Abstract
Prolonged mitosis often results in apoptosis1. Shortened mitosis causes tumorigenic aneuploidy, but it is unclear whether it also activates the apoptotic machinery2. Separase, a cysteine protease and trigger of all eukaryotic anaphases, has a caspase-like catalytic domain but has not previously been associated with cell death3,4. Here we show that human cells that enter mitosis with already active separase rapidly undergo death in mitosis owing to direct cleavage of anti-apoptotic MCL1 and BCL-XL by separase. Cleavage not only prevents MCL1 and BCL-XL from sequestering pro-apoptotic BAK, but also converts them into active promoters of death in mitosis. Our data strongly suggest that the deadliest cleavage fragment, the C-terminal half of MCL1, forms BAK/BAX-like pores in the mitochondrial outer membrane. MCL1 and BCL-XL are turned into separase substrates only upon phosphorylation by NEK2A. Early mitotic degradation of this kinase is therefore crucial for preventing apoptosis upon scheduled activation of separase in metaphase. Speeding up mitosis by abrogation of the spindle assembly checkpoint results in a temporal overlap of the enzymatic activities of NEK2A and separase and consequently in cell death. We propose that NEK2A and separase jointly check on spindle assembly checkpoint integrity and eliminate cells that are prone to chromosome missegregation owing to accelerated progression through early mitosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gascoigne, K. E. & Taylor, S. S. Cancer cells display profound intra- and interline variation following prolonged exposure to antimitotic drugs. Cancer Cell 14, 111–122 (2008).
Michel, L. S. et al. MAD2 haplo-insufficiency causes premature anaphase and chromosome instability in mammalian cells. Nature 409, 355–359 (2001).
Lin, Z., Luo, X. & Yu, H. Structural basis of cohesin cleavage by separase. Nature 532, 131–134 (2016).
Wirth, K. G. et al. Separase: a universal trigger for sister chromatid disjunction but not chromosome cycle progression. J. Cell Biol. 172, 847–860 (2006).
Galluzzi, L. et al. Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 25, 486–541 (2018).
Letai, A. et al. Distinct BH3 domains either sensitize or activate mitochondrial apoptosis, serving as prototype cancer therapeutics. Cancer Cell 2, 183–192 (2002).
Villunger, A. et al. p53- and drug-induced apoptotic responses mediated by BH3-only proteins puma and noxa. Science 302, 1036–1038 (2003).
Kamenz, J. & Hauf, S. Time to split up: dynamics of chromosome separation. Trends Cell Biol. 27, 42–54 (2017).
Hellmuth, S. et al. Securin-independent regulation of separase by checkpoint-induced shugoshin–MAD2. Nature https://www.doi.org/10.1038/s41586-020-2182-3 (2020).
Taylor, S. S. & McKeon, F. Kinetochore localization of murine Bub1 is required for normal mitotic timing and checkpoint response to spindle damage. Cell 89, 727–735 (1997).
Li, Y. & Benezra, R. Identification of a human mitotic checkpoint gene: hsMAD2. Science 274, 246–248 (1996).
Rankin, S., Ayad, N. G. & Kirschner, M. W. Sororin, a substrate of the anaphase-promoting complex, is required for sister chromatid cohesion in vertebrates. Mol. Cell 18, 185–200 (2005).
Tang, Z., Sun, Y., Harley, S. E., Zou, H. & Yu, H. Human Bub1 protects centromeric sister-chromatid cohesion through Shugoshin during mitosis. Proc. Natl Acad. Sci. USA 101, 18012–18017 (2004).
Bennett, A. et al. Inhibition of Bcl-xL sensitizes cells to mitotic blockers, but not mitotic drivers. Open Biol. 6, 160134 (2016).
Haschka, M. D. et al. The NOXA-MCL1-BIM axis defines lifespan on extended mitotic arrest. Nat. Commun. 6, 6891 (2015).
Sloss, O., Topham, C., Diez, M. & Taylor, S. Mcl-1 dynamics influence mitotic slippage and death in mitosis. Oncotarget 7, 5176–5192 (2016).
