Abstract
Metazoan development requires the robust proliferation of progenitor cells, the identities of which are established by tightly controlled transcriptional networks1. As gene expression is globally inhibited during mitosis, the transcriptional programs that define cell identity must be restarted in each cell cycle2,3,4,5 but how this is accomplished is poorly understood. Here we identify a ubiquitin-dependent mechanism that integrates gene expression with cell division to preserve cell identity. We found that WDR5 and TBP, which bind active interphase promoters6,7, recruit the anaphase-promoting complex (APC/C) to specific transcription start sites during mitosis. This allows APC/C to decorate histones with ubiquitin chains branched at Lys11 and Lys48 (K11/K48-branched ubiquitin chains) that recruit p97 (also known as VCP) and the proteasome, which ensures the rapid expression of pluripotency genes in the next cell cycle. Mitotic exit and the re-initiation of transcription are thus controlled by a single regulator (APC/C), which provides a robust mechanism for maintaining cell identity throughout cell division.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All original data are available from the corresponding author on request. ChIP–seq and RNA-sequencing data have been deposited at the Gene Expression Omnibus, accession code GSE122298.
Code availability
Custom Python scripts are available from the corresponding author on request.
References
Young, R. A. Control of the embryonic stem cell state. Cell 144, 940–954 (2011).
Michelotti, E. F., Sanford, S. & Levens, D. Marking of active genes on mitotic chromosomes. Nature 388, 895–899 (1997).
Teves, S. S. et al. A stable mode of bookmarking by TBP recruits RNA polymerase II to mitotic chromosomes. eLife 7, e35621 (2018).
Palozola, K. C. et al. Mitotic transcription and waves of gene reactivation during mitotic exit. Science 358, 119–122 (2017).
Hsiung, C. C. et al. A hyperactive transcriptional state marks genome reactivation at the mitosis–G1 transition. Genes Dev. 30, 1423–1439 (2016).
Thomas, L. R. et al. Interaction with WDR5 promotes target gene recognition and tumorigenesis by MYC. Mol. Cell 58, 440–452 (2015).
Wysocka, J. et al. WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell 121, 859–872 (2005).
Keyes, B. E. & Fuchs, E. Stem cells: aging and transcriptional fingerprints. J. Cell Biol. 217, 79–92 (2018).
Prescott, D. M. & Bender, M. A. Synthesis of RNA and protein during mitosis in mammalian tissue culture cells. Exp. Cell Res. 26, 260–268 (1962).
Martínez-Balbás, M. A., Dey, A., Rabindran, S. K., Ozato, K. & Wu, C. Displacement of sequence-specific transcription factors from mitotic chromatin. Cell 83, 29–38 (1995).
Caravaca, J. M. et al. Bookmarking by specific and nonspecific binding of FoxA1 pioneer factor to mitotic chromosomes. Genes Dev. 27, 251–260 (2013).
Festuccia, N. et al. Mitotic binding of Esrrb marks key regulatory regions of the pluripotency network. Nat. Cell Biol. 18, 1139–1148 (2016).
Kadauke, S. et al. Tissue-specific mitotic bookmarking by hematopoietic transcription factor GATA1. Cell 150, 725–737 (2012).
Rape, M. Ubiquitylation at the crossroads of development and disease. Nat. Rev. Mol. Cell Biol. 19, 59–70 (2018).
Buckley, S. M. et al. Regulation of pluripotency and cellular reprogramming by the ubiquitin-proteasome system. Cell Stem Cell 11, 783–798 (2012).
Gao, J. et al. The CUL4–DDB1 ubiquitin ligase complex controls adult and embryonic stem cell differentiation and homeostasis. eLife 4, e07539 (2015).
Hu, G. et al. A genome-wide RNAi screen identifies a new transcriptional module required for self-renewal. Genes Dev. 23, 837–848 (2009).
Yau, R. G. et al. Assembly and function of heterotypic ubiquitin chains in cell-cycle and protein quality control. Cell 171, 918–933 (2017).
Stegmeier, F. et al. Anaphase initiation is regulated by antagonistic ubiquitination and deubiquitination activities. Nature 446, 876–881 (2007).
Ang, Y. S. et al. Wdr5 mediates self-renewal and reprogramming via the embryonic stem cell core transcriptional network. Cell 145, 183–197 (2011).
