Abstract
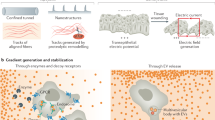
Eukaryotic cells migrate by coupling the intracellular force of the actin cytoskeleton to the environment. While force coupling is usually mediated by transmembrane adhesion receptors, especially those of the integrin family, amoeboid cells such as leukocytes can migrate extremely fast despite very low adhesive forces1. Here we show that leukocytes cannot only migrate under low adhesion but can also transmit forces in the complete absence of transmembrane force coupling. When confined within three-dimensional environments, they use the topographical features of the substrate to propel themselves. Here the retrograde flow of the actin cytoskeleton follows the texture of the substrate, creating retrograde shear forces that are sufficient to drive the cell body forwards. Notably, adhesion-dependent and adhesion-independent migration are not mutually exclusive, but rather are variants of the same principle of coupling retrograde actin flow to the environment and thus can potentially operate interchangeably and simultaneously. As adhesion-free migration is independent of the chemical composition of the environment, it renders cells completely autonomous in their locomotive behaviour.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
References
Lämmermann, T. & Sixt, M. Mechanical modes of ‘amoeboid’ cell migration. Curr. Opin. Cell Biol. 21, 636–644 (2009).
Abercrombie, M., Heaysman, J. E. & Pegrum, S. M. The locomotion of fibroblasts in culture. 3. Movements of particles on the dorsal surface of the leading lamella. Exp. Cell Res. 62, 389–398 (1970).
Liu, Y.-J. J. et al. Confinement and low adhesion induce fast amoeboid migration of slow mesenchymal cells. Cell 160, 659–672 (2015).
Friedl, P. & Wolf, K. Plasticity of cell migration: a multiscale tuning model. J. Cell Biol. 188, 11–19 (2010).
Lämmermann, T. et al. Rapid leukocyte migration by integrin-independent flowing and squeezing. Nature 453, 51–55 (2008).
Bergert, M. et al. Force transmission during adhesion-independent migration. Nat. Cell Biol. 17, 524–529 (2015).
Schmidt, S. & Friedl, P. Interstitial cell migration: integrin-dependent and alternative adhesion mechanisms. Cell Tissue Res. 339, 83–92 (2010).
Hons, M. et al. Chemokines and integrins independently tune actin flow and substrate friction during intranodal migration of T cells. Nat. Immunol. 19, 606–616 (2018).
Woolf, E. et al. Lymph node chemokines promote sustained T lymphocyte motility without triggering stable integrin adhesiveness in the absence of shear forces. Nat. Immunol. 8, 1076–1085 (2007).
Calderwood, D. A., Campbell, I. D. & Critchley, D. R. Talins and kindlins: partners in integrin-mediated adhesion. Nat. Rev. Mol. Cell Biol. 14, 503–517 (2013).
Le Berre, M., Aubertin, J. & Piel, M. Fine control of nuclear confinement identifies a threshold deformation leading to lamina rupture and induction of specific genes. Integr. Biol. (Camb.) 4, 1406–1414 (2012).
Renkawitz, J. et al. Adaptive force transmission in amoeboid cell migration. Nat. Cell Biol. 11, 1438–1443 (2009).
Renkawitz, J., Reversat, A., Leithner, A., Merrin, J. & Sixt, M. Micro-engineered “pillar forests” to study cell migration in complex but controlled 3D environments. Methods Cell Biol. 147, 79–91 (2018).
Vargas, P., Barbier, L., Sáez, P. J. & Piel, M. Mechanisms for fast cell migration in complex environments. Curr. Opin. Cell Biol. 48, 72–78 (2017).
Medeiros, N. A., Burnette, D. T. & Forscher, P. Myosin II functions in actin-bundle turnover in neuronal growth cones. Nat. Cell Biol. 8, 215–226 (2006).
Henson, J. H. et al. Two components of actin-based retrograde flow in sea urchin coelomocytes. Mol. Biol. Cell 10, 4075–4090 (1999).