Topham, C. et al. MYC is a major determinant of mitotic cell fate. Cancer Cell 28, 129–140 (2015).
Clem, R. J. et al. Modulation of cell death by Bcl-XL through caspase interaction. Proc. Natl Acad. Sci. USA 95, 554–559 (1998).
Michels, J. et al. Mcl-1 is required for Akata6 B-lymphoma cell survival and is converted to a cell death molecule by efficient caspase-mediated cleavage. Oncogene 23, 4818–4827 (2004).
Barclay, L. A. et al. Inhibition of pro-apoptotic BAX by a noncanonical interaction mechanism. Mol. Cell 57, 873–886 (2015).
Brouwer, J. M. et al. Conversion of Bim-BH3 from activator to inhibitor of Bak through structure-based design. Mol. Cell 68, 659–672.e659 (2017).
Czabotar, P. E. et al. Bax crystal structures reveal how BH3 domains activate Bax and nucleate its oligomerization to induce apoptosis. Cell 152, 519–531 (2013).
Dai, H. et al. Transient binding of an activator BH3 domain to the Bak BH3-binding groove initiates Bak oligomerization. J. Cell Biol. 194, 39–48 (2011).
Wang, C. & Youle, R. J. Predominant requirement of Bax for apoptosis in HCT116 cells is determined by Mcl-1’s inhibitory effect on Bak. Oncogene 31, 3177–3189 (2012).
Große, L. et al. Bax assembles into large ring-like structures remodeling the mitochondrial outer membrane in apoptosis. EMBO J. 35, 402–413 (2016).
Salvador-Gallego, R. et al. Bax assembly into rings and arcs in apoptotic mitochondria is linked to membrane pores. EMBO J. 35, 389–401 (2016).
McArthur, K. et al. BAK/BAX macropores facilitate mitochondrial herniation and mtDNA efflux during apoptosis. Science 359, eaao6047 (2018).
Day, C. L. et al. Solution structure of prosurvival Mcl-1 and characterization of its binding by proapoptotic BH3-only ligands. J. Biol. Chem. 280, 4738–4744 (2005).
Dewson, G. et al. Bak activation for apoptosis involves oligomerization of dimers via their α6 helices. Mol. Cell 36, 696–703 (2009).
Kudo, N. R. et al. Role of cleavage by separase of the Rec8 kleisin subunit of cohesin during mammalian meiosis I. J. Cell Sci. 122, 2686–2698 (2009).
Hayes, M. J. et al. Early mitotic degradation of Nek2A depends on Cdc20-independent interaction with the APC/C. Nat. Cell Biol. 8, 607–614 (2006).
Wolthuis, R. et al. Cdc20 and Cks direct the spindle checkpoint-independent destruction of cyclin A. Mol. Cell 30, 290–302 (2008).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Wild, T. et al. The spindle assembly checkpoint is not essential for viability of human cells with genetically lowered APC/C activity. Cell Rep. 14, 1829–1840 (2016).
Tighe, A., Johnson, V. L., Albertella, M. & Taylor, S. S. Aneuploid colon cancer cells have a robust spindle checkpoint. EMBO Rep. 2, 609–614 (2001).
Weitzel, D. H. & Vandré, D. D. Differential spindle assembly checkpoint response in human lung adenocarcinoma cells. Cell Tissue Res. 300, 57–65 (2000).
Bamford, S. et al. The COSMIC (Catalogue of Somatic Mutations in Cancer) database and website. Br. J. Cancer 91, 355–358 (2004).
Hernando, E. et al. Molecular analyses of the mitotic checkpoint components hsMAD2, hBUB1 and hBUB3 in human cancer. Int. J. Cancer 95, 223–227 (2001).
Stemmann, O., Zou, H., Gerber, S. A., Gygi, S. P. & Kirschner, M. W. Dual inhibition of sister chromatid separation at metaphase. Cell 107, 715–726 (2001).
