Abstract
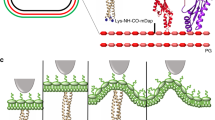
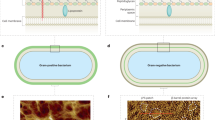
The primary structural component of the bacterial cell wall is peptidoglycan, which is essential for viability and the synthesis of which is the target for crucial antibiotics1,2. Peptidoglycan is a single macromolecule made of glycan chains crosslinked by peptide side branches that surrounds the cell, acting as a constraint to internal turgor1,3. In Gram-positive bacteria, peptidoglycan is tens of nanometres thick, generally portrayed as a homogeneous structure that provides mechanical strength4,5,6. Here we applied atomic force microscopy7,8,9,10,11,12 to interrogate the morphologically distinct Staphylococcus aureus and Bacillus subtilis species, using live cells and purified peptidoglycan. The mature surface of live cells is characterized by a landscape of large (up to 60 nm in diameter), deep (up to 23 nm) pores constituting a disordered gel of peptidoglycan. The inner peptidoglycan surface, consisting of more nascent material, is much denser, with glycan strand spacing typically less than 7 nm. The inner surface architecture is location dependent; the cylinder of B. subtilis has dense circumferential orientation, while in S. aureus and division septa for both species, peptidoglycan is dense but randomly oriented. Revealing the molecular architecture of the cell envelope frames our understanding of its mechanical properties and role as the environmental interface13,14, providing information complementary to traditional structural biology approaches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data that support the findings of this study are available in the Online Research Data (ORDA) figshare from the University of Sheffield with the identifier https://doi.org/10.15131/shef.data.11798898.
Code availability
The MATLAB code for determining glycan strand orientation can be found in Supplementary Information 2.
References
Turner, R. D., Vollmer, W. & Foster, S. J. Different walls for rods and balls: the diversity of peptidoglycan. Mol. Microbiol. 91, 862–874 (2014).
Vollmer, W. & Seligman, S. J. Architecture of peptidoglycan: more data and more models. Trends Microbiol. 18, 59–66 (2010).
Rojas, E. R. et al. The outer membrane is an essential load-bearing element in Gram-negative bacteria. Nature 559, 617–621 (2018).
Matias, V. R. F. & Beveridge, T. J. Native cell wall organization shown by cryo-electron microscopy confirms the existence of a periplasmic space in Staphylococcus aureus. J. Bacteriol. 188, 1011–1021 (2006).
Beeby, M., Gumbart, J. C., Roux, B. & Jensen, G. J. Architecture and assembly of the Gram-positive cell wall. Mol. Microbiol. 88, 664–672 (2013).
Misra, G., Rojas, E. R., Gopinathan, A. & Huang, K. C. Mechanical consequences of cell-wall turnover in the elongation of a Gram-positive bacterium. Biophys. J. 104, 2342–2352 (2013).
Turner, R. D. et al. Peptidoglycan architecture can specify division planes in Staphylococcus aureus. Nat. Commun. 1, 26 (2010).
Touhami, A., Jericho, M. H. & Beveridge, T. J. Atomic force microscopy of cell growth and division in Staphylococcus aureus. J. Bacteriol. 186, 3286–3295 (2004).
Dover, R. S., Bitler, A., Shimoni, E., Trieu-Cuot, P. & Shai, Y. Multiparametric AFM reveals turgor-responsive net-like peptidoglycan architecture in live streptococci. Nat. Commun. 6, 7193 (2015).
Andre, G. et al. Imaging the nanoscale organization of peptidoglycan in living Lactococcus lactis cells. Nat. Commun. 1, 27 (2010).
Turner, R. D., Mesnage, S., Hobbs, J. K. & Foster, S. J. Molecular imaging of glycan chains couples cell-wall polysaccharide architecture to bacterial cell morphology. Nat. Commun. 9, 1263 (2018).
Eskandarian, H. A. et al. Division site selection linked to inherited cell surface wave troughs in mycobacteria. Nat. Microbiol. 2, 17094 (2017).
Zhou, X. et al. Bacterial division. Mechanical crack propagation drives millisecond daughter cell separation in Staphylococcus aureus. Science 348, 574–578 (2015).
Green, E. R. & Mecsas, J. in Virulence Mechanisms of Bacterial Pathogens 5th edn (eds Kudva, I. T. et al.) 215–240 (ASM Press, 2016).
Bailey, R. G. et al. The interplay between cell wall mechanical properties and the cell cycle in Staphylococcus aureus. Biophys. J. 107, 2538–2545 (2014).
Weidenmaier, C. et al. Role of teichoic acids in Staphylococcus aureus nasal colonization, a major risk factor in nosocomial infections. Nat. Med. 10, 243–245 (2004).
Matias, V. R. F. & Beveridge, T. J. Lipoteichoic acid is a major component of the Bacillus subtilis periplasm. J. Bacteriol. 190, 7414–7418 (2008).