Pijnappel, W. W. et al. A central role for TFIID in the pluripotent transcription circuitry. Nature 495, 516–519 (2013).
Karatas, H. et al. High-affinity, small-molecule peptidomimetic inhibitors of MLL1/WDR5 protein–protein interaction. J. Am. Chem. Soc. 135, 669–682 (2013).
Mark, K. G., Loveless, T. B. & Toczyski, D. P. Isolation of ubiquitinated substrates by tandem affinity purification of E3 ligase-polyubiquitin-binding domain fusions (ligase traps). Nat. Protoc. 11, 291–301 (2016).
Fuchs, G. et al. RNF20 and USP44 regulate stem cell differentiation by modulating H2B monoubiquitylation. Mol. Cell 46, 662–673 (2012).
Chang, L. F., Zhang, Z., Yang, J., McLaughlin, S. H. & Barford, D. Molecular architecture and mechanism of the anaphase-promoting complex. Nature 513, 388–393 (2014).
Meyer, H. J. & Rape, M. Enhanced protein degradation by branched ubiquitin chains. Cell 157, 910–921 (2014).
Matsumoto, M. L. et al. K11-linked polyubiquitination in cell cycle control revealed by a K11 linkage-specific antibody. Mol. Cell 39, 477–484 (2010).
Aho, E. R. et al. Displacement of WDR5 from chromatin by a WIN site inhibitor with picomolar affinity. Cell Rep. 26, 2916–2928 (2019).
King, R. W. et al. A 20S complex containing CDC27 and CDC16 catalyzes the mitosis-specific conjugation of ubiquitin to cyclin B. Cell 81, 279–288 (1995).
Blobel, G. A. et al. A reconfigured pattern of MLL occupancy within mitotic chromatin promotes rapid transcriptional reactivation following mitotic exit. Mol. Cell 36, 970–983 (2009).
Pilaz, L. J. et al. Prolonged mitosis of neural progenitors alters cell fate in the developing Brain. Neuron 89, 83–99 (2016).
Halley-Stott, R. P., Jullien, J., Pasque, V. & Gurdon, J. Mitosis gives a brief window of opportunity for a change in gene transcription. PLoS Biol. 12, e1001914 (2014).
Egli, D., Birkhoff, G. & Eggan, K. Mediators of reprogramming: transcription factors and transitions through mitosis. Nat. Rev. Mol. Cell Biol. 9, 505–516 (2008).
Hockemeyer, D. et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat. Biotechnol. 29, 731–734 (2011).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Kampmann, M., Bassik, M. C. & Weissman, J. S. Functional genomics platform for pooled screening and generation of mammalian genetic interaction maps. Nat. Protoc. 9, 1825–1847 (2014).
McGourty, C. A. et al. Regulation of the CUL3 ubiquitin ligase by a calcium-dependent co-adaptor. Cell 167, 525–538 (2016).
Qiao, R. et al. Mechanism of APC/CCDC20 activation by mitotic phosphorylation. Proc. Natl Acad. Sci. USA 113, E2570–E2578 (2016).
Brown, N. G. et al. RING E3 mechanism for ubiquitin ligation to a disordered substrate visualized for human anaphase-promoting complex. Proc. Natl Acad. Sci. USA 112, 5272–5279 (2015).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Zhang, S. et al. Molecular mechanism of APC/C activation by mitotic phosphorylation. Nature 533, 260–264 (2016).
Zhang, P., Lee, H., Brunzelle, J. S. & Couture, J. F. The plasticity of WDR5 peptide-binding cleft enables the binding of the SET1 family of histone methyltransferases. Nucleic Acids Res. 40, 4237–4246 (2012).
Brown, N. G. et al. Mechanism of polyubiquitination by human anaphase-promoting complex: RING repurposing for ubiquitin chain assembly. Mol. Cell 56, 246–260 (2014).
Tsankov, A. M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015).
Acknowledgements
We thank N. Ingolia, R. Tjian, B. Schulman, J. Schaletzky and all members of M.R.’s laboratory for advice, helpful discussions and comments on the manuscript; and M. Matsumoto and V. Dixit for generously supplying us with linkage-specific ubiquitin antibodies. E.O. was funded by the Jane Coffin Childs Memorial Fund for Medical Research and the Siebel Stem Cell Institute. K.G.M. was funded by the NIH F32 postdoctoral fellowship (F32GM120956). A.M. was funded by the American Italian Cancer foundation and the California Institute for Regenerative Medicine. M.R. is an Investigator of the Howard Hughes Medical Institute. This work was also funded by an NIH grant (RO1GM083064) awarded to M.R.