Wolf, P. G., Cuba Ramos, A., Kenzel, J., Neumann, B. & Stemmann, O. Studying meiotic cohesin in somatic cells reveals that Rec8-containing cohesin requires Stag3 to function and is regulated by Wapl and sororin. J. Cell Sci. 131, jcs212100 (2018).
Hellmuth, S. et al. Human chromosome segregation involves multi-layered regulation of separase by the peptidyl-prolyl-isomerase Pin1. Mol. Cell 58, 495–506 (2015).
Hellmuth, S., Böttger, F., Pan, C., Mann, M. & Stemmann, O. PP2A delays APC/C-dependent degradation of separase-associated but not free securin. EMBO J. 33, 1134–1147 (2014).
Hellmuth, S., Gutiérrez-Caballero, C., Llano, E., Pendás, A. M. & Stemmann, O. Local activation of mammalian separase in interphase promotes double-strand break repair and prevents oncogenic transformation. EMBO J. 37, e99184 (2018).
To, T. L. et al. Rational design of a GFP-based fluorogenic caspase reporter for imaging apoptosis in vivo. Cell Chem. Biol. 23, 875–882 (2016).
Agudelo, D. et al. Marker-free coselection for CRISPR-driven genome editing in human cells. Nat. Methods 14, 615–620 (2017).
Wehr, M. C. et al. Monitoring regulated protein–protein interactions using split TEV. Nat. Methods 3, 985–993 (2006).
Murray, A. W. Cell cycle extracts. Methods Cell Biol. 36, 581–605 (1991).
McGuinness, B. E., Hirota, T., Kudo, N. R., Peters, J. M. & Nasmyth, K. Shugoshin prevents dissociation of cohesin from centromeres during mitosis in vertebrate cells. PLoS Biol 3, e86 (2005).
Lukinavičius, G. et al. SiR-Hoechst is a far-red DNA stain for live-cell nanoscopy. Nat. Commun. 6, 8497 (2015).
Hanson, K. M. & Finkelstein, J. N. An accessible and high-throughput strategy of continuously monitoring apoptosis by fluorescent detection of caspase activation. Anal. Biochem. 564–565, 96–101 (2019).
Gorr, I. H., Boos, D. & Stemmann, O. Mutual inhibition of separase and Cdk1 by two-step complex formation. Mol. Cell 19, 135–141 (2005).
Liu, L., Spurrier, J., Butt, T. R. & Strickler, J. E. Enhanced protein expression in the baculovirus/insect cell system using engineered SUMO fusions. Protein Expr. Purif. 62, 21–28 (2008).
Butt, T. R., Edavettal, S. C., Hall, J. P. & Mattern, M. R. SUMO fusion technology for difficult-to-express proteins. Protein Expr. Purif. 43, 1–9 (2005).
Hames, R. S. & Fry, A. M. Alternative splice variants of the human centrosome kinase Nek2 exhibit distinct patterns of expression in mitosis. Biochem. J. 361, 77–85 (2002).
Inoshita, S. et al. Phosphorylation and inactivation of myeloid cell leukemia 1 by JNK in response to oxidative stress. J. Biol. Chem. 277, 43730–43734 (2002).
Maurer, U., Charvet, C., Wagman, A. S., Dejardin, E. & Green, D. R. Glycogen synthase kinase-3 regulates mitochondrial outer membrane permeabilization and apoptosis by destabilization of MCL-1. Mol. Cell 21, 749–760 (2006).
Acknowledgements
We thank T. U. Mayer for suggesting the concept of the DMC, R. Youle and M. Orth for cell lines, D. Pfeiffer for help with the 2D SIM, S. Heidmann, T. Klecker, and P. Wolf for critical reading of the manuscript, and J. Hübner and M. Hermann for technical assistance. This work was supported by a grant (STE997/4-2) from the Deutsche Forschungsgemeinschaft (DFG) to O.S.