Kim, S. J., Chang, J. & Singh, M. Peptidoglycan architecture of Gram-positive bacteria by solid-state NMR. Biochim. Biophys. Acta 1848, 350–362 (2015).
Wheeler, R. et al. Bacterial cell enlargement requires control of cell wall stiffness mediated by peptidoglycan hydrolases. mBio 6, e00660 (2015).
Gan, L., Chen, S. & Jensen, G. J. Molecular organization of Gram-negative peptidoglycan. Proc. Natl Acad. Sci. USA 105, 18953–18957 (2008).
Hayhurst, E. J., Kailas, L., Hobbs, J. K. & Foster, S. J. Cell wall peptidoglycan architecture in Bacillus subtilis. Proc. Natl Acad. Sci. USA 105, 14603–14608 (2008).
Domínguez-Escobar, J. et al. Processive movement of MreB-associated cell wall biosynthetic complexes in bacteria. Science 333, 225–228 (2011).
Garner, E. C. et al. Coupled, circumferential motions of the cell wall synthesis machinery and MreB filaments in B. subtilis. Science 333, 222–225 (2011).
Dion, M. F. et al. Bacillus subtilis cell diameter is determined by the opposing actions of two distinct cell wall synthetic systems. Nat. Microbiol. 4, 1294–1305 (2019).
Mitchell, P. & Moyle, J. Autolytic release and osmotic properties of protoplasts from Staphylococcus aureus. J. Gen. Microbiol. 16, 184–194 (1957).
Daly, K. E., Huang, K. C., Wingreen, N. S. & Mukhopadhyay, R. Mechanics of membrane bulging during cell-wall disruption in Gram-negative bacteria. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 84, 041922 (2011).
Bisson-Filho, A. W. et al. Treadmilling by FtsZ filaments drives peptidoglycan synthesis and bacterial cell division. Science 355, 739–743 (2017).
Monteiro, J. M. et al. Peptidoglycan synthesis drives an FtsZ-treadmilling-independent step of cytokinesis. Nature 554, 528–532 (2018).
Lund, V. A. et al. Molecular coordination of Staphylococcus aureus cell division. eLife 7, e32057 (2018).
de Jong, I. G., Beilharz, K., Kuipers, O. P. & Veening, J. W. Live cell imaging of Bacillus subtilis and Streptococcus pneumoniae using automated time-lapse microscopy. J. Vis. Exp. 53, e3145 (2011).
Schirner, K. & Errington, J. The cell wall regulator σI specifically suppresses the lethal phenotype of mbl mutants in Bacillus subtilis. J. Bacteriol. 191, 1404–1413 (2009).
Turner, R. D., Foster, S. J. & Hobbs, J. K. in Bacterial Cell Wall Homeostasis: Methods and Protocols (ed. Hong, H. J.) 3–9 (Springer, 2016).
Kailas, L. et al. Immobilizing live bacteria for AFM imaging of cellular processes. Ultramicroscopy 109, 775–780 (2009).
Kumar, S. et al. Direct imaging of protein organization in an intact bacterial organelle using high-resolution atomic force microscopy. ACS Nano 11, 126–133 (2017).
Su, C. Mapping quantitative mechanical properties at molecular scale using Peak Force Tapping AFM. Microsc. Microanal. 16, 364–365 (2010).
Nečas, D. & Klapetek, P. Gwyddion: an open-source software for SPM data analysis. Cent. Eur. J. Phys. 10, 181–188 (2012).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
This work was funded by the BBSRC (grant no. BB/L006162/1), the EPSRC (grant nos. EP/M027430/1 and EP/J500124/1), The Wellcome Trust (grant no. 212197/Z/19/Z), the MRC (MR/N002679/1) and the UKRI Strategic Priorities Fund (grant no. EP/T002778/1). J.B. thanks The White Rose University Consortium for his studentship. L.P.-L. thanks The Florey Institute for her studentship. We acknowledge J. Errington and A. Grundling for provision of bacterial strains, and J. Sutton for preparation of the pbp3 and pbp4 sacculus samples. Electron microscopy was performed in the CryoEM Facility of the University of Sheffield.
Author information
Authors and Affiliations
Contributions
L.P.-L., J.B. and R.D.T. designed the study, performed the experiments (L.P.-L.: Figs. 2, 3, Extended Data Figs. 1–5, 7–10, Supplementary Information 2; J.B.: Figs. 1, 3, Extended Data Figs. 1, 2, 7; and R.D.T.: Fig. 3, Extended Data Figs. 7, 10), analysed and interpreted the data, and wrote the manuscript. S.K., J.S.W. and R.T. performed the experiments (S.K.: Extended Data Figs. 1, 7, 9; J.S.W.: Extended Data Fig. 6, Supplementary Information 1; and R.T.: Extended Data Fig. 7), and analysed and interpreted the data. N.M. developed the method for Fig. 1, provided support for the experiments (Fig. 1, Extended Data Figs. 1, 2) and wrote the manuscript. B.C. carried out the calculations in the paper (Supplementary Information 3) and wrote the manuscript. P.A.B. designed the study, interpreted the data and wrote the manuscript. S.J.F. and J.K.H. designed the study, interpreted the data, wrote the manuscript and directed the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Cécile Berne, Yves Dufrêne, Jan-Willem Veening, Kevin D. Young and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 S. aureus cells and sacculi: cell wall fine structure and other cell wall components.