Cramer, L. P. Molecular mechanism of actin-dependent retrograde flow in lamellipodia of motile cells. Front. Biosci. 2, d260–d270 (1997).
Driscoll, M. K. et al. Cell shape dynamics: from waves to migration. PLoS Comput. Biol. 8, e1002392 (2012).
Howe, J. D., Barry, N. P. & Bretscher, M. S. How do amoebae swim and crawl? PLoS One 8, e74382 (2013).
Bae, A. J. & Bodenschatz, E. On the swimming of Dictyostelium amoebae. Proc. Natl Acad. Sci. USA 107, E165–E166 (2010).
Mandeville, J. T. H., Lawson, M. A. & Maxfield, F. R. Dynamic imaging of neutrophil migration in three dimensions: mechanical interactions between cells and matrix. J. Leukoc. Biol. 61, 188–200 (1997).
Tozluoğlu, M. et al. Matrix geometry determines optimal cancer cell migration strategy and modulates response to interventions. Nat. Cell Biol. 15, 751–762 (2013).
Jankowiak, G., Peurichard, D., Reversat, A., Schmeiser, C. & Sixt, M. Modelling adhesion-independent cell migration. Math. Model. Methods Appl. Sci. 30, 513–537 (2020).
Riedl, J. et al. Lifeact: a versatile marker to visualize F-actin. Nat. Methods 5, 605–607 (2008).
Leithner, A. et al. Diversified actin protrusions promote environmental exploration but are dispensable for locomotion of leukocytes. Nat. Cell Biol. 18, 1253–1259 (2016).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Leithner, A., Merrin, J., Reversat, A. & Sixt, M. Geometrically complex microfluidic devices for the study of cell migration. Protoc. Exch. https://doi.org/10.1038/protex.2016.063 (2016).
Schwarz, J. et al. A microfluidic device for measuring cell migration towards substrate-bound and soluble chemokine gradients. Sci. Rep. 6, 36440 (2016).
Sixt, M. & Lämmermann, T. in Cell Migration: Developmental Methods and Protocols (eds. Wells, C. M. & Parsons, M.) 149–165 (Humana, 2011).
Képiró, M. et al. para-Nitroblebbistatin, the non-cytotoxic and photostable myosin II inhibitor. Angew. Chem. Int. Ed. Engl. 53, 8211–8215 (2014).
Tinevez, J. Y. et al. TrackMate: an open and extensible platform for single-particle tracking. Methods 115, 80–90 (2017).
Sommer, C., Straehle, C., Koethe, U. & Hamprecht, F. A. Ilastik: interactive learning and segmentation toolkit. In 2011 IEEE International Symposium on Biomedical Imaging: From Nano to Macro 230–233 (IEEE, 2011).
Acknowledgements
We thank A. Leithner and J. Renkawitz for discussion and critical reading of the manuscript; J. Schwarz and M. Mehling for establishing the microfluidic setups; the Bioimaging Facility of IST Austria for excellent support, as well as the Life Science Facility and the Miba Machine Shop of IST Austria; and F. N. Arslan, L. E. Burnett and L. Li for their work during their rotation in the IST PhD programme. This work was supported by the European Research Council (ERC StG 281556 and CoG 724373) to M.S. and grants from the Austrian Science Fund (FWF P29911) and the WWTF to M.S. M.H. was supported by the European Regional Development Fund Project (CZ.02.1.01/0.0/0.0/15_003/0000476). F.G. received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement no. 747687.