Author information
Authors and Affiliations
Contributions
S.H. carried out all experiments. S.H. and O.S. co-designed the research and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Andreas Villunger, Hongtao Yu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
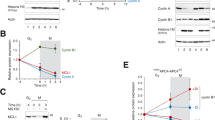
Extended Data Fig. 1 Premature activation of separase rather than loss of cohesion triggers DiM.
a, Premature sister chromatid separation (PCS) was quantified by chromosome spreading from siRNA-transfected HeLa-K cells at different times after release from RO-3306/G2 arrest. b, Immunoblots of cells from a. c–e, HeLa-K cells transfected with the indicated siRNAs and cultured in the presence of SiR-Hoechst49 and a fluorogenic caspase 3/7 reporter50 were analysed by video fluorescence microscopy. Shown are representative stills (c; scale bar, 10 μm), immunoblots (d), and cell fate profiles (e). f–h, HeLa-K cells expressing the caspase 3 reporter ZipGFP44 were transfected with the indicated siRNAs, supplemented with SiR-Hoechst and followed by video microscopy. At least 100 cells were counted per time point and condition. Shown are representative stills (f), immunoblots (g), and line graphs (h) of the percentages of GFP-positive (apoptotic) cells. The ZipGFP plasmid also expresses mCherry as a control. Scale bar, 10 μm.
Extended Data Fig. 2 Characterization of MCL1 (and BCL-XL) cleavage by separase (and caspase 3).
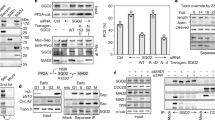
a, Sequence stretches of wild-type BCL-XL and MCL1 and TEV variants thereof. Differing amino acids are in bold. Arrowheads show protease cleavage sites. b, Separase and caspase 3 cleave MCL1 after Arg176 and Asp157, respectively, as mapped by in vitro-expressed fragments. Following incubation with NEK2A, CDK1–cyclin B1, PLK1 and ATP, separase, or caspase 3, in vitro-translated 35S-MCL1ΔTM was treated with a surplus of λ-PPase and analysed by SDS–PAGE and autogradiography. 35S-MCL1 fragments representing reported caspase 3 cleavage fragments and putative separase cleavage fragments served as molecular weight standards. c, Mouse MCL1 is cleaved by separase after 154-DXXR-157. Autoradiograph of in vitro-translated, NEK2A/separase-treated mouse 35S-MCL1. PD, protease-dead (C2029S); Ac, active (P1127A); ΔTM, transmembrane domain deleted. d, Immunoblots of taxol-arrested Hek293T cells transfected with siRNAs and expression plasmids as indicated. During separase-triggered DiM, MCL1 cleavage stimulates BCL-XL cleavage and vice versa, which—at least in the case of MCL1—is mediated by the corresponding N-terminal fragment (Extended Data Fig. 4b). e, Co-depletion of SGO2 and securin induces cleavage of MCL1 and BCL-XL followed by apoptosis in non-transformed cells. Immunoblots of siRNA-transfected, taxol-arrested hTERT RPE1 cells expressing ZipGFP. f, MCL1 is cleaved after R176 during DiM of SGO2- and securin-depleted cells. Immunoblots of extracts and MCL1 immunoprecipitates (IP) from taxol-arrested Hek293T cells transfected with the indicated siRNAs. IVT, in vitro translated. g, Mouse MCL1 and BCL-XL are cleaved and DiM is triggered upon separase deregulation in mouse cells. NIH/3T3s cells transfected with siRNAs and treated with taxol and staurosporine as indicated were analysed by immunoblotting.
Extended Data Fig. 3 Characterization of separase cleavage fragments of MCL1 and BCL-XL.