a, Overview of individual S. aureus wild-type (WT) live cells trapped inside silicon holes. Topographical DS = 1,100 nm. b, Individual cell showing the mature mesh covering the top surface, high-pass filtered; filter size = 0.23 μm, vertical. c, Higher-resolution image within b, with the blue dashed box denoting the location of Fig. 1b. DS = 35 nm. d, Example of mesh from another cell. DS = 58 nm. e, S. aureus tarO mutant cells (this mutant lacks WTAs) trapped in silicon holes (provided by NuNano; see Methods for sample preparation). DS = 2,033 nm. f, Higher-resolution image of an individual cell showing the mature mesh. DS = 580 nm. 3D details: z aspect ratio = 1.0, pitch = 16.5° and plot type = height. g, Higher-resolution image from f showing the mature mesh. DS = 34 nm (comparison to Fig. 1b). h, Concentric rings on the tarO mutant (comparison to Extended Data Fig. 2e). DS = 46 nm. Direct comparison between the WT and the tarO mutant shows that there is no substantial difference in the organization of the cell wall in the absence of WTAs and that the features observed in the WT are not WTAs. i, S. aureus ltaS mutant cells in an SH1000 background were purified and a sacculus with its external PG structure facing upwards is shown. DS = 121 nm. j, Higher-resolution image from within i highlighting the nascent PG structure of this sacculus with similar features to those seen in the WT (see Fig. 2d as an example). DS = 14 nm. k, Higher-resolution image from within i corresponding to the mature PG structure of this sacculus with no apparent difference from the same structure observed in the WT cells (see Fig. 2b as an example). DS = 33 nm. This direct comparison between the WT and the ltaS mutant shows that there is no apparent difference in the organization of the cell wall in the absence of lipoteichoic acids and that the features observed in the WT are not attributable to lipoteichoic acids or WTAs, so they must be PG. l, 3D view of the S. aureus cell that images m and n were taken from. DS = 398 nm. 3D details: z aspect ratio = 0.5, pitch = 35°, light rotation = 36°, light pitch = 40°, light intensity = 60% and plot type = mixed. m, The periodic features along the glycan chain direction (white arrows pointing at single glycan strands clearly showing the periodicity). DS = 14 nm. n, Higher-resolution image from m (dashed white square) showing the 3.7-nm periodicity (the white arrows are the same as in m). DS = 10 nm. o, Profile taken along the blue rectangle in n, showing a 4-nm separation between adjacent features. We suggest that the periodic bumps are uncrosslinked pentapeptide side chains presented at the surface of every helical repeat. p, Schematic of the helical repeat of PG, with statistics from 24 measurements similar to those in o; mean ± s.d. is 3.7 ± 0.8 nm for the periodicity. This is in agreement with NMR data18. The morphologies from a–d and l were also each detected in at least five other biological independent repeats, and similar features to those in m and n in at least three biological repeats (see repository). For e–g, two biological independent repeats were performed. h–k come from one biological repeat.
Extended Data Fig. 2 S. aureus live-cell architecture.
a, At the interface between the most recent division plane and the mature cell wall, we find a sharp topographical step between the mature (mesh) and recently synthesized cell wall (concentric rings), see black arrows. DS = 535 nm. 3D details: z aspect ratio = 1, pitch = 21°, light rotation = 103°, light pitch = 49°, light intensity = 70% and plot type = mixed. b, Same image as a but processed with a plane fit at the third order showing the sharp interface in more detail. DS = 27 nm. c, Profile taken from along the dotted black line in a, showing a sharp change in height at the interface between the mesh and the rings (see red arrow, which corresponds to the red arrowhead in a). d–g, A series of images, each from a different cell, showing the presumed time evolution of the ring architecture. Samples naturally contained cells with a distribution of times since their last division, and hence show differing levels of maturation of the PG at the extracellular surface of the most recent division plane. In d, a recently revealed outer septal surface showing dense concentric rings is shown. The blue dashed box shows the location of Fig. 1e. DS = 18 nm. In e, a different cell showing what is probably the next stage of maturation is shown. The rings are still prominent (white arrows), but show larger spacing, greater height variation and pores have appeared in the surface. Features oriented approximately along the radial direction are also apparent. DS = 23 nm. In f, a different cell showing the subsequent stage where, underneath the rings (white arrows), the pores of the mesh have started to appear. DS = 38 nm. In g, the last stage of maturation of the cell wall is the completely disordered mesh without any traces of former ring structures. DS = 30 nm. The morphologies from a and b were also detected in at least one other biological independent repeat (see repository) and from d–g were also each detected in at least two other biological independent repeats (see repository).