Author information
Authors and Affiliations
Contributions
A.R. and M.S. conceived the experiments and wrote the manuscript, with critical feedback from all authors. A.R. designed, performed and analysed the experiments, with the help of I.d.V., J.S., M.H., R.H. and S.T. A.R. and J.M. designed the microfluidic devices. J.M. performed the photolithography. F.G. designed and performed the nanoridge experiments. J.A. fabricated the nanoridge substrates. R.H. wrote the image analysis scripts. A.C.-J., R.V., J.M., R.H., M.S. and A.R. discussed the physical model. A.C.-J. and R.V. wrote the physical model.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Kenneth Yamada and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Effects of Tln1 deletion on T cell adhesion and migration.
a, Workflow scheme of virus production, cell infection and analysis. d, day. b, RNA-seq analysis of talin 1 and 2 expression in mouse CD4+ primary T cells, the T cell line and bone marrow-derived dendritic cells (BMDCs). c, Western blot analysis of talin 1 and talin 2 expression in T cells infected with lentivirus containing scrambled and talin 1-targeting CRISPR guides (guides 1–3 (Talin1g1–Talin1g3)). Data are representative of three independent experiments. d, Related to Fig. 1a. Phase-contrast images of control and talin-KO cells. Scale bars, 50 μm. Representative of four experiments. e, Related to Fig. 1b. Representative snapshots of epifluorescence (EPI) and TIRF microscopy of control cells (left) and talin-KO cells (middle) expressing mCherry and eGFP Lifeact fusions, respectively, plated on a Fc-ICAM1-coated surface. Representative of three experiments. The right panel shows bright field and merged fluorescence. Scale bar, 5 μm. f, Related to Fig. 1c. Representative snapshots of control and talin-KO cells embedded in 3D collagen gel at t = 120 min; individual tracks are displayed in different colours. Representative of three experiments. Scale bars, 100 μm. g, Related to Fig. 1d. Left, scheme of 5-μm high confinement used in f, g and j. Right, control cells (left) and talin-KO T cells (right) expressing Lifeact-GFP (red), stained with Hoechst (cyan), were placed under 5-μm height confinement and imaged by video microscopy. Representative of four experiments. Snapshot is at t = 60 min. Nucleus tracks are displayed in yellow. Scale bars, 50 μm. h, Cell speed of control T cells (n = 62) and T cells expressing Lifeact-GFP (n = 100) or Lifeact-mCherry (n = 138). Data are mean ± s.d.; ns P = 0.0775, Kruskal–Wallis one-way ANOVA followed by two-sided post hoc Dunn’s test; control versus Lifeact-GFP or Lifeact-mCherry: P > 0.999; Lifeact-GFP versus Lifeact-mCherry: P = 0.0717. i, Speed of T cells placed under 5-μm (n = 300) or 8-μm (n = 274) confinement. Data are mean ± s.d.; ns P = 0.8460, two-sided Mann–Whitney U-test. j, Perimeter (left) and area (right) of control and talin-KO cells placed under 3-μm confinement (control cells: n = 145, talin-KO cells: n = 127). Representative of three experiments. Data are mean ± s.d.; ns P = 0.4261 for perimeter and P = 0.2558 for area, two-sided Mann–Whitney U-test. k, Spreading area observed by TIRF microscopy of control (n = 27) and talin-KO (n = 20) cells under 5-μm confinement. Representative of two experiments. Data are mean ± s.d.; ns P = 0.2025, two-sided Mann–Whitney U-test. l, Scheme of the 5-μm high Fc-ICAM1-coated confinement (left) used to measure velocities and F-actin retrograde flow (right) in control cells (n = 49) and talin-KO T cells (n = 59). Representative of five experiments. ****P < 0.0001, two-sided Mann–Whitney U-test.
Extended Data Fig. 2 Topography rescues T cell locomotion in the absence of adhesion.
