Abstract
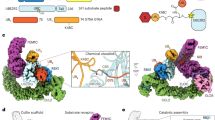
Eukaryotic cell biology depends on cullin–RING E3 ligase (CRL)-catalysed protein ubiquitylation1, which is tightly controlled by the modification of cullin with the ubiquitin-like protein NEDD82,3,4,5,6. However, how CRLs catalyse ubiquitylation, and the basis of NEDD8 activation, remain unknown. Here we report the cryo-electron microscopy structure of a chemically trapped complex that represents the ubiquitylation intermediate, in which the neddylated CRL1β-TRCP promotes the transfer of ubiquitin from the E2 ubiquitin-conjugating enzyme UBE2D to its recruited substrate, phosphorylated IκBα. NEDD8 acts as a nexus that binds disparate cullin elements and the RING-activated ubiquitin-linked UBE2D. Local structural remodelling of NEDD8 and large-scale movements of CRL domains converge to juxtapose the substrate and the ubiquitylation active site. These findings explain how a distinctive ubiquitin-like protein alters the functions of its targets, and show how numerous NEDD8-dependent interprotein interactions and conformational changes synergistically configure a catalytic CRL architecture that is both robust, to enable rapid ubiquitylation of the substrate, and fragile, to enable the subsequent functions of cullin–RING proteins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The atomic coordinates and electron microscopy maps have been deposited in the PDB with accession code 6TTU and the Electron Microscopy Data Bank with codes EMD-10585, EMD-10578, EMD-10579, EMD-10580, EMD-10581, EMD-10582 and EMD-10583. Uncropped gel source data are included as Supplementary Information. All other reagents and data (for example, raw gels of replicate experiments and raw movie electron microscopy data) are available from the corresponding author upon request.
References
Lydeard, J. R., Schulman, B. A. & Harper, J. W. Building and remodelling Cullin–RING E3 ubiquitin ligases. EMBO Rep. 14, 1050–1061 (2013).
Read, M. A. et al. Nedd8 modification of Cul-1 activates SCFβTrCP-dependent ubiquitination of IκBα. Mol. Cell. Biol. 20, 2326–2333 (2000).
Duda, D. M. et al. Structural insights into NEDD8 activation of cullin–RING ligases: conformational control of conjugation. Cell 134, 995–1006 (2008).
Saha, A. & Deshaies, R. J. Multimodal activation of the ubiquitin ligase SCF by Nedd8 conjugation. Mol. Cell 32, 21–31 (2008).
Yamoah, K. et al. Autoinhibitory regulation of SCF-mediated ubiquitination by human cullin 1’s C-terminal tail. Proc. Natl Acad. Sci. USA 105, 12230–12235 (2008).
Soucy, T. A. et al. An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature 458, 732–736 (2009).
Zheng, N. et al. Structure of the Cul1–Rbx1–Skp1–F boxSkp2 SCF ubiquitin ligase complex. Nature 416, 703–709 (2002).
Jin, J. et al. Systematic analysis and nomenclature of mammalian F-box proteins. Genes Dev. 18, 2573–2580 (2004).
Willems, A. R., Schwab, M. & Tyers, M. A hitchhiker’s guide to the cullin ubiquitin ligases: SCF and its kin. Biochim. Biophys. Acta 1695, 133–170 (2004).
Angers, S. et al. Molecular architecture and assembly of the DDB1–CUL4A ubiquitin ligase machinery. Nature 443, 590–593 (2006).
Jin, J., Arias, E. E., Chen, J., Harper, J. W. & Walter, J. C. A family of diverse Cul4-Ddb1-interacting proteins includes Cdt2, which is required for S phase destruction of the replication factor Cdt1. Mol. Cell 23, 709–721 (2006).
Yu, C. et al. Gln40 deamidation blocks structural reconfiguration and activation of SCF ubiquitin ligase complex by Nedd8. Nat. Commun. 6, 10053 (2015).
Pierce, N. W. et al. Cand1 promotes assembly of new SCF complexes through dynamic exchange of F box proteins. Cell 153, 206–215 (2013).
Stanley, D. J. et al. Inhibition of a NEDD8 cascade restores restriction of HIV by APOBEC3G. PLoS Pathog. 8, e1003085 (2012).
Yaron, A. et al. Identification of the receptor component of the IκBα-ubiquitin ligase. Nature 396, 590–594 (1998).
