Abstract
Arenaviruses can cause severe haemorrhagic fever and neurological diseases in humans and other animals, exemplified by Lassa mammarenavirus, Machupo mammarenavirus and lymphocytic choriomeningitis virus, posing great threats to public health1,2,3,4. These viruses encode a large multi-domain RNA-dependent RNA polymerase for transcription and replication of the viral genome5. Viral polymerases are one of the leading antiviral therapeutic targets. However, the structure of arenavirus polymerase is not yet known. Here we report the near-atomic resolution structures of Lassa and Machupo virus polymerases in both apo and promoter-bound forms. These structures display a similar overall architecture to influenza virus and bunyavirus polymerases but possess unique local features, including an arenavirus-specific insertion domain that regulates the polymerase activity. Notably, the ordered active site of arenavirus polymerase is inherently switched on, without the requirement for allosteric activation by 5′-viral RNA, which is a necessity for both influenza virus and bunyavirus polymerases6,7. Moreover, dimerization could facilitate the polymerase activity. These findings advance our understanding of the mechanism of arenavirus replication and provide an important basis for developing antiviral therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The cryo-EM density maps and corresponding coordinates have been deposited to the EMDB and the PDB, respectively. The accession numbers are listed as follows: apo LASV L (EMD-0706, 6KLC), apo MACV L monomer (EMD-0707, 6KLD), apo MACV L dimer (EMD-0708), MACV L–vRNA monomer (EMD-0709, 6KLE) and MACV L–vRNA dimer (EMD-0710, 6KLH). Details about RNA product characterization and uncropped images for autoradiographs are provided in the Supplementary information. All other data are available from the authors on reasonable request.
References
McCormick, J. B., Webb, P. A., Krebs, J. W., Johnson, K. M. & Smith, E. S. A prospective study of the epidemiology and ecology of Lassa fever. J. Infect. Dis. 155, 437–444 (1987).
Childs, J. E. et al. Human–rodent contact and infection with lymphocytic choriomeningitis and Seoul viruses in an inner-city population. Am. J. Trop. Med. Hyg. 44, 117–121 (1991).
Charrel, R. N. & de Lamballerie, X. Arenaviruses other than Lassa virus. Antiviral Res. 57, 89–100 (2003).
Gao, G. F. From “A”IV to “Z”IKV: attacks from emerging and re-emerging pathogens. Cell 172, 1157–1159 (2018).
Buchmeier, M. J., De La Torre, J. C. & Peters, C. J. in Fields Virology 5th edn, Vol. 2 (eds Knipe, D. M. & Howley, P. M.) 1791–1827 (Lippincott/The Williams & Wilkins, 2007).
Pflug, A., Guilligay, D., Reich, S. & Cusack, S. Structure of influenza A polymerase bound to the viral RNA promoter. Nature 516, 355–360 (2014).
Gerlach, P., Malet, H., Cusack, S. & Reguera, J. Structural insights into bunyavirus replication and its regulation by the vRNA promoter. Cell 161, 1267–1279 (2015).
Hass, M., Gölnitz, U., Müller, S., Becker-Ziaja, B. & Günther, S. Replicon system for Lassa virus. J. Virol. 78, 13793–13803 (2004).
Kranzusch, P. J. et al. Assembly of a functional Machupo virus polymerase complex. Proc. Natl Acad. Sci. USA 107, 20069–20074 (2010).
Brunotte, L. et al. Domain structure of Lassa virus L protein. J. Virol. 85, 324–333 (2011).
Raju, R. et al. Nontemplated bases at the 5′ ends of Tacaribe virus mRNAs. Virology 174, 53–59 (1990).
Polyak, S. J., Zheng, S. & Harnish, D. G. 5′ Termini of Pichinde arenavirus S RNAs and mRNAs contain nontemplated nucleotides. J. Virol. 69, 3211–3215 (1995).
Meyer, B. J. & Southern, P. J. Concurrent sequence analysis of 5′ and 3′ RNA termini by intramolecular circularization reveals 5′ nontemplated bases and 3′ terminal heterogeneity for lymphocytic choriomeningitis virus mRNAs. J. Virol. 67, 2621–2627 (1993).
