 dpc (Fig. 3i). A PRNT90 assay detected neutralizing antibody from the sera of one out of three fomite-contact hamsters at 16 dpc (titres at 1:320). Taken together, these results suggest that transmission of SARS-CoV-2 among hamsters was mediated mainly by aerosols, rather than by fomites.
a, Infectious viral load (log10(TCID50 per ml), shown as bars) and viral RNA copy numbers (log10(RNA copies per ml), shown as dots) detected in the nasal washes of donor hamsters (n = 3) inoculated with 8 × 104 TCID50 of SARS-CoV-2. Colour-matched bars and dots represent results from the same hamster. b, Infectious virus and viral RNA detected in the faecal samples of donor hamsters (n = 3). ND, not determined. c, Changes in body weight of donor hamsters (n = 3); individual data points and mean ± s.d. are shown. d, Aerosol transmission of SARS-CoV-2 to naive hamsters (n = 3) exposed to donors for 8 h at 1 dpi; infectious virus and viral RNA detected in the nasal washes of aerosol-contact hamsters are shown. e, Infectious virus and viral RNA detected in the faecal samples of aerosol-contact hamsters (n = 3). f, Changes in body weight (per cent weight change compared to the day of exposure) of aerosol-contact hamsters (n = 3). g, Fomite transmission of SARS-CoV-2 to naive hamsters (n = 3) that were single-housed in the soiled cage of a donor for 48 h; infectious virus and viral RNA detected in the nasal washes of fomite-contact hamsters are shown. h, Infectious virus and viral RNA detected in the faecal samples of fomite-contact hamsters (n = 3). i, Changes in body weight (per cent weight change compared to the day of exposure) of fomite-contact hamsters (n = 3). Aerosol transmission and fomite transmission experiments were each performed once, with three repeats.
Our results indicate that the golden hamster is a suitable experimental animal model for investigating infections with SARS-CoV-2, as there is apparent weight loss in the inoculated and naturally infected hamsters and evidence of efficient viral replication in the nasal mucosa and epithelial cells of the lower respiratory system. The ability of SARS-CoV-2 to infect olfactory sensory neurons at the nasal mucosa may explain the anosmia reported in patients with COVID-19. Hamsters support efficient transmission of SARS-CoV-2 from inoculated donor to naive contact hamsters by direct contact or via aerosols. We also show that transmission from the donor to naive hamsters may occur within a short period soon after inoculation; however, the donors have a short communicable period (of fewer than six days) despite of continuous detection of viral RNA in the nasal washes. Our findings are consistent with a recent report28 that was published while the current study was under peer review. Hamsters are easy to handle, and there are reagents to support immunological studies for vaccine development29,30,31. The results also highlighted similarity and differences between the SARS-CoV and SARS-CoV-2 in the hamster model. Both viruses replicated efficiently in respiratory epithelial cells with peak viral load detected soon after inoculation, followed by an infiltration of mononuclear inflammatory cells in the lungs and rapid clearance of infectious virus by 7 dpi. Understanding the host defence mechanism that leads to rapid viral clearance in the respiratory tissues in hamsters may aid the development of effective countermeasures for SARS-CoV-2. The efficient transmission of SARS-CoV-2 to naive hamsters by aerosols also provides an opportunity to understand the transmission dynamics for this coronavirus.
Methods
No statistical methods were used to predetermine sample size. The hamsters were randomized from different litters into experimental groups and investigators were not blinded to allocation during experiments and outcome assessment.
Virus
The BetaCoV/Hong Kong/VM20001061/2020 virus was isolated from the nasopharyngeal aspirate and throat swab of a confirmed patient with COVID-19 in Hong Kong (GISAID identifier EPI_ISL_412028), using Vero E6 cells at the BSL-3 core facility (LKS Faculty of Medicine, University of Hong Kong). Vero E6 cells were purchased from ATCC (CRL-1586) without further authentication, and the cells routinely tested negative for Mycoplasma sp. by real-time PCR. Stock virus (107.25 TCID50 per ml) was prepared after three serial passages in Vero E6 cells in Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 4.5 g/l d-glucose, 100 mg/l sodium pyruvate, 2% FBS, 100,000 U/l penicillin–streptomycin and 25mM HEPES. The sequence of the stock virus was identical to the original clinical isolate.