a–c, Migration of control (n = 71) and talin-KO (n = 74) cells under 3-μm confinement. Representative of four independent experiments. In a, the left panel shows the scheme of the 3-μm confinement used in a–c. The right panel shows representative snapshots of control and talin-KO cells under 3-μm confinement at t = 60 min. Individual tracks are displayed in different colours. Scale bars, 50 μm. In b, the speed and displacement of control and talin-KO cells under 3-μm confinement are shown. Data are mean ± s.d.; ****P < 0.0001, two-sided Mann–Whitney U-test. In c, the MSD of control cells (black) and talin-KO cells (red) is shown. Data are mean ± s.e.m.; P < 0.0001, two-sided Mann–Whitney U-test. d, Related to Fig. 2a–d. Scheme of multilayered microfluidic devices used to push cells into the pillar maze or into microchannels used in e and f. e, Snapshots of video microscopy at t = 60 min of control and talin-KO cells in the pillar device. The nucleus is shown in cyan (Hoechst), Lifeact-GFP reporter in red and the bright-field image in grey. Individual cell tracks are displayed in yellow. Scale bars, 50 μm. Time is in min:s. Representative of three experiments. f, Related to Fig. 2g, h. Velocities (μm/min) of control and talin-KO cells in serrated channels of different diameters (with a conserved serration period of 6 μm). Representative of three experiments. n = 57 for control cells and n = 9 for talin-KO cells. *P = 0.0356; otherwise not significant, Kruskal–Wallis one-way ANOVA followed by post hoc Dunn’s test. Boxes extend from the 25th to the 75th percentiles, with the middle line showing the median and the whiskers representing the minimum to maximum values.
Extended Data Fig. 3 Topography rescues dendritic cell locomotion in the absence of adhesion.
a, Top left, scheme of a mature dendritic cell migrating in a channel designed for the EDTA experiment. Mature dendritic cells migrate in a device containing channels with different serration periods, with or without 30 mM EDTA. Top right, single-cell migration speed with (+, blue) and without (-, grey) EDTA treatment. Bottom, histograms of cell speed in different serrations with (blue) or without (grey) EDTA. Representative of three independent experiments. n = 35 cells without EDTA, n = 17 cells with EDTA. Data are mean ± s.d.; **P = 0.0022, ****P < 0.0001 and ns P = 0.6363, one-way ANOVA with Kruskal–Wallis test followed by post hoc Dunn’s test. b, Top left, scheme of a mature dendritic cell migrating in a channel designed for the EDTA experiment, along a CCL19 gradient (indicated in red). Mature dendritic cells migrate towards the chemokine CCL19 into a device that contains microchannels with different serration periods, with or without 30 mM EDTA. Top right, migration speed with (+, blue) and without (-, grey) EDTA treatment. Bottom, histograms of cell speed in the different serrated channels with (blue) or without (grey) EDTA. Representative of three independent experiments. Without EDTA: n = 44 cells; with EDTA: n = 52 (6–12 μm), n = 44 (24 μm), n = 33 (smooth channels). Data are mean ± s.d.; ***P = 0.0001 and ****P < 0.0001, one-way ANOVA with Kruskal–Wallis test followed by post hoc Dunn’s test.
Extended Data Fig. 4 Effect of topographical features on T cell migration.
a, Related to Fig. 4d. Scheme of channel designs with decreasing complexity. In brief, serrations with a period of 6, 12 and 24 μm were arranged in consecutive zones of a 5-μm wide channel ending with a smooth zone. The left panel shows the different geometries. The right and middle panels show the velocity of cells migrating in zones with a different period. The total number of cells in each design is shown. Representative of four (control cells) and eight (talin-KO cells) independent experiments. Control: *P = 0.0415. KO arrow ‘right’: ***P = 0.0008 and **P = 0.0013; KO arrow ‘left’: ***P = 0.0003 and **P = 0.0018; KO bulb: *P = 0.0344; KO half-bulb: **P = 0.0016 (6 μm versus smooth) and **P = 0.0013 (12 μm versus smooth), otherwise not significant, one-way ANOVA with Kruskal–Wallis test followed by post hoc Dunn’s test. Boxes extend from the 25th to the 75th percentiles, with the middle line showing the median and the whiskers representing the minimum to maximum values. b, Top, scheme of cell and actin retrograde flow length. Bottom, control cells expressing the Lifeact-GFP reporter with 10 mM EDTA (grey) or talin-KO cells (red), under 5-μm confinement (left, n = 12) or in a 5 × 5-μm channel (right, n = 9 for control cells and n = 8 for talin-KO cells), were observed by TIRF microscopy, and the mean cell length (ns P = 0.0555, two-sided Mann–Whitney U-test) and the actin retrograde flow length (ns P = 0.7247, two-sided Mann–Whitney U-test) were measured using kymograph analysis. Representative of two independent experiments. Data are mean ± s.d.