Winston, J. T. et al. The SCFβ-TRCP-ubiquitin ligase complex associates specifically with phosphorylated destruction motifs in IκBα and β-catenin and stimulates IκBα ubiquitination in vitro. Genes Dev. 13, 270–283 (1999).
Spencer, E., Jiang, J. & Chen, Z. J. Signal-induced ubiquitination of IκBα by the F-box protein Slimb/β-TrCP. Genes Dev. 13, 284–294 (1999).
Hart, M. et al. The F-box protein β-TrCP associates with phosphorylated β-catenin and regulates its activity in the cell. Curr. Biol. 9, 207–211 (1999).
Latres, E., Chiaur, D. S. & Pagano, M. The human F box protein β-Trcp associates with the Cul1/Skp1 complex and regulates the stability of β-catenin. Oncogene 18, 849–854 (1999).
Wu, G. et al. Structure of a β-TrCP1–Skp1–β-catenin complex: destruction motif binding and lysine specificity of the SCFβ-TrCP1 ubiquitin ligase. Mol. Cell 11, 1445–1456 (2003).
Wu, K., Kovacev, J. & Pan, Z. Q. Priming and extending: a UbcH5/Cdc34 E2 handoff mechanism for polyubiquitination on a SCF substrate. Mol. Cell 37, 784–796 (2010).
Frescas, D. & Pagano, M. Deregulated proteolysis by the F-box proteins SKP2 and β-TrCP: tipping the scales of cancer. Nat. Rev. Cancer 8, 438–449 (2008).
Margottin, F. et al. A novel human WD protein, h-βTrCp, that interacts with HIV-1 Vpu connects CD4 to the ER degradation pathway through an F-box motif. Mol. Cell 1, 565–574 (1998).
Cui, J. et al. Glutamine deamidation and dysfunction of ubiquitin/NEDD8 induced by a bacterial effector family. Science 329, 1215–1218 (2010).
Jubelin, G. et al. Pathogenic bacteria target NEDD8-conjugated cullins to hijack host-cell signaling pathways. PLoS Pathog. 6, e1001128 (2010).
Morikawa, H. et al. The bacterial effector Cif interferes with SCF ubiquitin ligase function by inhibiting deneddylation of Cullin1. Biochem. Biophys. Res. Commun. 401, 268–274 (2010).
Tang, X. et al. Suprafacial orientation of the SCFCdc4 dimer accommodates multiple geometries for substrate ubiquitination. Cell 129, 1165–1176 (2007).
Dou, H., Buetow, L., Sibbet, G. J., Cameron, K. & Huang, D. T. BIRC7–E2 ubiquitin conjugate structure reveals the mechanism of ubiquitin transfer by a RING dimer. Nat. Struct. Mol. Biol. 19, 876–883 (2012).
Plechanovová, A., Jaffray, E. G., Tatham, M. H., Naismith, J. H. & Hay, R. T. Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis. Nature 489, 115–120 (2012).
Pruneda, J. N. et al. Structure of an E3:E2~Ub complex reveals an allosteric mechanism shared among RING/U-box ligases. Mol. Cell 47, 933–942 (2012).
Brzovic, P. S. & Klevit, R. E. Ubiquitin transfer from the E2 perspective: why is UbcH5 so promiscuous? Cell Cycle 5, 2867–2873 (2006).
Low, T. Y. et al. A systems-wide screen identifies substrates of the SCFβTrCP ubiquitin ligase. Sci. Signal. 7, rs8 (2014).
Lumb, K. J. & Kim, P. S. A buried polar interaction imparts structural uniqueness in a designed heterodimeric coiled coil. Biochemistry 34, 8642–8648 (1995).
Kawakami, T. et al. NEDD8 recruits E2-ubiquitin to SCF E3 ligase. EMBO J. 20, 4003–4012 (2001).
Ozkan, E., Yu, H. & Deisenhofer, J. Mechanistic insight into the allosteric activation of a ubiquitin-conjugating enzyme by RING-type ubiquitin ligases. Proc. Natl Acad. Sci. USA 102, 18890–18895 (2005).
Brzovic, P. S., Lissounov, A., Christensen, D. E., Hoyt, D. W. & Klevit, R. E. A. A UbcH5/ubiquitin noncovalent complex is required for processive BRCA1-directed ubiquitination. Mol. Cell 21, 873–880 (2006).
Sakata, E. et al. Direct interactions between NEDD8 and ubiquitin E2 conjugating enzymes upregulate cullin-based E3 ligase activity. Nat. Struct. Mol. Biol. 14, 167–168 (2007).