Hamster experiments
Male golden hamsters at 4–5 weeks old were obtained from the Laboratory Animal Services Centre (Chinese University of Hong Kong). The hamsters were originally imported from Harlan (Envigo) in 1998. All experiments were performed at the BSL-3 core facility, (LKS Faculty of Medicine). The hamsters were randomized from different litters into experimental groups, and were acclimatized at the BSL-3 facility for 4–6 d before the experiments. The study protocol was reviewed and approved by the Committee on the Use of Live Animals in Teaching and Research, The University of Hong Kong (CULATR no. 5323-20). Experiments were performed in compliance with all relevant ethical regulations. For challenge studies, hamsters were anaesthetized with ketamine (150 mg/kg) and xylazine (10 mg/kg) via intraperitoneal injection and were intranasally inoculated with 8 × 104 TCID50 of SARS-CoV-2 in 80 μl DMEM. On days 2, 5 and 7, three hamsters were euthanized by intraperitoneal injection of pentobarbital at 200 mg/kg. No blinding was done and a sample size of three hamsters was selected to assess the level of variation between animals. The left lungs and one kidney were collected for viral load determination and were homogenized in 1 ml PBS. Brain, nasal turbinate, right lungs, liver, heart, spleen, duodenum and kidney were fixed in 4% paraformaldehyde for histopathological examination. To collect faecal samples, hamsters were transferred to a new cage one day in advance and fresh faecal samples (10 pieces) were collected for quantitative real-time RT–PCR and TCID50 assay. To evaluate SARS-CoV-2 transmissibility by direct contact, donor hamsters were anaesthetized and inoculated with 8 × 104 TCID50 of SARS-CoV-2. At 1 or 6 dpi, one inoculated donor was transferred to cohouse with one naive hamster in a clean cage; the cohousing of the hamsters continued for at least 13 days. Experiments were repeated with three pairs of donors:direct contact at 1:1 ratio32,33. Body weight and clinical signs of the hamsters were monitored daily. To evaluate SARS-CoV-2 transmissibility via aerosols, one naive hamster was exposed to one inoculated donor hamster in two adjacent stainless steel wired cages at 1 dpi (of the donor) for 8 h (Extended Data Fig. 3). DietGel76A (ClearH2O) was provided to the hamsters during the 8-h exposure. Exposure was done by holding the hamsters inside individually ventilated cages (IsoCage N, Techniplast) with 70 air changes per h. Experiments were repeated with three pairs of donors:aerosol contact at 1:1 ratio. After exposure, the hamsters were single-housed in separate cages and were monitored daily for 14 d. To evaluate the transmission potential of SARS-CoV-2 virus via fomites, three naive fomite contact hamsters were each introduced to a soiled donor cage at 2 dpi. The fomite contact hamsters were single-housed for 48 h inside the soiled cages and then were each transferred to a new cage at 4 dpi (of the donor). All hamsters were monitored daily for 14 d. For nasal wash collection, hamsters were anaesthetized using ketamine (100 mg/kg) and xylazine (10 mg/kg) via intraperitoneal injection and 160 μl of PBS containing 0.3% BSA was used to collect nasal washes from both nostrils of each hamster. Collected nasal washes were diluted 1:1 by volume and aliquoted for TCID50 assay in Vero E6 cells and for quantitative real-time RT–PCR. The contact hamsters were handled first, followed by surface decontamination using 1% virkon and handling of the donor hamster.
Environmental sampling
To monitor the level of fomite contamination of SARS-CoV-2 virus in soiled cages, surface samples (5 cm × 5 cm; the whole water bottle nozzle was also swabbed) were collected using flocked polyester swabs (Puritan). Swabs were stored in 0.5 ml of viral transport medium (VTM, containing 0.45% bovine serum albumin, vancomycin, amikacin and nystatin) at −80 °C. In addition, 10 pieces of corn cob bedding were collected from the soiled cage and were soaked in 1 ml VTM for 30 min before titration of infectious virus and viral RNA extraction. Infectious viral loads were determined in Vero E6 cells, and viral RNA copy numbers were determined by quantitative real-time RT–PCR.
Viral load determination by quantitative real-time RT–PCR
RNA was extracted from 140-μl samples using QIAamp viral RNA mini kit (Qiagen) and eluted with 60 μl of water. Two μl RNA was used for quantitative real-time RT–PCR to detect and quantify the N gene of SARS-CoV-2 using TaqMan Fast Virus 1-Step Master Mix, as previously described34.