Extended Data Fig. 5 Actin polymerization versus contractility during topography-based locomotion.
a, Scheme of the device used for the experiments in Fig. 2i, j and Supplementary Figs. 3, 5. b, Control T cells migrating under confinement (2.5D) or in smooth versus serrated channels as shown in the top panels. After 60 min, 50 μM para-nitroblebbistatin (pNB) was added. The migration speed before (-) and after (+) treatment is shown. Representative of four independent experiments. n = 68 for 2.5D, n = 79 for serrated channel and n = 44 for smooth channel. ****P < 0.0001, two-tailed, Wilcoxon matched-pairs signed-rank test. c, Migration speed change after versus before pNB treatment. Data are mean ± s.d. ns P > 0.9999 (2.5D versus serrated channel), ns P = 0.1414 (2.5D versus smooth channel) and ns P = 0.1044 (serrated vs smooth channel), one-way ANOVA with Kruskal–Wallis test followed by post hoc Dunn’s test. d, Control T cells migrating under confinement or in smooth versus serrated channels as shown in the top panels. After 60 min, 50 μM pNB and 10 mM EDTA were added. The migration speed before (-) and after (+) treatment is shown. Representative of four independent experiments. n = 166 for 2.5D, n = 45 for serrated channel and n = 22 for smooth channel. ****P < 0.0001, two-tailed, Wilcoxon matched-pairs signed-rank test. e, Migration speed change after versus before pNB + EDTA treatment. Data are mean ± s.d. ns P = 0.1170 and ****P < 0.0001, one-way ANOVA with Kruskal–Wallis test followed by post hoc Dunn’s test. f, Histogram of migration speed variation after versus before pNB treatment alone (left, see b, c), EDTA treatment alone (middle, related to Fig. 2i, j) or pNB + EDTA treatment (right, see d, e). g, Related to Fig. 4e–g. Left, snapshots of a control T cell expressing Lifeact-mCherry incubated with 10 mM EDTA in different channel zones. The left panel from top to bottom shows bright field, Lifeact-mCherry in black, colour-coded snapshots of F-actin in the different channel zones (15-s intervals), and the corresponding kymograph of a cell migrating in the channel. Scale bars, 5 μm. Right, cell and actin retrograde flow velocities observed by wide-field microscopy and measured with kymograph analysis. n = 8 cells; 3 cells in all zones, 4 cells in the 6–12–24-μm zone and 1 cell in the 12–24-µm zone. *P = 0.0138 and ***P = 0.0002, one-way ANOVA with Kruskal–Wallis test followed by post hoc Dunn’s test. Boxes extend from the 25th to the 75th percentiles, with the middle line showing the median and the whiskers representing the minimum to maximum values.
Extended Data Fig. 6 Topography-based migration of primary T cells.