Buetow, L. et al. Activation of a primed RING E3-E2–ubiquitin complex by non-covalent ubiquitin. Mol. Cell 58, 297–310 (2015).
Scott, D. C. et al. Structure of a RING E3 trapped in action reveals ligation mechanism for the ubiquitin-like protein NEDD8. Cell 157, 1671–1684 (2014).
Hospenthal, M. K., Freund, S. M. & Komander, D. Assembly, analysis and architecture of atypical ubiquitin chains. Nat. Struct. Mol. Biol. 20, 555–565 (2013).
Scott, D. C. et al. Two distinct types of E3 ligases work in unison to regulate substrate ubiquitylation. Cell 166, 1198–1214.e24 (2016).
Huttenhain, R. et al. ARIH2 is a Vif-dependent regulator of CUL5-mediated APOBEC3G degradation in HIV infection. Cell Host Microbe 26, 86–99.e7 (2019).
Hill, S. et al. Robust cullin–RING ligase function is established by a multiplicity of poly-ubiquitylation pathways. eLife 8, e51163 (2019).
den Besten, W., Verma, R., Kleiger, G., Oania, R. S. & Deshaies, R. J. NEDD8 links cullin–RING ubiquitin ligase function to the p97 pathway. Nat. Struct. Mol. Biol. 19, 511–516 (2012).
Schapira, M., Calabrese, M. F., Bullock, A. N. & Crews, C. M. Targeted protein degradation: expanding the toolbox. Nat. Rev. Drug Discov. 18, 949–963 (2019).
Goldenberg, S. J. et al. Structure of the Cand1–Cul1–Roc1 complex reveals regulatory mechanisms for the assembly of the multisubunit cullin-dependent ubiquitin ligases. Cell 119, 517–528 (2004).
Mosadeghi, R. et al. Structural and kinetic analysis of the COP9-signalosome activation and the cullin–RING ubiquitin ligase deneddylation cycle. eLife 5, e12102 (2016).
Cavadini, S. et al. Cullin–RING ubiquitin E3 ligase regulation by the COP9 signalosome. Nature 531, 598–603 (2016).
Liu, X. et al. Cand1-mediated adaptive exchange mechanism enables variation in F-box protein expression. Mol. Cell 69, 773–786.e6 (2018).
Kamadurai, H. B. et al. Insights into ubiquitin transfer cascades from a structure of a UbcH5B~ubiquitin-HECTNEDD4L complex. Mol. Cell 36, 1095–1102 (2009).
Brown, N. G. et al. Mechanism of polyubiquitination by human anaphase-promoting complex: RING repurposing for ubiquitin chain assembly. Mol. Cell 56, 246–260 (2014).
Weissmann, F. et al. biGBac enables rapid gene assembly for the expression of large multisubunit protein complexes. Proc. Natl Acad. Sci. USA 113, E2564–E2569 (2016).
Hao, B., Oehlmann, S., Sowa, M. E., Harper, J. W. & Pavletich, N. P. Structure of a Fbw7–Skp1–cyclin E complex: multisite-phosphorylated substrate recognition by SCF ubiquitin ligases. Mol. Cell 26, 131–143 (2007).
Schulman, B. A. et al. Insights into SCF ubiquitin ligases from the structure of the Skp1–Skp2 complex. Nature 408, 381–386 (2000).
Walden, H. et al. The structure of the APPBP1–UBA3–NEDD8–ATP complex reveals the basis for selective ubiquitin-like protein activation by an E1. Mol. Cell 12, 1427–1437 (2003).
Whitby, F. G., Xia, G., Pickart, C. M. & Hill, C. P. Crystal structure of the human ubiquitin-like protein NEDD8 and interactions with ubiquitin pathway enzymes. J. Biol. Chem. 273, 34983–34991 (1998).
Koduri, V. et al. Peptidic degron for IMiD-induced degradation of heterologous proteins. Proc. Natl Acad. Sci. USA 116, 2539–2544 (2019).
Starita, L. M. et al. Activity-enhancing mutations in an E3 ubiquitin ligase identified by high-throughput mutagenesis. Proc. Natl Acad. Sci. USA 110, E1263–E1272 (2013).
Pierce, N. W., Kleiger, G., Shan, S. O. & Deshaies, R. J. Detection of sequential polyubiquitylation on a millisecond timescale. Nature 462, 615–619 (2009).
Sievers, Q. L. et al. Defining the human C2H2 zinc finger degrome targeted by thalidomide analogs through CRBN. Science 362 eaat0572 (2018).