PRNT90 assay
The experiments were carried out in duplicate using Vero E6 cells seeded in 24-well culture plates. Serum samples were heat-inactivated at 56 °C for 30 min and were serially diluted and incubated with 30–40 plaque-forming units of SARS-CoV-2 for 1 h at 37 °C. The virus–serum mixtures were added to the cells and incubated 1 h at 37 °C in 5% CO2 incubator. The plates were overlaid with 1% agarose in cell culture medium and incubated for 3 d. Thereafter, the plates were fixed and stained with 1% crystal violet. Antibody titres were defined as the highest serum dilution that resulted in >90% reduction in the number of plaques (PRNT90).
Histopathology and immunohistochemistry
Tissue (hearts, livers, spleens, duodenums, brains, right lungs and kidneys) were fixed in 4% paraformaldehyde and were processed for paraffin embedding. The 4-μm sections were stained with haematoxylin and eosin for histopathological examinations. For immunohistochemistry, SARS-CoV-2 N protein was detected using monoclonal antibody (4D11)35; CD3 was detected using polyclonal rabbit anti-human CD3 antibodies (DAKO); and neuron-specific β-III tubulin was detected using monoclonal antibody clone TuJ1 (R&D Systems). Images were captured using a Leica DFC 5400 digital camera and were processed using Leica Application Suite v.4.13.
Statistics and reproducibility
Kruskal–Wallis test and Dunn’s multiple comparisons test were used to compare viral loads in the lungs and kidney at 2, 5 and 7 dpi. The area under the curve was calculated from the nasal washes of the donor and contact hamsters followed by Mann–Whiteny test. Data were analysed in Microsoft Excel for Mac, version 16.35 and GraphPad Prism version 8.4.1. For the detection of viral replication in hamsters, 9 hamsters were inoculated and tissues were collected from hamsters at 2 (n = 3), 5 (n = 3) and 7 (n = 3) dpi; the results from the three hamsters were similar (Fig. 1a, b). Inoculation of the donor hamsters was independently performed twice and the inoculated hamsters showed comparable weight loss and shed comparable amount of virus in the nasal washes (Figs. 2a, b, 3a, b). Transmissions by direct contact, aerosols or fomites were performed with three pairs of donor:contacts at 1:1 ratio.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The sequence of SARS-CoV-2 virus BetaCoV/Hong Kong/VM20001061/2020 can be accessed at www.gisaid.org (identifier: EPI_ISL_412028). All experimental data shown in Figs. 1–3, Extended Data Figs. 2, and Extended Data Table 1 are available from the corresponding author upon request.
References
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
WHO. Coronavirus Disease (COVID-2019) Situation Reports. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports (2020).
Li, Q. et al. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. N. Engl. J. Med. 382, 1199–1207 (2020).
He, X. et al. Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat. Med. 26, 672–675 (2020).
Wu, Z. & McGoogan, J. M. Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in China: summary of a report of 72 314 cases from the Chinese Center for Disease Control and Prevention. J. Am. Med. Assoc. 323, 1239–1242 (2020).
Wu, J. T. et al. Estimating clinical severity of COVID-19 from the transmission dynamics in Wuhan, China. Nat. Med. 26, 506–510 (2020).
McCray, P. B. Jr et al. Lethal infection of K18-hACE2 mice infected with severe acute respiratory syndrome coronavirus. J. Virol. 81, 813–821 (2007).
Menachery, V. D. et al. SARS-like WIV1-CoV poised for human emergence. Proc. Natl Acad. Sci. USA 113, 3048–3053 (2016).
Tseng, C. T. et al. Severe acute respiratory syndrome coronavirus infection of mice transgenic for the human angiotensin-converting enzyme 2 virus receptor. J. Virol. 81, 1162–1173 (2007).
Yang, X. H. et al. Mice transgenic for human angiotensin-converting enzyme 2 provide a model for SARS coronavirus infection. Comp. Med. 57, 450–459 (2007).
Baas, T. et al. Genomic analysis reveals age-dependent innate immune responses to severe acute respiratory syndrome coronavirus. J. Virol. 82, 9465–9476 (2008).
Glass, W. G., Subbarao, K., Murphy, B. & Murphy, P. M. Mechanisms of host defense following severe acute respiratory syndrome-coronavirus (SARS-CoV) pulmonary infection of mice. J. Immunol. 173, 4030–4039 (2004).
Hogan, R. J. et al. Resolution of primary severe acute respiratory syndrome-associated coronavirus infection requires Stat1. J. Virol. 78, 11416–11421 (2004).