Morin, B. et al. The N-terminal domain of the arenavirus L protein is an RNA endonuclease essential in mRNA transcription. PLoS Pathog. 6, e1001038 (2010).
Reguera, J. et al. Comparative structural and functional analysis of bunyavirus and arenavirus cap-snatching endonucleases. PLoS Pathog. 12, e1005636 (2016).
Lehmann, M. et al. Role of the C terminus of Lassa virus L protein in viral mRNA synthesis. J. Virol. 88, 8713–8717 (2014).
Rosenthal, M. et al. Structural insights into reptarenavirus cap-snatching machinery. PLoS Pathog. 13, e1006400 (2017).
Reich, S. et al. Structural insight into cap-snatching and RNA synthesis by influenza polymerase. Nature 516, 361–366 (2014).
Thierry, E. et al. Influenza polymerase can adopt an alternative configuration involving a radical repacking of PB2 domains. Mol. Cell 61, 125–137 (2016).
Hengrung, N. et al. Crystal structure of the RNA-dependent RNA polymerase from influenza C virus. Nature 527, 114–117 (2015).
Pyle, J. D. & Whelan, S. P. J. RNA ligands activate the Machupo virus polymerase and guide promoter usage. Proc. Natl Acad. Sci. USA 116, 10518–10524 (2019).
Peng, Q. et al. Structural insight into RNA synthesis by influenza D polymerase. Nat. Microbiol. 4, 1750–1759 (2019).
Fan, H. et al. Structures of influenza A virus RNA polymerase offer insight into viral genome replication. Nature 573, 287–290 (2019).
Sánchez, A. B. & de la Torre, J. C. Genetic and biochemical evidence for an oligomeric structure of the functional L polymerase of the prototypic arenavirus lymphocytic choriomeningitis virus. J. Virol. 79, 7262–7268 (2005).
Morin, B., Kranzusch, P. J., Rahmeh, A. A. & Whelan, S. P. The polymerase of negative-stranded RNA viruses. Curr. Opin. Virol. 3, 103–110 (2013).
Liang, B. et al. Structure of the L protein of vesicular stomatitis virus from electron cryomicroscopy. Cell 162, 314–327 (2015).
Vogel, D., Rosenthal, M., Gogrefe, N., Reindl, S. & Günther, S. Biochemical characterization of the Lassa virus L protein. J. Biol. Chem. 294, 8088–8100 (2019).
Perez, J. T. et al. Influenza A virus-generated small RNAs regulate the switch from transcription to replication. Proc. Natl Acad. Sci. USA 107, 11525–11530 (2010).
Wilda, M., Lopez, N., Casabona, J. C. & Franze-Fernandez, M. T. Mapping of the Tacaribe arenavirus Z-protein binding sites on the L protein identified both amino acids within the putative polymerase domain and a region at the N terminus of L that are critically involved in binding. J. Virol. 82, 11454–11460 (2008).
Gong, P. & Peersen, O. B. Structural basis for active site closure by the poliovirus RNA-dependent RNA polymerase. Proc. Natl Acad. Sci. USA 107, 22505–22510 (2010).
Te Velthuis, A. J. W. & Oymans, J. Initiation, elongation, and realignment during influenza virus mRNA synthesis. J. Virol. 92, e01775-17 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Sehnal, D. et al. MOLE 2.0: advanced approach for analysis of biomacromolecular channels. J. Cheminform. 5, 39 (2013).
Acknowledgements
We thank all staff members at the Center of Biological Imaging (CBI), Institute of Biophysics (IBP), Chinese Academy of Sciences (CAS), the Cryo-EM Center of Southern University of Science and Technology (SUST), and the Cryo-EM department of the National Center for Protein Science Shanghai (NCPSS), CAS, for assistance with data collection; are grateful to G. Wang (The Core Facilities at School of Life Sciences, Peking University), T. Yang and staff members in the EM department of the State Key Laboratory of Membrane Biology, Institute of Zoology (IOZ), CAS, Beijing, for technical support in electron microscope operation; and appreciate the assistance of H. Song and F. Zhao (the Research Facility Center at Beijing Institutes of Life Science, CAS) in the FortéBio Octet experiment. This study was supported by the Strategic Priority Research Program of CAS (XDB29010000), the National Science and Technology Major Project (2018ZX10101004), the External Cooperation Program of CAS (153211KYSB20160001), the Advanced Customer Cultivation Project of Wuhan National Biosafety Laboratory, CAS (no. 2018ACCP-MS11) and the National Natural Science Foundation of China (NSFC) (81871658). M.W. is supported by the National Science and Technology Major Project (2018ZX09711003). R.P. is supported by the Young Elite Scientist Sponsorship Program (YESS) by China Association for Science and Technology (CAST) (2018QNRC001). G.F.G. is supported partly as a leading principal investigator of the NSFC Innovative Research Group (81621091). Y.S. is supported by the Excellent Young Scientist Program, the NSFC (81622031) and the Youth Innovation Promotion Association of CAS (2015078).