a, Scheme (top and side view) of control and Lifeact-GFP cells confined under agarose on a smooth surface coated with Fc-ICAM1 or passivated with PLL-PEG. WT, wild type. b, Scheme (top and side view) of control and Lifeact-GFP cells confined under agarose on a coverslip with linear ridges passivated with PLL-PEG. c, Minimum-intensity projection showing the trace of control cells migrating for 15 min under agarose on Fc-ICAM1-coated smooth surfaces (left), PLL-PEG-coated smooth surfaces (middle) and PLL-PEG-coated ridge-baring surfaces (right). Representative of six independent experiments. Scale bars, 50 μm. d, Randomly selected tracks of mixed control and Lifeact-GFP cells migrating under agarose on Fc-ICAM1-coated smooth surfaces (left, n = 183), PLL-PEG-coated smooth surfaces (middle, n = 183) and PLL-PEG-coated ridge-baring surfaces (right, n = 183). The orientation of ridges is indicated in the top right corner. e, Rose diagram (with circular lines from the centre showing 6%, 12% and 18% of total cells) of cells migrating on Fc-ICAM1-coated smooth surfaces (left, n = 419), PLL-PEG-coated smooth surfaces (middle, n = 650) and PLL-PEG-coated ridge-baring surfaces (right, n = 687). The orientation of ridges is indicated in the top right corner. n = 6 independent experiments. f, g, Velocities of cells migrating under agarose on Fc-ICAM1-coated smooth surfaces (n = 419), PLL-PEG-coated smooth surfaces (n = 650) and PLL-PEG-coated ridge-baring surfaces (n = 687). n = 6 independent experiments. Cell velocities (cell displacement divided by track duration) (f) and mean instantaneous velocities (g) are shown. ****P < 0.0001 and ns P = 0.0757, both one-way ANOVA with Kruskal–Wallis test followed by post hoc Dunn’s test.
Extended Data Fig. 7 Actin-based force transmission in primary T cells.
a, Scheme (top and side view) of cells migrating under agarose on a non-adhesive, topography-baring substrate. b, Tracks of migrating cells analysed in c–f. n = 18; mean cell speed = 6.58 μm/min; mean actin retrograde flow speed = 6.53 μm/min. Starting points of tracks are shifted to the origin. c, Mean speed of cells in relation to their mean actin retrograde flow speed (in relation to the substrate). Pearson's rank correlation coefficient r = −0.6484. A drop in retrograde flow with increasing locomotion speed demonstrates that, as in the transmembrane clutch paradigm of force transmission, actin slippage is inversely related to locomotion. d, Snapshots of actin flow analysis on ridged surfaces of cells expressing Lifeact-GFP. Left, cell core segmented with Ilastik (pink); the yellow arrow indicates cell direction, the blue arrow indicates the mean optical flow direction and the green arrow indicates actin retrograde flow. Middle, optical flow analysis (obtained with a customized MATLAB code). Right, actin single-spot tracking (TrackMate). Representative of three independent experiments. e, Angular distribution of actin retrograde flow and forwards locomotion of cells migrating on nanoridges. Only frames where optical flow analysis was confirming automatic particle tracking (deviation of <30°; see d) were subjected to analysis. The histogram on the left shows predominant retrograde actin flow when aligned along the ridges (90°). Retrograde actin flow declines towards 0° (perpendicular to the ridges), while forwards locomotion (right histogram) steadily increases, indicating that retrograde actin flow couples to topographical barriers and drives locomotion. n = 422 events of actin flow and cell movements obtained in n = 18 cells from three independent experiments. f, Frame-to-frame speed of cells migrating on nanoridges increases when forwards locomotion aligns perpendicular to topographical barriers (90° → 0° ≃ 15% increase). The orientation-dependent increase of cell speed is paralleled by a steady decrease of retrograde actin flow (90° → 0° ≃ 55%), indicating that retrograde actin flow couples to topographical barriers and drives locomotion. Frame-to-frame cell speed and actin retrograde flow speed were recorded in 18 cells from 3 independent experiments, and pooled to n = 412 (cell speed) and n = 422 (actin retrograde flow speed) events. The centre shows the mean, and both s.e.m. (solid lines) and s.d. (thick transparent lines) are shown. Before pooling, cell speed was normalized to the mean speed of the tracked cell, and actin retrograde flow speed was normalized to the mean actin retrograde flow speed.
Extended Data Fig. 8 Retrograde actin flow and shape changes in adhesion-free confinement.