a, The C-terminal separase cleavage fragments of MCL1 and BCL-XL preferentially bind BIM and BAD, respectively. Experimental setup and immunoblots of the indicated immunoprecipitation from taxol-arrested Hek293T cells co-expressing His6–MCL1-CΔTM, BCL-XL-CΔTM–Flag, HA–BAD, and HA–BIM. b, BCL-XL and BCL-XL-C bind BAK and BAD, respectively. Immunoblots of Flag immunoprecipitation from transfected, mitotic Hek293T cells expressing the indicated Flag-tagged BCL-XL fragments together with HA-tagged BAK and BAD. c, MCL1-N interacts with BAK. Experimental setup and immunoblots of lysate and consecutive Flag and His6 immunoprecipitations from transfected, mitotic Hek293T cells co-expressing Flag–TEV-BAK, His6–BAK, and MCL1-N. TEV protease supplementation of lysate served as a negative control. Self-interaction of BAK is mutually exclusive with binding of MCL1-N. d, Separase-induced DiM is suppressed by knock-down of MCL1 and BCL-XL. Quantification and representative images of siRNA-transfected, mitotic Hek293T cells cultivated in the presence of mitoTracker and a fluorogenic caspase 3/7 reporter before fixation and Hoechst staining. At least 100 cells each were counted. Scale bar, 5 μm. e, Immunoblots of cells from d. f, MCL1-N and -C promote DiM. Plot of early (dark grey) and late (light grey) apoptosis as judged by flow cytometric analysis of propidium iodide and annexin V staining of Hek293T cells transfected with siRNAs and expression vectors for transgenic MCL1 (fragments) and supplemented with taxol and BCL-XL inhibitor WEHI-539 (WEHI) as indicated. g, Representative 2D scatter plots of cells from f and their interpretation. h, Induction of DiM by MCL1-C does not require taxol treatment but does require Thr301. Cell fate profiles of HeLa-K cells expressing the indicated MCL1 fragments and cultured in the presence of SiR-Hoechst and a fluorogenic caspase 3/7 reporter. i, Immunoblot of cells from h and Extended Data Fig. 10a.
Extended Data Fig. 4 MCL1-N enables BAD to replace BAK as an interactor of BCL-XL and enhances cleavage of BCL-XL by separase.
a, MCL1-N causes a switch in the binding partner of BCL-XL from BAK to BAD. Immunoblots of BCL-XL immunoprecipitation from MCL1-depleted or control-treated, mitotic Hek293T cells co-expressing HA–BAK, HA–BAD and, where indicated, various MCL1 fragments. Separase was not deregulated in this experiment, which is why BCL-XL is not cleaved. b, Separase-dependent BCL-XL cleavage in cells is primarily stimulated by MCL1-N. Immunoblots of SGO2/securin/MCL1 triple-depleted or control-treated, prometaphase Hek293T cells expressing the indicated transgenic MCL1 variants. c, Separase prefers BCL-XL in complex with BAD as a substrate rather than BAK. Experimental setup and immunoblots of HA immunoprecipitation from taxol-arrested Hek293T cells expressing NEK2A-ΔMR and either HA–BAK or HA–BAD plus MCL1-N. Before SDS–PAGE, samples were incubated with inactive (PD) or active (ac) separase or control treated (−). d, MCL1-N promotes apoptosis by a positive feedback mechanism. MCL1-N competitively displaces the BH4 domain of BCL-XL from BAK. Similar to cleavage by separase, this breaks the cooperative binding of BCL-XL to BAK, resulting in BH3-only proteins, such as BAD, excluding BAK from BCL-XL. At the same time, this renders BCL-XL a better separase substrate. MCL1-N acts catalytically, being released from BAK upon self-interaction and pore formation by the latter (dotted arrow; Extended Data Fig. 3c).
Extended Data Fig. 5 Self-interaction of pro-apoptotic MCL1-C shares characteristics with pore formation by BAK/BAX but requires mitosis-specific phosphorylation.
a–c, MCL1-C exhibits mitosis-specific self-interaction, which requires the transmembrane domain and an accessible BH3-binding groove. Immunoblots of Flag immunoprecipitation from mitotic or interphase Hek293T cells expressing MCL1 variants and supplemented with the MCL1 inhibitor A-1210477 as indicated. Blockade of the MCL1–BAK interaction served as a control for the effectiveness of A-1210477. d, The homotypic interaction of MCL1-C is mutually exclusive with BIM binding. Top, experimental setup; bottom, immunoblots of consecutive Flag and His6 immunoprecipitations from Hek293T cells expressing HA–BIM together with Flag–TEV- and His6-tagged forms of either wild-type MCL1-C or the T301A variant. X, irrelevant lane between HA–BIM control (asterisk) and His6 immunoprecipitation samples. The T301A mutation prevents self-interaction of MCL1-C but not its association with BIM.