Lu, G. et al. UBE2G1 governs the destruction of cereblon neomorphic substrates. eLife 7, e40958 (2018).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Streich, F. C. Jr & Lima, C. D. Capturing a substrate in an activated RING E3/E2–SUMO complex. Nature 536, 304–308 (2016).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Palovcak, E. et al. A simple and robust procedure for preparing graphene-oxide cryo-EM grids. J. Struct. Biol. 204, 80–84 (2018).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Hohn, M. et al. SPARX, a new environment for cryo-EM image processing. J. Struct. Biol. 157, 47–55 (2007).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D 74, 814–840 (2018).
Duda, D. M. et al. Structure of a glomulin–RBX1–CUL1 complex: inhibition of a RING E3 ligase through masking of its E2-binding surface. Mol. Cell 47, 371–382 (2012).
Yunus, A. A. & Lima, C. D. Lysine activation and functional analysis of E2-mediated conjugation in the SUMO pathway. Nat. Struct. Mol. Biol. 13, 491–499 (2006).
Acknowledgements
We thank D. Scott, J. Kellermann, J. Liwocha, S. Kostrhon, D. Horn-Ghetko, J. W. Harper, R. V. Farese Jr., H. Stark, D. Haselbach, S. Raunser, T. Raisch, S. Scheres, S. Uebel and S. Pettera for assistance, reagents and helpful discussions; and M. Strauss, D. Bollschweiler, T. Schäfer and the cryo-EM facility at the Max Planck Institute of Biochemistry. This study was supported by the Max Planck Gesellschaft and by a grant of the European Commission (ERC Advanced Investigator Grant Nedd8Activate) to B.A.S. S.H. and G.K. were supported by a grant from the National Institutes of Health (R15GM117555-02).
Author information
Authors and Affiliations
Contributions
K.B., D.T.K., M.K., L.-M.N. and S.v.G. generated protein complexes and assayed quality for suitability for cryo-EM. D.T.K. conceived of, designed and generated stable proxies for ubiquitylation intermediates. K.B., D.T.K., S.H. and G.K. performed ubiquitylation assays. K.B. and J.R.P. collected, processed and refined cryo-EM data and built and refined structure. K.B., D.T.K., G.K. and B.A.S. prepared the manuscript with input from other authors. B.A.S. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks David Barford, Ronald Hay and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Quantitative pre-steady-state enzyme kinetics of neddylated CRL1β-TRCP- and UBE2D-dependent ubiquitylation.
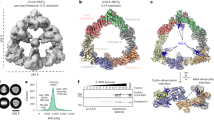
Gel images are representative of independent technical replicates (n = 2); the symbols on the graphs show the data from independent experiments (n = 2). a, Autoradiogram of SDS–PAGE gel showing products of ubiquitylation reactions under single-encounter conditions for the interaction of radiolabelled substrate (medium β-catenin substrate peptide derived from β-catenin) with neddylated CRL1β-TRCP, titrating UBE2D3 (hereafter denoted UBE2D). Each lane represents a single ubiquitylation reaction that was used to estimate the fraction of peptide that had been converted into ubiquitylated products as a function of UBE2D concentration. b, Plots of the fraction of substrate that had been converted to ubiquitylated products against UBE2D concentration for ubiquitylation reactions containing either wild-type UBE2D (as shown in a), or the mutants UBE2D(S22R) or UBE2D(H32A). Various CRL1β-TRCP complexes were assayed that contained either wild-type neddylated CRL1β-TRCP (red), CRL1β-TRCP complexes modified by NEDD8 variants containing I44A (orange), Q40E (green) or ‘ubiquitylizing’ (L2Q/K4F/E14T/D16E/G63K/G64E) substitutions (blue), or CRL1β-TRCP modified by Ub(R72A) that is competent for ligation to CUL1 (purple) or unmodified CRL1β-TRCP (CUL1 with the neddylation-site mutation K720R, black). Duplicate data points from independent experiments performed with identical samples are shown and were fit to the Michaelis–Menten model to estimate the Km of UBE2D for CRL1β-TRCP using nonlinear curve fitting (GraphPad Prism). c, Plots of the fraction of substrate that had been converted to ubiquitylated products against UBE2D concentration for ubiquitylation reactions with various substrate peptides: derived from IκBα (but with a single acceptor Lys); derived from β-catenin; derived from β-catenin with different spacing between the phosphodegron motif and a potential acceptor Lys (a medium β-catenin substrate peptide with a nine-residue spacer between the β-catenin phosphodegron and acceptor, matching the relative position of these moieties in IκBα; and a short β-catenin substrate in which the four