Extended Data Fig. 3 S. aureus sacculi.
a, Height image of the same sacculus shown in Fig. 2a. This sacculus has been broken on the left side and emptied of all of the cell’s contents, allowing the difference in PG structure between the internal and the external surfaces to be clearly seen. Imaging buffer: 10 mM Tris pH 7.8 + 300 mM KCl. DS = 192 nm. b, The same sacculus as in a but imaged in air (dried with nitrogen flow). DS = 84 nm. c, Higher-resolution image from a taken in buffer showing the mature mesh (white arrows, 1–6, lower regions on the image, and blue arrows, 7–13, higher regions on the image). DS = 33 nm. d, The same region of mesh as in c but in air. This morphology has been previously termed ‘knobbles’7. The arrows were manually placed to point to the corresponding features (1–13) of c. DS = 11 nm. e, A sacculus fragment in buffer (10 mM Tris pH 7.8 + 300 mM KCl). The height of the sacculus (32.4 nm) was measured from the spacing of the modal values of Gaussian fits to the peaks corresponding to the sacculus and the substrate in a height histogram generated using the 1D statistical analysis tool in Gwyddion. f, The same sacculus fragment as e but imaged in air with an overall height of 17.5 nm measured in the same way as in e. Both images (e and f) have the same DS (65 nm). The same process was repeated with 24 other sacculi to produce the thickness data for Fig. 2k, using a positioning marker on either the sample or the substrate beneath to enable the same fragments to be located in both air and liquid. g–j, Analysis of the septation process. In g, the graph shows decreasing aperture size in the septum with constant septum diameter (n = 5 measures) of septation aperture for each septum shown in circles. The centre values represent the mean value and the error bars represent the maximum value excluding outliers. Septa 2, 3 and 5 are shown in h–j, respectively. Images in h–j show different stages of septal growth; each image was collected from a separate S. aureus sacculus. Images are presented in order of apparent septation progress, with the internal aperture in the middle of the septa getting smaller for cells at later stages of septation. DS: h, 548 nm; i, 512 nm; and j, 627 nm. The area corresponding to Fig. 2h was taken from the blue dashed box in h. The black arrow in j indicates the internal aperture in the centre of this nearly completed septum. Images in h–j were taken from cells grown at OD600 = 0.1, because this condition provided a higher number of partially formed septa in the sample. k, Higher-resolution image from a of the internal structure of the S. aureus cell wall, showing a randomly orientated dense mesh (black dashed square in Fig. 2a). DS = 10 nm.
Extended Data Fig. 4 Quantitative analysis of pores.
a, Image of Extended Data Fig. 2g of the mature PG structure from an S. aureus live-cell image represented in 3D and then tilted. The lateral resolution is lost and no clear information about the pores is visible with this representation. Therefore, a new analysis method was designed. The first step is to measure the maximum depth of this 3D image; the Zmax is calculated neglecting the areas below 0.3% and above 99% because they might contain artefacts and signal errors from the AFM. b, A 2D sectioning process is performed in the xy plane going down from Zmax, highlighting a set of certain depths. c, Slices were taken at intervals of approximately 3% of the total Zmax, including slices with the selected depths from b. d, All slices of different depths were super-imposed on top of the original image in Extended Data Fig. 2g, and five of the slices are highlighted in different colours (blue = 5 nm, green = 10 nm, orange = 15 nm, purple = 20 nm and white = 23 nm) showing the different areas covered by different depths. This is the complete image from which Fig. 1c was cropped. e, After converting all of the slices into a stack of images in ‘.tiff’ format, the AvizoLite software was used to represent the shape of the pores in 3D (view from the bottom) and the chosen set of depths were marked with the same colours as in d; the black arrow highlights the deepest pore in this image. f, This procedure was repeated for the mature PG structure from the B. subtilis live cell (Extended Data Fig. 7d). The 2D version of f, analogous to d, is shown in Fig. 3c. The deepest pore in the 3D representation is marked with a black arrow. g, h, A direct comparison of lateral slices in the xz plane, so that the lateral profile of the pores can be visualized. PG is coloured in grey in these images. For g, the lateral slice from S. aureus is extracted from the red dashed line marked in d, and for B. subtilis in h, it corresponds to the black dashed line in Fig. 3c. i–k, The steps followed to quantitatively compare the area of the pores on the outside of the cell and the inside of purified PG. In i, the raw data extracted from the AFM are shown. In j, the image has been processed with the Nanoscope Analysis software (third-order plane-fit) to remove the offset, tilt and curvature of the cell. The image has been converted to greyscale, and a despeckle filter (3 × 3 median filter) has been applied with ImageJ/Fiji38. In k, a 2D slice in the xy plane was made at a depth of 50% of Zmax for six images of live S. aureus and four images of internal S. aureus sacculi using the ImageJ/Fiji threshold tool. Slices were then binarized with pores in black and PG in white. The ‘analyse particle’ tool from ImageJ/Fiji was used to measure the pores. This outputs a list of individual pores with their associated area and a number for each pore. l, In some cases, in particular when several pores lie within a larger depression on the surface, pores are artificially merged. These were manually removed from the data set by comparing the binary image with the greyscale image. Correctly assigned (retained) pores are shown in blue, and three artificially merged (discarded) pores are shown in black. m–o, Histograms of pore area. In m, a pore area histogram was calculated for 310 pores from 6 separate images for the external surface of live S. aureus cells. The bin width is 150 nm2. In n, a subset of the same data set as m (denoted by the blue dashed box) binned at 10 nm2 intervals is shown. In o, a pore area histogram was calculated for 158 pores from 4 separate images of the internal surface of S. aureus sacculi, binned at 2 nm2 intervals. The distributions of pore area are non-normal and positively skewed for both the internal and the external surfaces. Of the pores in the external surface, 90% have an area of <900 nm2 (grey dashed line in m). For the internal surface, 90% of the pores have an area of <47 nm2 (grey dashed line in o). The mode and quantiles Q1–3 (25%, 50% and 75%, respectively) are marked on the figure distributions in n and o with dashed lines.