Author information
Authors and Affiliations
Contributions
E.O. performed work with stem cells (including knockdowns and infections), and performed and analysed the ultracomplex screen, cytometry-based studies, microscopy-based studies, qPCR-based studies and ChIP–seq and RNA-sequencing experiments. K.G.M. performed in vitro assays. E.O., K.G.M., A.M. and D.D.C. prepared samples for mass spectrometry analyses, performed immunoprecipitation experiments and maintained cell culture. E.R.W. and J.R.P. performed the cryo-EM of APC/C–WDR5. M.K. helped to analyse the ultracomplex screen. N.G. and C.Y.Z. prepared recombinant histones for in vitro assays. E.O., K.G.M. and M.R. interpreted the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.R. is a cofounder and consultant to Nurix Therapeutics, a biotechnology company working in the ubiquitin space.
Additional information
Peer review information Nature thanks William P. Tansey and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Ultracomplex shRNA screen identifies APC/C and USP44 as regulators of human ES cell biology.
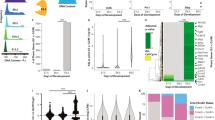
a, Karyotype analysis of H1 OCT4–GFP cell line shows normal chromosome architecture. This line was karyotyped before performing the screen by a third party vendor (WiCell). Twenty cells were counted, 8 were analysed and 4 were karyotyped as normal. No clonal abnormalities were detected at the band resolution of 450–475. b, H1 OCT4–GFP cells undergo neural conversion with an efficiency similar to that of the unmodified parent line. This experiment was performed three independent times with similar results. c, Deep sequencing read counts (log2-transformed) for individual shRNAs (red dots) targeting the indicated gene from the screen in Fig. 1b. Grey dots represent negative-control shRNAs. P values (two-sided Mann–Whitney U test, not corrected for multiple hypothesis testing) are indicated for each gene. d, Mass spectrometry analysis shows that many quality-control enzymes associate with K11/K48-branched chains in human ES cells. H1 human ES cells were synchronized in mitosis before being subjected to affinity purification using K11/K48-bispecific antibodies under denaturing conditions. Values listed in brackets are total spectral counts of tryptic peptides for each protein. e, CRISPR–Cas9-edited USP44 H1 human ES cells show impaired rates of neural conversion. Expression of markers of pluripotency (OCT4), neural crest cells (SNAIL2) or neural progenitors (PAX6) were determined at indicated times of differentiation by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) and western blotting using specific antibodies. This experiment was performed two independent times with similar results.
Extended Data Fig. 2 Characterization of APC/C and the role of USP44 in pluripotency.
a, Western blot of OCT4 and NANOG upon APC2 knockdown in asynchronous H1 human ES cells. This experiment was performed two independent times with similar results. b, Western blot of OCT4 and NANOG upon knockdown of APC/C subunits in asynchronous H1 human ES cells. This experiment was performed three independent times with similar results. c, Real-time qPCR of OCT4 and NANOG upon APC2 knockdown in asynchronous H1 human ES cells (mean of n = 4 independent experiments, ± s.d.). d, Flow cytometry analysis of APC2 depletion in H1 OCT4–GFP human ES cells. H1 OCT4–GFP human ES cells were transfected with siRNA against APC2 for 48 h before cytometry analysis. This experiment was performed two independent times with similar results. e, Loss of the pluripotency marker OCT4 upon depletion of APC2 or WDR5 requires entry into mitosis. H1 human ES cells were transfected with indicated siRNAs for 36 h and treated with DMSO (asynchronous), 5 μM STLC (mitotic arrest) or 200 mM thymidine (S-phase arrest) for an additional 12 h before collection for western blot analysis. This experiment was performed three independent times with similar results. f, Flow cytometry analysis of asynchronous H1 human ES cells transfected with indicated siRNAs for 72 h. This experiment was performed three independent times with similar results.