Author information
Authors and Affiliations
Contributions
R.P., G.F.G. and Y.S. conceived the study. X.X., J.J., M.W., Y.W. and X.B. purified the protein samples and conducted biochemical studies. R.P., Q.P., S.L. and P.W. prepared the cryo-EM specimens and collected data. R.P. conducted the image processing and reconstruction. R.P. and J.Q. built the atomic models. R.P., X.X., M.W., J.Q., G.F.G. and Y.S. analysed the structures. R.P., X.X., Q.P., G.F.G. and Y.S. wrote the manuscript. All authors participated in the discussion and manuscript editing. R.P., X.X., J.J. and M.W. contributed equally to this work. Y.S. supervised all of the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Juan C. de la Torre, Ming Luo and Aartjan te Velthuis for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cryo-EM analysis of apo LASV and MACV L proteins.
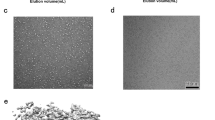
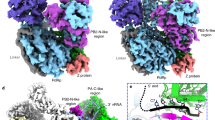
a, A representative cryo-EM micrograph of MACV L (out of ~2,500 micrographs). b, 2D class average images of LASV and MACV L particles (at least two rounds of classification were performed). A few MACV L particles were observed to form homodimers (red boxes, right panel). Some potential LASV L dimeric particles were also observed (red boxes, left panel) but the abundance was very low and did not result in distinguished 2D class averages. c–f, Brief image processing workflows for LASV and MACV L data sets and Euler angle distribution of each reconstruction. g, Local resolution maps for monomeric structures of LASV and MACV L proteins. h, The FSC curves for each reconstruction. The FSC 0.143 and 0.5 cut-off values are indicated by blue and orange dashed lines, respectively. i, j, Representative density of LASV (i) and MACV (j) L monomeric reconstructions.
Extended Data Fig. 2 Comparison of individual domains of FluPol, LACV, LASV and MACV polymerases.
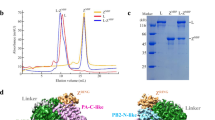
a, Similar overall architecture of these polymerases. b, LASV and MACV L proteins lack the arch loop as in FluBPol and LACV L, which is highlighted by a red dashed oval. c, Arenavirus polymerases lack a bridge domain in the C-terminal region of RdRp but possess a pendant domain insertion between the α-ribbon and the finger domain compared with LACV L. The pendant domain in LASV L was not resolved and its supposed location is represented by a pink dashed oval. d, In LACV L (PDB ID: 5AMQ), the CTD was truncated in protein expression. For FluBPol (PDB ID: 4WRT) and LASV and MACV L, the CTD is present in the protein sample but not resolved due to flexibility. NTD, N-terminal domain. e, SEC and SDS–PAGE profiles of LASV and MACV L pendant deletion mutants. OD, optical density. f, Binding kinetics of LASV and MACV L pendant deletion mutants to the 3′-vRNA or 3′-cRNA promoter strands determined by BLI experiments. KD, dissociation constant. g, In vitro activity assays for WT polymerase and mutants. The correct and incomplete-denatured product bands are indicated by red asterisks and blue arrowheads, respectively. The uncropped autoradiograph images are shown in Supplementary Fig. 1. The data shown in e–g are representative of three independent experiments using different protein preparations.