Extended Data Fig. 5 Mechanistic insight into PG hydrolysis and synthesis.
a, Sacculi of S. aureus WT imaged in air, with different fragments containing single layers of PG (green dashed arrows) and double layers (black dotted arrow). DS = 75 nm. b, Sacculi from the S. aureus sagB mutant in an SH1000 background imaged in air. The fragments have two distinct regions with different thicknesses (pink arrows and orange arrowheads). These are distinct from the double-layer fragments (black dotted arrow). DS = 75 nm. c, Graph comparing the thickness of the cell wall in air and liquid between the sagB mutant and the WT. The box plot consists of boxes from the 25–75% percentile, the middle line represents the median, the black star represents the mean, and the error bars are the maximum and minimum values excluding any outliers (using the 1.5 interquartile range). The two different regions of the sagB cell wall are separately binned and labelled ‘sagB thin’ and ‘sagB thick’. Measurements were taken using the same approach as the values for the graph in Fig. 2k, Extended Data Fig. 3. The number of sacculi measured n = 25 for each bin. Mean ± s.d.: sagB thin air = 12 ± 3 nm, sagB thin liquid (liq) = 24 ± 7 nm, sagB thick air = 21 ± 1.5 nm, sagB thick liquid = 42 ± 5.7 nm, WT air and WT liquid corresponded to the same values as in Fig. 2k (left data, PG + WTA). All measurements in liquid were performed in buffer: 300 mM KCl + 10 mM Tris, pH 8. d, S. aureus sagB mutant sacculus in liquid with its external surface facing upwards. This sacculus is clearly divided into two sections: the region on the left is thinner than the region on the right. This irregularity in height was visualized in several sacculi from cells with this mutation. DS = 64 nm. e, Higher-resolution image from the thicker section in d corresponding to the finer mature cell wall architecture. This mesh is still in an early transition from rings, with some features reminiscent of rings that are visualized following the same orientation (blue arrows). These sorts of structures are uncommon on WT sacculi at the same stage in the cell cycle (exponential phase OD600 = 0.5–0.7). However, in the sagB mutant, it was more common to find this early transition from rings to mesh. DS = 11 nm. f, Higher-resolution image from the thinner section in d showing the finer nascent cell wall architecture (that is, rings). The inset shows a lower-resolution image of the ring architecture in a different sacculus. The features from this image are not significantly different from the WT nascent PG architecture, such as in Fig. 2d. However, almost every sacculus containing nascent architecture in the sagB mutant had this part of the cell wall being thinner than the rest of the sacculus, something that is not seen in the WT. DS = 9 nm, inset DS = 29 nm. g, Sacculus fragment with the inner cell wall architecture facing upwards. DS = 14 nm. h, Higher-resolution image from g showing the fine inner structure of the sagB mutant, with no clear difference from the very tight and disordered structure seen in WT sacculi (for example, see Fig. 2f). DS = 4 nm. i, Sacculi from S. aureus pbp3 mutant cells in an SH1000 background. Here we show the fine external structure, with no apparent difference between this and the S. aureus WT mature external structure. It is important to highlight that PBP3 is not an essential PG synthesis enzyme, and the cells can survive without it and still maintain their spheroid shape. DS = 55 nm. j, The fine structure of an S. aureus pbp3 mutant sacculus with the internal PG surface exposed, which is similar to that seen in WT S. aureus. DS = 4 nm. k, Sacculus from S. aureus pbp4 mutant cells in an SH1000 background showing the nascent PG structure, with the concentric rings facing upwards. It is important to highlight that PBP4 is not an essential PG synthesis enzyme either, and the cells can survive without it and still maintain their spheroid shape. DS = 28 nm. l, Higher-resolution image taken within k showing the fine structure of the nascent PG surface: the concentric rings. There is no significant difference compared to the S. aureus WT nascent structure, except the fraction of sacculi with rings in a random area is higher in this sample than in the WT. DS = 8 nm. m, Sacculus showing a small area of the mature PG structure (green arrow), with its external surface facing upwards. The rest of the sacculus is covered by concentric rings that are already in the late phase of the transition to mesh (black arrow pointing to the centre of the concentric rings; the boundary with the mature structure is indicated by the red dashed line) and the newest generation fragment with rings (blue arrowhead; the boundary with the oldest rings is marked by the red dotted line). It is important to highlight that this is the first time that two perpendicular planes of rings have been visualized at exponential phase in our experience. OD600 = 0.6. DS = 79 nm. n, Higher-resolution image from m showing the fine structure of the mature external cell wall of an S. aureus pbp4 mutant, with no apparent difference between this and the S. aureus WT mature external structure (see Fig. 2b, Extended Data Fig. 1). DS = 44 nm. o, Sacculus showing the fine internal structure of the S. aureus pbp4 mutant cell wall, with no apparent difference compared to the S. aureus WT internal PG architecture (see Fig. 2f). DS = 10 nm. These data show that removal of non-essential PBPs makes only relatively minor differences to the wall architecture.