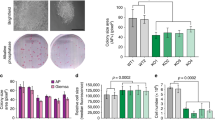
Extended Data Fig. 3 APC/C and WDR5 are required for human ES cell survival.
a, Mass spectrometry analysis of Flag–WDR5 purified from mitotic HEK293T cells. Values listed in brackets are total spectral counts of tryptic peptides of indicated proteins. b, Depletion of WDR5 phenocopies the depletion of APC2 in H1 human ES cells. H1 cells were depleted with the indicated siRNAs for 72 h before collection for western blot analysis. This experiment was performed once. c, A cumulative fraction curve measuring the length of each metaphase-to-anaphase transition. n = 112 cells for control siRNA; n = 105 cells for siRNA against APC2; n = 106 cells for siRNA against WDR5; and n = 217 cells for siRNAs against both APC2 and WDR5. d, Depletion of APC2 or WDR5 causes cell death in H1 human ES cells. Cell death was measured by trypan blue staining of dead cells (mean of n = 4 independent experiments ± s.d.). e, Quantifying cell survival using chromosome catastrophe as a proxy for cell death. H1 human ES cells virally expressing H2B–mCherry were transfected with the indicated siRNAs for 24 h before imaging by confocal microscopy. n = 97 cells for control siRNA; n = 104 cells for siRNA against APC2; n = 90 cells for siRNA against WDR5; and n = 213 cells for siRNAs against both APC2 and WDR5. f, Sister cells die immediately following mitotic exit when depleted of APC2 and WDR5. H1 human ES cells virally expressing H2B–mCherry were transfected with siRNA against APC2 and/or siRNA against WDR5 for 24 h before imaging by confocal microscopy. The time of death, as defined by cells undergoing chromosome catastrophe, was measured for each sister (mean of n = 57 pairs of cells ± s.d.). g, Representative frames of live-cell imaging from four independent experiments (in minutes) tracking the nuclei of siRNA-depleted H1 human ES cells virally expressing H2B–mCherry. Arrows mark individual sister cells upon mitotic exit. Chromosome catastrophe was used a proxy for cell death (time points 198 and 342). h, Flag–WDR5 associates with APC/C in mitotic H1 human ES cells. Flag–WDR5 immunoprecipitations were performed on asynchronous H1 human ES cells (A) or H1 human ES cells arrested in mitosis (M). Bound proteins were determined by SDS–PAGE and western blotting. This experiment was performed two independent times with similar results. i, Overexpressed haemagglutinin (HA)-tagged USP44 associates with Flag–WDR5 in both asynchronous and mitotic HEK293T cells. MYC–WDR5 was used as the control vector. This experiment was performed three independent times with similar results.
Extended Data Fig. 4 WDR5 associates with APC/C and TBP on distinct surfaces.
a, The WIN-motif binding site on WDR5 is critical for APC/C engagement, whereas the surface that binds the WBM is dispensable. A secondary binding surface (4A) is also important for the association of WDR5 with APC/C. The surface on WDR5 that binds the WBM is critical for TFIID association, whereas the WIN-motif binding site is dispensable. HEK293T cells were transfected with the indicated Flag–WDR5 variants, and cells were synchronized in mitosis. Flag–WDR5 was affinity-purified, and bound proteins were determined by western blotting. This experiment was performed five independent times with similar results. b, Reciprocal immunoprecipitations show that APC/C binds WDR5 through its WIN-motif binding site. Endogenous APC/C was purified from HEK293T cells expressing the indicated Flag–WDR5 variants, and bound proteins were determined by SDS–PAGE and western blotting. This experiment was performed three independent times with similar results. c, Heat map of bait-normalized total spectral counts identified from Flag–WDR5-purified mass spectrometry experiments. HeLa cells were transfected with Flag–WDR5 for 24 h before mitotic synchronization. d, The WDR5 inhibitor MM-102 impairs the association of WDR5with APC/C. Mitotic HeLa S3 cells were released into MM-102 for 2 h before immunoprecipitation experiments. Under these conditions, MM-102 did not prevent the association of WDR5 with MLL and RBBP5. This experiment was performed two independent times with similar results. e, Expression of wild-type WDR5 but not WDR5(∆WIN) rescues the pluripotency defect caused by WDR5 depletion in H1 human ES cells. H1 human ES cells virally expressing siRNA-resistant WDR5 variants (WDR5 versus WDR5(∆WIN)) were depleted of endogenous WDR5 (W) or treated with control siRNA (C). Expression of OCT4 and NANOG was determined by western blotting. This experiment was performed once.