Extended Data Fig. 3 Knock-on effect of 5′-vRNA hook binding for FluPol and LACV polymerase.
a–h, The B-factor maps for apo FluCPol (a), vRNA-bound FluAPol (b) or FluBPol (c), 5′-hook-free (d) or 5′-hook-bound (e) LACV L, and apo LASV (f), apo MACV (g) and apo vesicular stomatitis virus (VSV) (h) L structures are shown as tube models. The radius and colour are correlated to the B-factor of corresponding atoms, with a larger radius and red colour representing high B-factor values and a smaller radius and blue colour representing low B-factor values. The finger-tip loop in LACV L without 5′-hook binding is disordered and is represented by a red dotted line. Corresponding regions in apo LASV, MACV or VSV L are highly ordered.
Extended Data Fig. 4 mRNA sequencing and verification by denatured PAGE experiment.
a, The migration profiles of the same transcription reaction samples in gels containing different concentrations of urea. b, The expected sequence of correct mRNA product and sequencing results for the products generated by LASV and MACV polymerases. The capped primer sequence is underlined with three terminal residues (red) complementary to the template. See Supplementary Methods and Supplementary Discussion for details about these experiments. The images show the complete gels without cropped features. The data shown are representative of two independent experiments using different protein preparations.
Extended Data Fig. 5 Biochemical characterization of LASV and MACV L proteins and their interactions with RNA promoters.
a, b, SEC profiles of LASV and MACV L and their interactions with vRNA promoter strands. c, Schematic diagram of the arenavirus vRNA promoter. The non-templated G (position 0) at the 5′-vRNA terminus is coloured in red. Residues 1–7 of the 3′-vRNA that are responsible for L protein interaction are highlighted with a green background. The 5′ terminus (residues 1–12) binds to L protein in a separate binding site, indicated with a grey background, for which the complementarity with the 3′-vRNA terminus is represented by dotted lines. The distal ends (residues 13–19) of 5′-vRNA and 3′-vRNA are supposed to form a duplex, for which the base pairs are represented by solid lines. d–g, Binding kinetics of LASV and MACV L proteins to 3′-vRNA or 3′-cRNA strands determined by BLI experiments. The data shown in this figure are representative of three independent experiments using different protein preparations.
Extended Data Fig. 6 Cryo-EM analysis of the MACV L–vRNA complex.
a, A representative cryo-EM micrograph of promoter-bound MACV L (out of ~3,000 micrographs). b, Representative 2D class average images (three rounds of classification were performed). Most of the particles exist as homodimers (red boxes). c–f, Brief 3D classification and refinement procedures for the dimeric and monomeric particles. The masks for splitting dimeric particles into monomeric forms are indicated by red dotted outlines in c. The Euler angle distribution of each reconstruction is shown. g, Local resolution maps for monomeric (left) and dimeric (right) structures of the MACV L–vRNA complex. The RNA-binding groove is indicated by a black arrowhead in each structure. The densities of the α-ribbon and pendant domains (highlighted by red dashed circles) become less ordered than the apo form, indicating conformational changes in this region. h, The FSC curves for each reconstruction. The FSC 0.143 and 0.5 cut-off values are indicated by blue and orange dashed lines, respectively. i, Representative density of the dimeric interface and 3′-vRNA.
Extended Data Fig. 7 The 3′-vRNA-binding site and conservation analysis for arenavirus polymerases.
a–c, The similar 3′-vRNA-binding site in MACV polymerase (a), LACV polymerase (PDB ID: 5AMQ) (b) and FluPol (PDB ID: 6KUT) (c). The protein structures are shown as cartoons (coloured by domains), and the bound RNA is shown as a surface model. d, e, Electrostatic surface of the 3′-vRNA-binding site in MACV (d) and LASV (e) L proteins. f, Conservation analysis for arenavirus polymerases. Residues H1189 and D1329 (numbered according to Tacaribe virus (TCRV) polymerase), which are potentially involved in Z protein interaction29, are located near the NTP entrance. The clamp domain (blue dotted oval) shows high variability, whereas the dimerization interface (red dotted oval) and the 3′-vRNA-binding site (black dotted rectangle) are relatively conserved. g, Sequence alignment of the clamp region. Residues with >70% similarity in one site are coloured in red and indicated by blue rectangles. The strictly identical residues are highlighted with a red background. The two insertions of Old World arenavirus polymerases relative to New World arenaviruses are highlighted with green boxes. GTOV, Guanarito virus; IPPYV, Ippy virus; JUNV, Junin virus; LCMV, lymphocytic choriomeningitis virus; LUJV, Lujo virus; MBLV, Mobala virus; MOPV, Mopeia virus; OLIV, Oliveros virus; PIRV, Pirital virus; SABV, Sabia virus; WWAV, Whitewater Arroyo virus.