Extended Data Fig. 6 Tomogram of purified frozen hydrated S. aureus sacculus.
S. aureus WT cells were purified and treated with HF to remove all WTAs, so that only a pure PG sacculus was imaged. Tomograms were captured of five sacculi, and an example of a single sacculus is given in Supplementary Video 1. a, A 13-nm thick tomogram slice from the first stop point on Supplementary Video 1, which corresponds to the external cell wall surface showing disordered structures throughout the area of the sacculus (blue arrows). b, A 13-nm thick tomogram slice from the second stop point on Supplementary Video 1 corresponding to the other side of the cell wall, the smooth internal structure, which is in agreement with the AFM data presented in Fig. 2. c, An intermediate 13-nm thick tomogram slice between a and b where the outline of the exterior surface of cell wall is visualized showing a disordered structure presumably formed by pores. d, As in c but with the exterior surface highlighted in yellow; examples of individual pores analysed are indicated by the blue arrows. e, Graph plotting the area of each pore versus the cumulative fraction of the total pore area of all pores analysed. The blue circles correspond to measures from the AFM images of S. aureus cells, measured at a height of 50% of the total image height scale (data also presented in Fig. 2i), and the black dashed line is the linear fit to these data. The orange squares are individual measurements of pore diameter from the image in d converted into pore area by assuming the pores are circular, and the black dotted line is the linear fit to these data, showing a similar slope to the AFM data. The red stars show 50% of the cumulative area, that is, half of the total pore area consists of pores smaller or larger than this value. The values calculated from the linear equations for both types of data are: 39 nm for AFM data and 42 nm for the cryo-EM data. This slight difference is most likely due to the lack of smaller pores analysed by cryo-EM, which is highlighted with a transparent green area in the graph. The total pores analysed by AFM were n = 311 from n = 6 images, and by cryo-EM were n = 30 from n = 1 image. For more details on Supplementary Video 1, see Supplementary Information 1. f, The same tomogram slice as a highlighting a region of the sacculus that is smooth on both the inside and the outside. Cryo-EM does not discern the ring structure as it views in transverse section, but we suggest that this region of the sacculus is most probably made up of rings. g, The region highlighted in yellow in f, which has been traced with a spline function that was then used to straighten the section. h, The same tomogram slice as a highlighting in yellow a region with a rough external surface, which we suggest corresponds to the mesh architecture observed with AFM. i, The yellow region of h straightened as in g. j, An AFM image of a sacculus showing two distinguishable areas (reproducing Fig. 2c). The top part consists of a smooth surface with occasional pores, where the glycan strands are organized as concentric rings. On the bottom part of the sacculus, the mature cell wall shows a highly topographical surface. k, Image from f cropped to the same size as the AFM profile shown in l. l, AFM profile in yellow from the red dashed line in j, showing an almost flat surface. m, Image from i cropped to the same size as the AFM profile in n. n, AFM profile taken along the blue dotted line in j. We note the qualitative similarity between the two data sets, despite the very different contrast mechanisms of the two techniques. o, An example tomogram slice from a tomogram of a different sacculus. p, The region of the cell wall highlighted in yellow, straightened as in g. q, Another example AFM image of the mesh architecture. r, The area indicated with the blue line in p, cropped to the same size as the AFM profile in s, highlighting the rough texture of the external cell wall. s, AFM profile taken along the blue dashed line in q.