Extended Data Fig. 5 WDR5 binds near the catalytic core of APC/C.
a, WDR5 forms BMB-dependent crosslinks with APC2, APC11 and CDC20. APC/C–CDC20 was affinity-purified from prometaphase-arrested HeLa cells and incubated with recombinant WDR5 or WDR5(ΔWIN) before the addition of crosslinker. Crosslinked APC/C subunits were detected by SDS–PAGE and western blotting using specific antibodies. This experiment was performed two independent times with similar results. b, In vitro translation binding assays reveal that APC2 directly interacts with recombinant WDR5. MBP-tagged WDR5 or WDR5(ΔWIN) were immobilized on amylose resin and incubated with 35S-labelled APC/C subunits produced by in vitro transcription and translation. Bound proteins were detected by SDS–PAGE and autoradiography. APC3 did not synthesize by in vitro transcription and translation. This experiment was performed once for the full set of APC/C subunits. APC2 binding was validated three independent times. c, Quantification of autoradiography blot shown in b.
Extended Data Fig. 6 WDR5 associates with active APC/C.
a, APC/C does not ubiquitylate WDR5 in vitro. Recombinant WDR5 was incubated with active APC/C, E1, UBE2C, UBE2S and ubiquitin, and potential reaction products were detected by western blotting against WDR5. This experiment was performed once. b, APC/C-dependent ubiquitylation of geminin is outcompeted by recombinant securin (comp), a canonical substrate, but not by recombinant WDR5. Securin or WDR5 was added to APC/C-dependent geminin ubiquitylation reactions at the indicated concentrations, and various reaction products were detected using western blotting. Asterisks represent cross-reactive bands. This experiment was performed once. c, APC/C–WDR5-dependent ubiquitylation of geminin is inhibited by EMI1. WDR5 affinity purifications from mitotic HeLa cells were incubated with E1, the APC/C-specific E2 enzymes UBE2C and UBE2S, and ubiquitin. EMI1 was added at indicated concentrations, and reaction products were detected by western blotting using antibodies against geminin. This experiment was performed two independent times with similar results. d, Immunoprecipitation of Flag–WDR5 from mitotic HEK293T cells coprecipitates K11-linked ubiquitin chains. HEK293T cells arrested in prometaphase were released into fresh medium, and WDR5 was affinity-purified at the indicated time points. Bound proteins were detected by western blotting. This experiment was performed once. e, Depletion of UBE2S eliminates WDR5-associated K11-linked ubiquitin chains in mitotic HEK293T cells. This experiment was performed once.
Extended Data Fig. 7 Mitotic APC/C–WDR5 complexes are catalytically active.
a, APC/C–WDR5 ubiquitylates human H3 in H3–H4 tetramers in vitro. APC/C was affinity-purified from mitotic HeLa cells and incubated with E1, UBE2C, UBE2S, ubiquitin and human H3–H4 tetramers, as indicated. Reaction products were detected by western blotting using antibodies against H3 and H4. This experiment was performed once. b, APC/C ubiquitylation of H2B in X. laevis H2A–H2B histone dimers. APC/C–WDR5 was purified from mitotic HeLa cells by Flag–WDR5 affinity purification and incubated with E1, UBE2C, UBE2S, ubiquitin and X. laevis histone octamers. Ubiquitylation was detected by western blotting against ubiquitylated H2B. This experiment was performed three independent times with similar results. c, APC/C–WDR5 ubiquitylation of H2B in X. laevis H2A–H2B–H3–H4 histone octamers. Reactions were performed as described in b. This experiment was performed two independent times with similar results. d, APC/C purified from H1 human ES cells is competent to ubiquitylate human H2B. This experiment was performed two independent times with similar results. e, APC/C purified from mitotic, but not S-phase, extracts can ubiquitylate H2B in vitro. APC/C was purified from HeLa cells synchronized at the indicated cell-cycle stages and incubated with E1, UBE2C, UBE2S, ubiquitin and X. laevis H2A–H2B dimers. Histone ubiquitylation was detected by western blotting using antibodies against ubiquitylated H2B. This experiment was performed once. f, APC/C-dependent ubiquitylation of H2B requires K11 residue on ubiquitin for chain elongation. Ubiquitylation of H2A–H2B dimers by APC/C–WDR5 was performed as described in e, but with ubiquitin variants. This experiment was performed once. g, APC/C-dependent ubiquitylation of H2B requires both K11 and K48 on ubiquitin for synthesis of branched chains. This experiment was performed two independent times with similar results. h, Securin, a canonical APC/C substrate, outcompetes H2A–H2B dimers for APC/C-dependent ubiquitylation. The D-box motif (an APC/C–CDC20-specific degron) is required for full competition, whereas the KEN motif (an APC/C–CDH1-specific degron) is not. This experiment was performed two independent times with similar results. i, Polyubiquitylated H2B is degraded by the proteasome. K11/K48-branched chains were purified under denaturing conditions from mitotic HeLa cells either in the presence or absence of MG132, and modified H2B was detected using western blotting. Proteasome inhibition with MG132 was found to stabilize endogenous polyubiquitylated H2B. This experiment was performed four independent times with similar results.