Extended Data Fig. 6 BAK/BAX-independent release of cytochrome c by separase deregulation or MCL1-C expression.
a, b, Time-resolved immunoblots of cytosol and mitochondria-containing fractions from SGO2/securin-depleted or mock-transfected BAK−/−, BAX−/− and parental HCT116 cells released from a G2-arrest at t = 0 min. c, Immunoblots of taxol-arrested BAK−/−, BAX−/− and parental HCT116 cells that were transfected with siRNA and supplemented with staurosporine as indicated. Note the absence of MCL1 and BCL-XL cleavage during staurosporine-induced apoptosis and suppression of siSGO2/siPTTG1-induced DiM by co-depletion of MCL1 and BCL-XL. Grey lines within panels are between lanes that were not directly juxtaposed but nevertheless originated from the same gel. d, Time-resolved immunoblots of cytosol and mitochondria-containing fractions from Flag–MCL1-C-expressing BAK−/−, BAX−/− and parental HCT116 cells released from G2-arrest at t = 0 min.
Extended Data Fig. 7 The pro-apoptotic effect of MCL1-C and BAX requires a negative charge at the end of α-helix 6.
a, b, Immunofluorescence micrographs of taxol-arrested Hek293T cells transfected with the indicated siRNAs. Shown are maximum projections of 20 z-stacks (a) or a single, deconvoluted plane (b). The interruption of TOM20 rings by MCL1 dots is consistent with a cross-section through a fragmented mitochondrion containing an MCL1 ring. Scale bars, 3 μm. c, MCL1-C is likely to form macropores into the mitochondrial outer membrane. Immunofluorescence 2D SIM of SGO2/securin-depleted Hek293T cells undergoing DiM. Note the absence of the mitochondrial outer membrane marker TOM20 from the centres of MCL1 rings. Scale bar, 0.5 μm. d, Immunoblots of extracts and Flag immunoprecipitations from interphase or mitotic Hek293T cells expressing Flag-tagged MCL1-C-WT (T), MCL1-C(T301A) (A), or MCL1-C(T301E) (E). e, Identifying serine and threonine residues that affect the pro-apoptotic nature of MCL1-N and MCL1-C. Immunoblot of MCL1-depleted, taxol-arrested Hek293T cells expressing the indicated siRNA-resistant variants of MCL1-N or MCL1-C. BCL-XL is not cleaved during apoptosis if separase remains inhibited. f, g, Aurora B kinase phosphorylates MCL1-C in vitro, probably at position 301 primarily. Immunoblots and autoradiographs of in vitro-translated wild-type MCL1-CΔTM and variants thereof after incubation with the indicated kinases and inhibitors in the presence of γ33P-ATP. Activity of the recombinant kinase was confirmed using model substrates (f, lower panels). KD, kinase-dead; RO, RO-3306; BI, BI-2536; ZM, ZM-447439; MBP, myelin basic protein; H1, histone H1. h, Local alignment of MCL1 and BAX. Vertical lines and colons mark identical and chemically similar residues, respectively; dashes represent gaps. i, Immunoblots of interphase or mitotic Hek293T cells expressing BAX with (+) or without (Δ) TM and with Glu (E), Ala (A), or Thr (T) at position 146. j, Model of MOMP by MCL1-C homo-oligomerization. The indicated conformational changes are inspired by knowledge about BAK/BAX pore formation and their hierarchy was chosen to best explain our data; that is, why the T301A mutation abolished self-interaction of MCL1-C but left BIM binding unaffected (Extended Data Fig. 5d). Minus signs represents phosphorylated Thr301 and plus signs represent a nearby basic residue.