Related to Fig. 4h. a–c, Primary T cells expressing Lifeact-GFP confined on PLL-PEG. n = 51 cells from three independent experiments: n = 22 cells for 100 nM CCL19, n = 16 cells for 10 nM CCL19 and n = 13 cells for 1 nM CCL19. a, Representative snapshots (right, t = 20, 25 and 30 s) and kymographs (left), of primary T cells expressing Lifeact-GFP confined on PLL-PEG, showing actin retrograde flow (red arrows). Scale bar, 5 μm. b, Representative snapshots (right, t = 20, 25 and 30 s) of the cell outline and the colour-coded curvature. Kymograph analysis (left) of the migrating T cell shows retrograde movement of cell body deformations (right, red arrowhead). c, Top, scheme of the travelling cell body deformations. Bottom, retrograde actin flow versus curvature flow velocities in primary T cells confined on an inert substrate and exposed to the indicated chemokine (CCL19) concentrations. n = 51 cells from three independent experiments: n = 22 cells for 100 nM CCL19, n = 16 cells for 10 nM CCL19 and n = 13 cells for 1 nM CCL19. Pearson's rank correlation coefficient r = 0.5542. d, e, Cell curvature versus cortical actin retrograde flow analysis in channels (obtained from three cells in three experiments). A scheme of the segmentation used to measure channel-dependent cell curvature and cortical actin retrograde flow velocity used in e is shown (d). Fluorescent images were binarized and the local curvature was calculated from a spline fit to the cell outline. Measurement of channel-dependent cell curvature versus cortical actin retrograde flow shows no correlation (Pearson's correlation coefficient c ≈ −0.006425) (e). The green line shows mean ± s.d.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary Text and a Supplementary Table 1.
Video 1: Control and talin KO T cells migrating in 3D collagen
Related to Fig. 1c. Control (left panel) and talin KO cells (right panel) migrating in 3D collagen. Individual cell tracks are marked in colors. Time-lapse over 90 min. Time stamp in h:min. Scale bar 100 µm. Representative of 3 independent experiments.
Video 2: Control and talin KO T cells migrating in 2.5D confinement
First part, related to Fig. 1d. Control (left panel) and talin KO cells (right panel) under 5 µm confinement. Lifeact-GFP reporter displayed in red, cell nucleus in cyan (Hoechst 33342). Individual cell tracks are marked in yellow. Second part related to Extended Data Fig. 2a-c: Control (left panel) and talin KO cells (right panel) migrating under 3 µm confinement. Individual cell tracks are highlighted with colors. Time-lapses of 60 min movie. Time stamps in min:sec. Scale bar 50 µm. Representative of 4 independent experiments.
Video 3: Actin dynamics in control and talin KO T cells in 2.5D confinement
Related to Fig. 1e-g. Examples of total internal reflection (TIRF) microscopy of control and talin KO cells expressing Lifeact-GFP under 5 µm confinement. First two examples show brightfield (left panel), Lifeact-GFP TIRF in black (middle panel) and merged images (brightfield in gray, TIRF in green, right panel) acquired every 3 sec. Example 3 shows cells from Fig. 1e-g, where TIRF of Lifeact-GFP (black) was acquired every sec. Time stamps in min:sec. Scale bars 5 µm. Representative of 3 independent experiments.
Video 4: Migration of T cells in pillar arrays
Related to Fig. 2a-d. Control and talin KO cells (upper and lower panel, respectively) migrating under 5 µm confined pillar arrays with separation of 3, 4, 5 and 6 µm. Lifeact-GFP reporter in red, cell nucleus in cyan (Hoechst 33342). Individual cell tracks are marked in yellow. Time-lapse of 60 min. Time stamp in min:sec. Scale bar 50 µm. Representative of at least 3 independent experiments.
Video 5: Control and talin KO T cells migrating in smooth and serrated channel
First part, related to Fig. 2e-f. Control and talin KO cells (upper and lower panel, respectively) in 5x5 µm smooth microchannel. Individual cell tracks are marked in yellow. Second part, related to Fig. 2g-h. talin KO and control cells in 5x5 µm serrated microchannels. Lifeact-GFP reporter displayed in red nucleus in cyan (Hoechst 33342). Time-lapse of 60 sec. Time stamp in min:sec. Scale bar 20 µm. All representative of at least 3 independent experiments.