Roberts, A. et al. Aged BALB/c mice as a model for increased severity of severe acute respiratory syndrome in elderly humans. J. Virol. 79, 5833–5838 (2005).
Hamming, I. et al. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J. Pathol. 203, 631–637 (2004).
Bao, L. et al. The pathogenicity of SARS-CoV-2 in hACE2 transgenic mice. Nature https://doi.org/10.1038/s41586-020-2312-y (2020).
Rockx, B. et al. Comparative pathogenesis of COVID-19, MERS, and SARS in a nonhuman primate model. Science 368, 1012–1015 (2020).
Munster, V. J. et al. Respiratory disease and virus shedding in rhesus macaques inoculated with SARS-CoV-2. Nature https://doi.org/10.1038/s41586-020-2324-7 (2020).
Roberts, A. et al. Animal models and vaccines for SARS-CoV infection. Virus Res. 133, 20–32 (2008).
Roberts, A. et al. Severe acute respiratory syndrome coronavirus infection of golden Syrian hamsters. J. Virol. 79, 503–511 (2005).
de Wit, E. et al. The Middle East respiratory syndrome coronavirus (MERS-CoV) does not replicate in Syrian hamsters. PLoS ONE 8, e69127 (2013).
MacPhail, M. et al. Identification of small-animal and primate models for evaluation of vaccine candidates for human metapneumovirus (hMPV) and implications for hMPV vaccine design. J. Gen. Virol. 85, 1655–1663 (2004).
Buthala, D. A. & Soret, M. G. Parainfluenza type 3 virus infection in hamsters: virologic, serologic, and pathologic studies. J. Infect. Dis. 114, 226–234 (1964).
Ali, M. J., Teh, C. Z., Jennings, R. & Potter, C. W. Transmissibility of influenza viruses in hamsters. Arch. Virol. 72, 187–197 (1982).
Iwatsuki-Horimoto, K. et al. Syrian hamster as an animal model for the study of human influenza virus infection. J. Virol. 92, e01693-17 (2018).
Leung, W. K. et al. Enteric involvement of severe acute respiratory syndrome-associated coronavirus infection. Gastroenterology 125, 1011–1017 (2003).
Jones, R. M. & Brosseau, L. M. Aerosol transmission of infectious disease. J. Occup. Environ. Med. 57, 501–508 (2015).
Chan, J. F. et al. Simulation of the clinical and pathological manifestations of coronavirus disease 2019 (COVID-19) in golden Syrian hamster model: implications for disease pathogenesis and transmissibility. Clin. Infect. Dis. ciaa325 (2020).
Miao, J., Chard, L. S., Wang, Z. & Wang, Y. Syrian hamster as an animal model for the study on infectious diseases. Front. Immunol. 10, 2329 (2019).
Warner, B. M., Safronetz, D. & Kobinger, G. P. in Emerging and Re-emerging Viral Infections (Advances in Experimental Medicine and Biology) Vol. 972 (eds Rezza, G. & Ippolito, G.) 87–101 (Springer, 2016).
Zivcec, M., Safronetz, D., Haddock, E., Feldmann, H. & Ebihara, H. Validation of assays to monitor immune responses in the Syrian golden hamster (Mesocricetus auratus). J. Immunol. Methods 368, 24–35 (2011).
Belser, J. A., Maines, T. R., Katz, J. M. & Tumpey, T. M. Considerations regarding appropriate sample size for conducting ferret transmission experiments. Future Microbiol. 8, 961–965 (2013).
Nishiura, H., Yen, H. L. & Cowling, B. J. Sample size considerations for one-to-one animal transmission studies of the influenza A viruses. PLoS ONE 8, e55358 (2013).
Chu, D. K. W. et al. Molecular diagnosis of a novel coronavirus (2019-nCoV) causing an outbreak of pneumonia. Clin. Chem. 66, 549–555 (2020).
Nicholls, J. M. et al. Time course and cellular localization of SARS-CoV nucleoprotein and RNA in lungs from fatal cases of SARS. PLoS Med. 3, e27 (2006).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Li, W. et al. Receptor and viral determinants of SARS-coronavirus adaptation to human ACE2. EMBO J. 24, 1634–1643 (2005).