Extended Data Fig. 8 NEK2A kinase turns MCL1 and BCL-XL into separase substrates.
a, Autoradiographs and immunoblots of combined kinase (33P-labelling) and cleavage (fragment-generation) assays using bacterially expressed, purified MCL1ΔTM and kinases, specific inhibitors and active (ac) or inactive (PD) separase, as indicated. b, CDK1–cyclin B1 and PLK1 enhance separase-dependent cleavage of NEK2A-phosphorylated MCL1. Immunoblots of cleavage assays using bacterially expressed, purified MCL1ΔTM and the indicated combination of kinases and separase variants. c, When immunoprecipitated from NEK2A-ΔMR-expressing, SAC-arrested cells, endogenous MCL1 is efficiently cleaved by separase in vitro. Left, experimental setup; right, immunoblots of cleavage assay combining active or inactive separase with MCL1 immunoprecipitation from prometaphase Hek293T cells expressing MCL1ΔTM–Flag and, where indicated, Flag–NEK2A-ΔMR or N-terminally truncated (Δ86) Flag–cyclin A2. d, Cleavage of mouse (M.m) MCL1 by separase also requires NEK2A-dependent phosphorylation. In vitro-translated 35S-MYC6-M.m.MCL1ΔTM was incubated with separase, caspase 3 and NEK2A/ATP as indicated. Reactions were resolved by SDS–PAGE and analysed by autoradiography and immunoblotting. KD, kinase-dead (K37M). e, NEK2A and (to a lesser extent) CDK1/2–cyclin A2 sensitize BCL-XL to separase. Autoradiography of cleavage assay combining in vitro-translated 35S-RAD21 (positive control) or 35S-BCL-XLΔTM with kinases/ATP and separase variants as indicated. f, Autoradiography of kinase assays (33P-labelling) using model substrates and the kinases from d. g–i, The NEK2A-related NEK2B does not support separase-dependent MCL1 cleavage. g, Schematics and C-terminal sequences (dashed box) of NEK2A and NEK2B, which arise by alternative splicing of the same gene54. NEK2A-specific, C-terminal degrons (KEN box and MR-tail) are underlined. h, Both NEK2A and NEK2B can phosphorylate the model substrate MBP. i, NEK2B cannot phosphorylate MCL1. Left, experimental setup; top right, kinase assay; bottom right, cleavage assay combining MCL1ΔTM with NEK2A, NEK2B, staurosporine (kinase inhibitor) and active separase as indicated.
Extended Data Fig. 9 Mapping cleavage-relevant phosphorylation sites within MCL1 and BCL-XL.
a, b, Cleavage of MCL1 by separase essentially requires NEK2A-dependent phosphorylation of Ser60 and Thr163. Autoradiographs and immunoblots of combined kinase (33P-labelling) and cleavage (fragment-generation) assays. Prior to analysis, in vitro-translated, wild-type MCL1ΔTM and variants thereof were incubated with active NEK2A, γ33P-ATP, staurosporine, and active (ac) or inactive (PD) separase as indicated. In vivo phosphorylation of Ser159 and Thr163 has previously been reported55,56. In vivo phosphorylation of Ser60 was detected by a phosphorylation-specific antibody (Fig. 4c, Extended Data Fig. 11d, e). c, Separase-dependent cleavage of MCL1(S/T60,159,163D/E) is independent of NEK2A. Immunoblots and autoradiography of combined kinase (33P) and cleavage assay. d–f, In vitro-translated, 35S-labelled wild-type BCL-XLΔTM and variants thereof were incubated with the indicated kinases (+) or reference buffers (−) and active separase before SDS–PAGE and autoradiography. d, NEK2A-stimulated cleavage of BCL-XL by separase essentially requires Ser4 and Ser164. e, CDK1/2–cyclin A2-stimulated cleavage of BCL-XL by separase essentially requires Ser62. In vivo phosphorylation of Ser62 has been previously reported48. f, Separase-dependent cleavage of BCL-XL(S4,62,164D) occurs independently of NEK2A and CDK1/2–cyclin A2. g, Autoradiograph and immunoblot of kinase assay using in vitro-translated, wild-type BCL-XLΔTM or its S4,62,164A variant and γ33P-ATP in combination with the indicated kinases (+) or reference buffers (−).