Video 6: Adhesion-free T cell migration in textured channels
Related to Fig. 2i-j. Control cells migrating in various microchannels, before and after 10 mM EDTA (added after 90 min). Lifeact-GFP reporter displayed in red, cell nucleus is marked in cyan (Hoechst 33342). Time stamp in min:sec. Scale bar 20 µm. Representative of 3 independent experiments.
Video 7: Adhesion-free neutrophil and dendritic cell migration in varying geometries
First part: PLB-985 (neutrophil cell line) in 5 µm-high serrated microchannels with 6 µm, 12 µm, 24 µm and no serrations. Lifeact-GFP reporter displayed in red (middle panel). 10 mM EDTA was added prior to recording. Time interval of 10 sec. Scale bar 20 µm. Second part, related to Extended Data Fig. 3: Mature bone marrow-derived dendritic cells migrating in channels toward CCL19 added on left side. Left panel without and right panel with 30 mM EDTA. Time interval 30 sec. Scale bar 20 µm. Time stamps in min:sec. Representative of 3 independent experiments.
Video 8: Force transmission of T cells in varying geometries
Related to Fig. 4a-d. Control and talin KO cells in 5 µm-high serrated channels with 6 µm, 12 µm, 24 µm and no serration. Lifeact-GFP reporter displayed in red, cell nucleus in cyan (Hoechst 33342). Individual cell tracks are marked in a speed-dependent color code (red: fast; blue:slow). Time interval of 60 sec. Time stamp in min:sec. Scale bar 20 µm. Representative of at least 3 independent experiments.
Video 9: Actin dynamics in varying geometries
Related to Fig. 4e-g and to Extended Data Fig. 5g. First part: T cells expressing Lifeact-mCherry migrating in a 5 µm-high microchannels with 6 µm, 12 µm, 24 µm and no serration, after addition of 10 mM EDTA. Brightfield (left panel) and Lifeact-mCherry (right panel, in black). Second part: Talin KO T cell expressing Lifeact-GFP in 5 µm-high serrated microchannel with 6 µm, 12 µm, 24 µm and no serration. Brightfield (left panel) and Lifeact-GFP TIRF (right panel, in black). Time interval of 2-3 sec, time stamps in min:sec. Scale bars 5 µm. Representatives of at least 3 independent experiments.
Video 10: Actin dynamics of primary T cells on smooth or nanoridged surfaces
Related to Extended Data Fig. 7: High-magnification (100X, inverted spinning-disc confocal microscope) of primary T cells expressing Lifeact-GFP placed under agarose on a non-adhesive, smooth substrate (first part) and on a non-adhesive substrate with engraved linear ridges (second part). Scale bar 5 µm. Time stamp in sec. Representative of at least 3 independent experiments.
Source data
Rights and permissions
About this article
Cite this article
Reversat, A., Gaertner, F., Merrin, J. et al. Cellular locomotion using environmental topography. Nature 582, 582–585 (2020). https://doi.org/10.1038/s41586-020-2283-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2283-z
This article is cited by
-
KLF5 regulates actin remodeling to enhance the metastasis of nasopharyngeal carcinoma
Oncogene (2024)
-
The amoeboid migration of monocytes in confining channels requires the local remodeling of the cortical actin cytoskeleton by cofilin-1
Scientific Reports (2024)
-
In silico analysis shows that dynamic changes in curvature guide cell migration over long distances
Biomechanics and Modeling in Mechanobiology (2024)
-
Biomimetic natural biomaterials for tissue engineering and regenerative medicine: new biosynthesis methods, recent advances, and emerging applications
Military Medical Research (2023)
-
Switch of cell migration modes orchestrated by changes of three-dimensional lamellipodium structure and intracellular diffusion
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.