Extended Data Fig. 7 B. subtilis live cells and sacculi.
a, High-pass-filtered image (filter size: 0.2 μm, horizontal) of the B. subtilis cell cylinder in exponential phase attached to a Cell-Tak-coated substrate. The inset (I) shows the unfiltered height data. DS = 1,417 nm. b–d, Higher-resolution images of the mature cell wall (mesh) within a in locations marked with white dashed squares. DS = 31 nm (b), 27 nm (c) and 21 nm (d). The external surface of B. subtilis is covered by disordered mesh all along the cylinder with the exception of the newly synthesized poles (which have a concentric ring architecture). e, A sacculus from a B. subtilis cell, showing the internal surface of a pole. We can differentiate between poles and completely formed septa because of the folded material on top (blue arrow). No septa were observed with this feature. DS = 130 nm. f, Higher-resolution image from within e showing the fine structure of the internal pole, similar to the internal structure of S. aureus, suggesting that this architecture is characteristic of the spherical part of Gram-positive cells. DS = 87 nm. g, External part of the pole facing upwards, with the interface between the pole and the rest of the cylinder indicated by blue arrows. DS = 118 nm. We presume that this image corresponds to a mature pole and that the concentric rings have become mesh, suggesting that the structural reorganization in time from rings to mesh also occurs in B. subtilis poles. h, Example of a newly synthesized pole facing upwards with a sharp interface between the pole and the rest of the cylinder (blue arrows). This pole displays very tight concentric rings in contrast to g and similar to Fig. 3e, f. DS = 800 nm. i, The same image as in h but processed with Gwyddion and JPK software (flattened to 0th order with the mask drawn manually over the sacculus and path level to remove lines), allowing the concentric rings (white arrow) to be visualized more clearly. The results from images shown in e–i support a common process of synthesis and maturation of the cell wall on the spherical parts of Gram-positive bacteria in both spherical and rod-shaped species. Images shown in e–i were taken with JPK NanoWizard 3 in QI mode (see Methods). Images shown in e–h were processed with Gwyddion36, the image shown in i was processed with JPK data processing software, and the colour palettes are different. j, Two fragments from B. subtilis sacculi. The fragment on the left corresponds to the internal structure facing upwards (white arrowhead), and the fragment on the right has the external structure of the cylinder, mesh, facing upwards (blue arrow). The assumed cylindrical axis is marked (red dashed arrow). DS = 118 nm. k, Higher-resolution image from within j of the internal cell wall. Some larger pores are visualized here and there is some predominant orientation indicated by the angle distribution of the fibres shown in the inset, in agreement with the cylindrical axis (red dashed arrow). The method for obtaining these distributions is explained in Extended Data Fig. 8. DS = 46 nm. l, Example of B. subtilis sacculus fragments showing the two different structures: the internal cell wall in the two fragments (white arrowheads) and the top part of the cylinder corresponding to the external cell wall (blue arrow). DS = 146 nm. The distinction between this image and that shown in j is that these fragments are part of the cylinder that once belonged to the same cell. The blue dashed box marks the area corresponding to Fig. 3i, and the assumed cylindrical axis is marked (red dashed arrow). m, Higher-resolution image from l of the internal cell wall. This strongly aligned structure (see inset (I)) is the most commonly found in these sacculi in contrast with k. As the fragments in l are not totally broken, we assume that this will be more similar to the native structure of the cell wall. DS = 19 nm. n–p, The different structures from B. subtilis sacculi that have been HF treated so the WTAs have been removed: the mesh seen on the external surface (DS = 30 nm) (n); strands inside (DS = 30 nm), the circumferential axis direction is given by the red dashed arrow (strand orientation is shown in the inset (I)) (o); and the partially formed septum (DS = 130 nm) (p). This shows that none of the previous structures was due to teichoic acid organization, which is consistent with our findings for S. aureus cells.
Extended Data Fig. 8 B. subtilis strand orientation analysis and mechanistic insights.
a, B. subtilis mreB, mbl, mreBH, rsgI mutant sacculus of the internal structure of the cell wall, with a defined ridge (green arrow). This mutant lacks the MreB family of proteins, which leads to a spheroidal shape. DS = 411 nm. b, Higher-resolution image from a showing the fine structure of the internal surface of a B. subtilis mreB, mbl, mreBH, rsgI mutant sacculus, with a disordered structure (see inset (I), which indicates a broad distribution of strand orientations) similar to the internal structure of S. aureus, suggesting that when Gram-positive cells cannot maintain a rod shape, no cylindrical orientation is required. DS = 52 nm. c, Data used to obtain the orientation distributions for the glycan strands shown in the inset in b. Lines were hand drawn over the images along the clearly visible strands, using short straight strokes. A custom MATLAB routine (Supplementary Information 2) was then used to obtain the orientation distribution of these lines. The routine successfully recognized ~70% of the lines and determined their orientation. d, Mutations encoding the MreB family, shown in a–c, were created in a B. subtilis rsgI background strain. We prepared a purified sample from this B. subtilis rsgI mutant as a control. This image corresponds to the internal cell wall structure (the red dashed arrow indicates the circumferential axis direction). DS = 76 nm. e, Higher-resolution image from within d showing glycan strand orientation along the short axis of the rod morphology of the bacteria (dashed red arrow and inset (I)). This sample had the same characteristics and structures as the WT strain. DS = 23 nm. f, Data used to plot the orientation distribution rose from the inset in e. g, As an additional control, a sacculi sample of B. subtilis WT was grown as the mutants. This image shows another internal cell wall structure from a broken sacculus (the red dashed arrow indicates the circumferential axis direction). DS = 125 nm. h, Higher-resolution image from within g showing glycan strand orientation cylindrical along the short axis of the rod morphology of the bacteria (see dashed red arrow and inset (I)). DS = 16 nm. i, The same strand orientation analysis as shown in h was applied to the external surface of B. subtilis cells (see Fig. 3c) and the angle distribution is broad (inset (I)), indicating that the glycan strands on the external surface have no predominant orientation in contrast to the internal surface. j, Data used to plot the orientation distribution rose from the inset in i, using the same procedure as c and f. This experiment provides mechanistic insight into the cylindrical orientation architecture in the inside of the rod shape of the Gram-positive species B. subtilis. The protein complex of MreB, Mbl and MBH is vital for the synthesis of the PG in this cylindrical architecture22,23,24.