Extended Data Fig. 8 Different conformations of LASV and MACV L proteins for RNA synthesis.
a, Superposition of LASV (dark) and MACV (light) L structures. The four major domains with different conformations are highlighted, including the endonuclease (blue), α-ribbon (yellow), pendant (pink) and clamp (green) domains. The rest of the main body is coloured in grey. The pendant domain of LASV is modelled by superimposing the α-ribbon and pendant of MACV L with the α-ribbon helices of LASV and is highlighted by a dotted oval. The CTD is not resolved in both structures and is represented with a black dotted oval. The predicted template entrance is indicated by a black arrow, which is closed by the α-ribbon and pendant in MACV L. b, Schematic model of the different conformational states of LASV and MACV L proteins. The domains of LASV L are represented by solid colours, and those of MACV L are shown with dotted outlines. c, Model of replicating RdRp based on the structures of LASV L and the poliovirus elongation complex30. The catalytic site is indicated by a red asterisk. The 3′-vRNA (black) and product (blue) strands are separated by the lid domain. The NTP entrance and template or product exits are indicated by arrows. d, Inside tunnels within the LASV L structure for RNA synthesis, including the template entrance, the NTP entrance and the template (black) or product (blue) exits. The catalytic site is indicated by a red asterisk.
Extended Data Fig. 9 Modelling of the LASV L homodimer and the contacting interfaces.
a, Structural model of the LASV L homodimer. The two protomers are coloured by domains: one in dark colours and the other in light colours. b, Close-up view of the predicted dimeric interface. The electrostatic surface of one protomer is shown to visualize the complementary charge patterns. Positive and negative charges are represented in blue and red, respectively. The key contacting residues are shown as sticks. c, Sequence alignment for key regions of the dimeric interface, with three key motifs highlighted by red boxes. Residues with >70% similarity in one site are coloured in red and indicated by blue rectangles. The strictly identical residues are highlighted with a red background. d, Close-up view of the steric clash between endonuclease domains (blue) within the two LASV L protomers, indicated by a dotted oval.
Extended Data Fig. 10 Purification and dynamic light scattering analysis of LASV, MACV L and mutants.
a–d, SEC and SDS–PAGE profiles of LASV and MACV L mutants. All of these mutants display similar behaviour to the WT proteins, indicating predominant monomeric forms in solution. e, f, Dynamic light scattering patterns of the WT LASV and MACV L and mutants. Both mut_A and mut_B mutants showed sharper and smaller radius distribution patterns than the WT proteins, indicating attenuated dimeric interactions and thus better homogeneity than the WT. The upper limit of molecular radius for mut_B proteins is indicated by red dashed lines to facilitate comparison. The data shown in this figure are representative of three independent experiments using different protein preparations.
Supplementary information
Supplementary Information
This file contains supplementary methods and discussions about RNA product characterization and different conformations of the structures, and Supplementary Fig. 1 presenting uncropped autoradiographs used for preparing Figs. 2 and 4, and Extended Data Fig. 2.
Rights and permissions
About this article
Cite this article
Peng, R., Xu, X., Jing, J. et al. Structural insight into arenavirus replication machinery. Nature 579, 615–619 (2020). https://doi.org/10.1038/s41586-020-2114-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2114-2
This article is cited by
-
Structural characterization of the oligomerization of full-length Hantaan virus polymerase into symmetric dimers and hexamers
Nature Communications (2024)
-
An intermediate state allows influenza polymerase to switch smoothly between transcription and replication cycles
Nature Structural & Molecular Biology (2023)
-
Structures of active Hantaan virus polymerase uncover the mechanisms of Hantaviridae genome replication
Nature Communications (2023)
-
Molecular mechanism of de novo replication by the Ebola virus polymerase
Nature (2023)
-
Structural snapshots of La Crosse virus polymerase reveal the mechanisms underlying Peribunyaviridae replication and transcription
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.