Extended Data Fig. 10 Stabilized NEK2A and constitutively cleavable MCL1- and BCL-XL variants result in DiM upon activation of separase in anaphase.
a, Cell fate profiles of HeLa-K cells expressing NEK2A-WT or NEK2A-ΔMR and cultured in the presence of SiR-Hoechst and a fluorogenic caspase 3/7 reporter. b, Immunoblots of time-resolved separase and MCL1 immunoprecipitations from NEK2A-ΔMR-expressing or control HeLa-K cells released from taxol arrest by addition of ZM-447439 (ZM) at t = 0 min. c, d, HeLa-K cells expressing the indicated variants of MCL1 and BCL-XL were analysed as in a. (Dephosphorylated) CDC27 served as a marker for late mitosis.
Extended Data Fig. 11 NEK2A stabilization preferentially kills MCL1-overexpressing cells.
a, b, Representative images and immunoblots of quantitative analysis shown in Fig. 4b. Scale bar, 5 μm. c, Immunoblots of NEK2A-ΔMR-expressing Hek293T cells transfected with siMCL1 and expression plasmids for His6–MCL1 as indicated. MCL1 and PARP cleavage were quantified densitometrically. n.d., not determined. d, MCL1-Ser60 is phosphorylated in early mitosis only. Untransfected or NEK2A-ΔMR-expressing HeLa-K cells were released from thymidine arrest for 8 h and then analysed by (immuno)fluorescence microscopy using Hoechst and the indicated antibodies. Scale bar, 5 μm. e, Chemical abrogation of the SAC triggers DiM. Immunoblots of reversine- or siRNA-treated HeLa-K cells synchronized as in Fig. 4c. Dephosphorylation of CDC27 into a sharp, fast-migrating band serves as a marker of late mitosis. f, TP53−/− cells and parental HCT116 cells were depleted of SGO2 and securin, taxol-arrested and analysed by immunoblotting.
Extended Data Fig. 12 MCL1 cleavage by separase and the pro-apoptotic effect of MCL1-C is conserved in non-mammalian vertebrates.
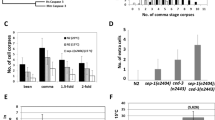
a, 35S-labelled, NEK2A/ATP-treated full-length Mcl1 from X. laevis, X. tropicalis and D. rerio were incubated with separase variants and caspase 3 as indicated, and analysed by autoradiography. b, c, Separase can cleave after ExxxR motifs. b, X. laevis and D. rerio Mcl1 are cleaved by separase after 136-ExxxR-140 and 84-ExxR-87, respectively. The indicated full-length Mcl1 or fragments thereof were analysed as in a. c, Changing the pseudo-substrate sequence of securin to ExxxR turns it into a separase substrate. Endogenous human SGO2 and securin were depleted by RNAi and replaced by the indicated variants9. These were then assessed for cleavage by immunoprecipitation and western blotting. d, Xenopus S3 cells transfected to express ZipGFP and the indicated forms of X. laevis Mcl1 were analysed by immunoblotting (left) and video microscopy (representative phase contrast images on right). Scale bar, 50 μm.
Supplementary information
Supplementary Data
This file contains gel source data.
Rights and permissions
About this article
Cite this article
Hellmuth, S., Stemmann, O. Separase-triggered apoptosis enforces minimal length of mitosis. Nature 580, 542–547 (2020). https://doi.org/10.1038/s41586-020-2187-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2187-y
This article is cited by
-
Mechanisms of BCL-2 family proteins in mitochondrial apoptosis
Nature Reviews Molecular Cell Biology (2023)
-
Marker-free co-selection for successive rounds of prime editing in human cells
Nature Communications (2022)
-
Severe cellular stress activates apoptosis independently of p53 in osteosarcoma
Cell Death Discovery (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.