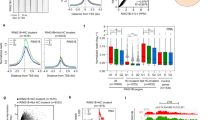
Extended Data Fig. 8 ChIP–seq analyses of APC/C- and WDR5-occupied gene targets.
a, Overall histone levels do not change upon release from mitosis. H1 cells were synchronized in mitosis by STLC and released into fresh medium. Indicated proteins were monitored by western blotting. This experiment was performed two independent times with similar results. b, Comparison between ChIP–seq and MNase ChIP–seq against anti-K11 from mitotic H1 human ES cells reveals that sonication shears polymeric ubiquitin linkages. c, Venn diagram of anti-K11 and anti-WDR5 ChIP peaks that colocalize with TSSs from MNase ChIP–seq experiments. MNase ChIP–seq experiments were performed from mitotic H1 human ES cells. d, Heat map of MNase ChIP–seq data from mitotic H1 human ES cells. Cluster 1 includes sites that are co-occupied by K11 and WDR5 near TSSs (within 100 bp); cluster 2 includes sites that are co-occupied by K11 and WDR5 outside of TSSs; and cluster 3 includes sites occupied only by K11, regardless of colocalization with TSSs. e, ChIP–qPCR analysis of candidate targets using K11- or K11/K48-linkage specific ubiquitin antibodies from mitotic H1 human ES cells. Mean of independent replicates ± s.d. n = 3 for K11/K48; n = 5 for IGG and K11 (except n = 4 for PUM1). f, ChIP–qPCR analysis of mitotic H1 human ES cells shows that K11 linkages synthesized at candidate sites are dependent on UBE2S and WDR5. This experiment was performed once. g, WDR5 inhibition prevents K11-ubiquitin chain formation at APC/C–WDR5-bound TSSs. H1 human ES cells were treated with or without 50 μM MM102 during mitotic synchronization with STLC before anti-K11 MNase ChIP–seq. Heat map of all APC/C–WDR5-bound TSSs are shown. h, Heat map of ChIP–seq peaks of individual genes co-occupied by Flag–CDC20 and Flag–WDR5. ChIP–seq against anti-Flag was performed on mitotic HEK293T cells that overexpress Flag–CDC20 or Flag–WDR5. i, Spatial profile of PUM1 of factor occupancy by ChIP–qPCR. This experiment was performed once. j, Spatial profile of E2F3 of factor occupancy by ChIP–qPCR. This experiment was performed once. k, Heat map of MNase ChIP–seq data of transcription-factor binding. Previously published MNase ChIP–seq data were obtained47, and APC/C-bound sites were analysed as described in d.
Extended Data Fig. 9 Select transcription factors are found at APC/C–WDR5-bound sites.
Heat map of MNase ChIP–seq data of transcription-factor binding. Previously published MNase ChIP–seq data were obtained47, and APC/C-bound sites were analysed as follows: cluster 1 includes sites that are co-occupied by K11 and WDR5 near TSSs (within 100 bp); cluster 2 includes sites that are co-occupied by K11 and WDR5 outside of TSSs; and cluster 3 includes sites only occupied by K11, regardless of colocalization with TSSs.