Acknowledgements
We thank N. Ip for comments on the interpretation of nasal mucosal data; D. Rowlands, C. Y. H. Leung and J. Rudds for comments on hamster handling; and the Laboratory Animal Unit (University of Hong Kong) and Laboratory Animal Services Centre (Chinese University of Hong Kong) for animal husbandry support. L.L.M.P. was supported by Croucher Foundation. This study was supported by contract HHSN272201400006C from the National Institute of Allergy and Infectious Diseases of the National Institutes of Health (USA) and the theme-based research scheme (T11-705/14N) from the Research Grants Council (Hong Kong SAR, China).
Author information
Authors and Affiliations
Contributions
S.F.S., L.-M.Y., A.W.H.C. and H.-L.Y. designed and performed the experiments; K.-T.C., A.Y.L.W., P.K. and R.A.P.M.P. performed the experiments, K.F. and J.M.N. performed immunohistochemistry and histopathological examination; L.L.M.P., J.M.N., M.P. and H.-L.Y. analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Emmie de Wit, Stanley Perlman and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sequence alignment of ACE2 proteins (1–420) from human, macaque, hamster and mouse.
Asterisks denote amino acid residues of human ACE2 that have been shown by structural analysis to interact with the receptor-binding domain of SARS-CoV-236. Amino acid residues that are important for the interaction between human ACE2 and receptor-binding domain of SARS-CoV are highlighted by red boxes37.
Extended Data Fig. 2 Haematoxylin and eosin staining and immunohistochemistry on hamster tissues challenged with SARS-CoV-2.
a, Hyperplasia of the pneumocytes detected at 7 dpi. b, Detection of CD3-positive cells (using rabbit anti-human CD3 polyclonal antibody) in the lungs at 5 dpi. c, Detection of SARS-CoV-2 N protein (red staining, using monoclonal antibody 4D11) and olfactory neurons (brown staining, using monoclonal antibody TuJ1) from the nasal turbinate at 5 dpi. d, Detection of olfactory neurons (using monoclonal antibody TuJ1) from the nasal turbinate of a mock-infected hamster (n = 1). e, Nasal epithelial cells from the nasal turbinate of a mock-infected hamster (n = 1) showed negative staining for TuJ1. f, Detection of olfactory neurons from nasal turbinate at 2 dpi. g, Detection of olfactory neurons from nasal turbinate at 7 dpi. h, Detection of olfactory neurons from nasal turbinate at 14 dpi. i, Haematoxylin and eosin (H&E) staining of the brain tissue at 5 dpi. j, H&E staining of the heart at 5 dpi. k, H&E staining of the liver at 5 dpi. l, H&E staining of the kidney at 5 dpi. Hamsters were intranasally inoculated with PBS (mock infection, n = 1) or with 8 × 104 TCID50 of SARS-CoV-2 (n = 9), and the tissues were collected at 2 (n = 3), 5 (n = 3) and 7 (n = 3) dpi. H&E and immunohistochemistry performed on tissues from three hamsters at day 2, 5 and 7 dpi showed similar results; representative results are shown.
Extended Data Fig. 3 Experimental layout for the aerosol transmission experiment in hamsters.
To evaluate SARS-CoV-2 transmissibility via aerosols, one naive hamster was exposed to one inoculated donor hamster in two adjacent stainless steel wired cages at 1 dpi for 8 h. DietGel76A (ClearH2O) was provided to the hamsters during the 8-h exposure. Exposure was done by holding the hamsters inside individually ventilated cages (IsoCage N, Techniplast) with 70 air changes per h. Experiments were repeated with three pairs of donors:aerosol contact at 1:1 ratio. After exposure, the hamsters were single-housed in separate cages and were monitored daily for 14 d.
Supplementary information
Rights and permissions
About this article
Cite this article
Sia, S.F., Yan, LM., Chin, A.W.H. et al. Pathogenesis and transmission of SARS-CoV-2 in golden hamsters. Nature 583, 834–838 (2020). https://doi.org/10.1038/s41586-020-2342-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2342-5
This article is cited by
-
Seroevidence of SARS-CoV-2 spillback to rodents in Sarawak, Malaysian Borneo
BMC Veterinary Research (2024)
-
Modulation of alveolar macrophage and mitochondrial fitness by medicinal plant-derived nanovesicles to mitigate acute lung injury and viral pneumonia
Journal of Nanobiotechnology (2024)
-
Transmission dynamics of MERS-CoV in a transgenic human DPP4 mouse model
npj Viruses (2024)
-
SARS-CoV-2 immunity in animal models
Cellular & Molecular Immunology (2024)
-
Inhalation of ACE2-expressing lung exosomes provides prophylactic protection against SARS-CoV-2
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.