Extended Data Fig. 9 B. subtilis septa.
A comparison of B. subtilis cell wall fragments that were acquired in liquid with the same fragments acquired in air, allowing our current findings to be understood in the context of our previous study21. a, b, B. subtilis septa at different stages of formation, imaged in liquid. DS = 70 nm (a) and 158 nm (b). In both cases, a disordered mesh structure is apparent with pores penetrating deep into the cell wall. c, d, Corresponding images from the same septa as a and b in the same orientation in air. DS = 31 nm (c) and 60 nm (d). Some features that were observed in our previous work, such as the concentric rings around the circumference of the septa (green arrowheads) and the ridges around the outer diameter of the septa (blue arrow), were identified. We attribute these features to the superposition of the mesh structures on both sides of the septa, with the concentric rings formed in the middle between them, coupled with the influence of differential drying. The experiment of correlating S. aureus sacculi in ambient and liquid environments (Extended Data Fig. 3) to measure the difference in thickness, was repeated for B. subtilis sacculus fragments and also showed a significant difference. e, f, Examples of single-layer fragments, marked 1 and 2, imaged in water (e) and air (f). The thickness of each individual fragment was measured using the 1D statistical analysis from the Gwyddion software (see Extended Data Fig. 3e for details). g, Graph showing the thickness for 19 sacculi measured in both air and liquid. There is a significant increase in thickness in liquid after a two-tailed, paired t-test was performed (P = 1.2 × 10−9; see Methods). The box plot consists of boxes from the 25–75% percentile, the middle line represents the median, the black star indicates the mean, and the error bars are the maximum and minimum values excluding any outliers (using the 1.5 interquartile range). Mean ± s.d.: 9 ± 1 nm (air) and 34 ± 5 nm (liquid). These results are in agreement with the sacculus thickness of B. subtilis measured by transmission electron microscopy (figure 4 of ref. 5). h, Higher-resolution image from Fig. 3j. The internal surface of this partially formed septum is composed of randomly orientated mesh. Large pores are present that go partially through the cell wall (blue arrows). DS = 137 nm. i, In contrast to h, in this partially formed septum, most of the material has been synthesized already, and the pores are not as deep or as wide as in h (yellow arrows). DS = 160 nm. These two septa differ in the size of their aperture by 100 nm, meaning that they are probably representing consecutive stages of septal formation. The structural difference between them suggests that there is more than one type of synthesis happening simultaneously: one type that occurs at the leading edge of the aperture making it smaller, and the other type that is responsible for in-filling the pores that go partially through the cell wall as seen in h, finally resulting in the tighter mesh seen in i.
Extended Data Fig. 10 Schematic structure of the molecular architecture of the Gram-positive bacterial cell wall.
The proposed PG organization derived from AFM imaging for both S. aureus and B. subtilis. The top-centre and bottom-left indicate areas of distinct morphology within the cell wall during the division process. Insets show schematized representative AFM images of the different architectures within the cell wall, all displayed at the same scale.
Supplementary information
Supplementary Information
This file contains: 1, legend for Video showing a tomogram of a purified frozen hydrated S. aureus sacculus; 2, MATLAB code for obtaining peptidoglycan strand orientation; and 3, Estimation of the critical pore size to maintain plasma membrane integrity.
Supplementary Video 1
Video showing a tomogram of a purified frozen hydrated S. aureus sacculus. (Seconds 0 to 4) The video “Video_SI1.mp4” shows the tomogram from the bottom to the top. There is a first stopping point which corresponds to where image ED6b was extracted from and then a second stopping point which corresponding to ED6a. Each slice shown on the tomogram correspond to 1.3 nm. Images ED6a-d correspond to 10 slices thick, which means they represent 13 nm of the tomogram. (Seconds 4 to 7) Later on in the video, the slices are shown from top to bottom, with an isosurface (in green) manually drawn showing the outline of the sacculus.
Rights and permissions
About this article
Cite this article
Pasquina-Lemonche, L., Burns, J., Turner, R.D. et al. The architecture of the Gram-positive bacterial cell wall. Nature 582, 294–297 (2020). https://doi.org/10.1038/s41586-020-2236-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2236-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.