Extended Data Fig. 10 Regulation of chromatin and transcription by APC/C–WDR5.
a, Comparison of genes co-occupied by Flag–CDC20 and Flag–WDR5 from mitotic HEK293T cells with known gene-expression profiles reveals a strong overlap with ES cell and medulloblastoma cancer cell lines. n = 1,628 genes were analysed (P values represent a one-sided Fisher’s exact test with Bonferroni correction). b, Loss of APC/C–WDR5 function interferes with the expression of genes marked with K11-linked ubiquitin chains in H1 human ES cells. Poly(A)-selected RNA was purified from asynchronous H1 human ES cells transfected with control siRNA or siRNA against WDR5 for 48 h, and subjected to RNA-sequencing analysis (a biological replicate of Fig. 4h). c, Transcript analysis of WDR5 depletion on APC/C–WDR5-dependent genes (from Fig. 4h and b). Box plots include the median TPM value (n = 90 genes) with quartile ranges Q1–Q3; top whiskers represent the 3rd quartile + 1.5× interquartile range; bottom whiskers represent the 1st quartile − 1.5× interquartile range. P values were calculated from comparing individual TPM values of APC/C–WDR5-regulated genes (n = 90) versus all transcripts (n = 18,791) using a two-sided Student’s t-test (unpaired). d, Real-time qPCR analysis of nascent RNA reveals APC/C–WDR5 target genes are reactivated upon mitotic exit and gene reactivation is dependent on WDR5. Initial screening from a single experiment. e, The RNA levels of genes regulated by APC/C–WDR5 do not change upon mitotic exit. RNA-sequencing analysis was performed on poly(A)-selected RNA purified from H1 human ES cells at the indicated cell-cycle stages. Box plots were derived as described in c (n = 90 genes). f, Ubiquitylated H2B preferentially associates with p97–UBXN7 in vitro. H2B was preubiquitylated by APC/C in vitro, and incubated with immobilized p97 or p97–UBXN7 complexes. Bound histone H2B was detected by western blotting. This experiment was performed three independent times with similar results. g, Flag–UBXN7 associates with polyubiquitylated H2B, p97 and K11/K48-linked branched ubiquitin chains in mitosis. Native Flag–UBXN7 immunoprecipitations were performed on mitotic HEK293T cells and bound proteins were detected by western blotting or Ponceau staining. This experiment was performed three independent times with similar results. h, H2B ubiquitylation is stabilized by p97 inhibition in cells. Denaturing K11/K48 immunoprecipitations were performed on H1 human ES cells synchronized in prometaphase or released into 10 μM NMS-873 for 2 h. This experiment was performed four independent times with similar results. i, p97 inhibition restores K11 deposition at sites regulated by APC/C–WDR5 upon mitotic exit. Anti-K11 MNase ChIP–seq was performed from H1 human ES cells synchronized in mitosis (0 h) or released into fresh medium without (2 h + DMSO) or with p97 inhibition (2 h + 10 μM NMS-873). j, Anti-K11 MNase ChIP–qPCR of candidate targets from mitotic H1 human ES cells (mean of n = 3 independent replicates ± s.d.). H1 human ES cells were synchronized in mitosis (0 h) and released into fresh medium for 2 h with the indicated drugs. k, Model of APC/C-dependent gene activation upon mitotic exit.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-2 showing the uncropped gel data and Supplementary Tables 1-3, showing lists of plasmids and antibodies used in the study as well as oligonucleotide sequences used for siRNA knockdown, ChIP-qPCR and RT-qPCR.
Rights and permissions
About this article
Cite this article
Oh, E., Mark, K.G., Mocciaro, A. et al. Gene expression and cell identity controlled by anaphase-promoting complex. Nature 579, 136–140 (2020). https://doi.org/10.1038/s41586-020-2034-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2034-1
This article is cited by
-
Mitotic gene regulation by the N-MYC-WDR5-PDPK1 nexus
BMC Genomics (2024)
-
Mitotic bookmarking redundancy by nuclear receptors in pluripotent cells
Nature Structural & Molecular Biology (2024)
-
Structural snapshots along K48-linked ubiquitin chain formation by the HECT E3 UBR5
Nature Chemical Biology (2024)
-
ASPSCR1-TFE3 reprograms transcription by organizing enhancer loops around hexameric VCP/p97
Nature Communications (2024)
-
Exploiting E3 ubiquitin ligases to reeducate the tumor microenvironment for cancer therapy
Experimental Hematology & Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.