residues between these moieties are too few to bridge the structurally observed gap between the substrate receptor and UBE2D~Ub active site); and a homogeneous ubiquitin linked-β-catenin generated by sortase-mediated transpeptidation wherein the only lysines are from ubiquitin. d, Autoradiogram of SDS–PAGE gel showing results from rapid quench-flow reactions under pre-steady-state single encounter conditions for the interaction of radiolabelled substrate (a medium β-catenin phosphopeptide) with CRL1β-TRCP. The representative raw data are from a reaction using wild-type UBE2D and wild-type neddylated CRL1β-TRCP, and show time-resolved conjugation of increasing numbers of individual ubiquitin molecules. S0, substrate with 0 ubiquitins; S1, substrate with one ubiquitin; S2, substrate with two ubiquitins; and so on. e, Plots comparing various substrate peptides described in c, showing disappearance of unmodified substrate (S0) with black circles, and the appearance of mono-ubiquitylated substrate (S1) with grey triangles, in rapid quench-flow reactions all performed as in d and under single-encounter conditions as in a. Duplicate data points from independent experiments performed with identical samples are shown. The data were fit to closed form equations (Mathematica) as previously described59 to obtain both the rates for the transfer of the first ubiquitin to substrate (kobsS0–S1) and of the second ubiquitin to the singly ubiquitin-modified substrate (kobsS1–S2) as well as their associated standard error (Extended Data Table 1). f, Plots from experiments performed and analysed as described in e, except with radiolabelled medium β-catenin peptide substrate, CRL1β-TRCP variants containing the indicated versions of CUL1–RBX1, and with either wild-type or indicated mutant versions of UBE2D.
Extended Data Fig. 2 In CRL1β-TRCP, the RING domain of RBX1 and the WHB domain of CUL1—without or with a covalently linked NEDD8— are dynamic in the absence of other factors and are harnessed in the catalytic architecture for substrate ubiquitylation with UBE2D.
a, Cryo-EM density corresponding to substrate-scaffolding regions of unneddylated CRL1β-TRCP is shown in surface representation, with the density encompassing the RING domain of RBX1 and the WHB domain of CUL1 outlined for different 3D classes in different colours corresponding to the percentage of particles in that 3D class. b, As in a but with neddylated CRL1β-TRCP, with its surfaces outlined in different classes encompassing the RING domain of RBX1, the WHB domain of CUL1, and covalently modified NEDD8. c, Refined cryo-EM density from CRL1β-TRCP reveals the substrate-scaffolding module bridging the substrate recruited to substrate receptor β-TRCP with the intermolecular C/R domain, readily fitted with crystal structures of SKP1–β-TRCP (PDB: 1P22)20 and the N-terminal domain of CUL1 and the C/R domain of CUL1–RBX1 (PDB: 1LDK)7. d, Schematic showing dynamics of NEDD8, its linked CUL1 WHB domain and the RING domain of RBX1 based on cryo-EM data in b for substrate-bound neddylated CRL1β-TRCP, and model for varying locations of the RBX1 RING-bound UBE2D~Ub relative to the substrate awaiting ubiquitylation. e, Left, schematic of the catalytic architecture based on the cryo-EM data shown in Fig. 2, representing neddylated CRL1β-TRCP-catalysed ubiquitin transfer from E2 UBE2D to an IκBα-derived substrate peptide. Right, semi-transparent version of the schematic, highlighting the three modules (substrate-scaffolding, catalytic and activation modules), their constituents and locations establishing the catalytic architecture for substrate priming by neddylated CRL1β-TRCP and UBE2D.
Extended Data Fig. 3 Generation of a stable proxy for the UBE2D~Ub–substrate intermediate, and characterization in complexes with neddylated CRL1β-TRCP by cryo-EM and biochemistry.
Gel panels in this figure are representative of two independent experiments; n = 2. a, Our strategy for trapping a mimic of the transient neddylated CRL E2~Ub–substrate complex requires that the E2 UBE2D contain only a single cysteine at the active site. However, UBE2D contains three additional cysteines (Cys21, Cys107 and Cys111). Standard replacements of cysteine by serine or alanine severely compromised activity. On the basis of the structural locations of these cysteines, we presumed that their mutation hindered formation of the RING-activated, closed, active UBE2D~Ub conformation28,29,30. We thus devised a systematic structure- and random-based approach to identify suitable replacements that qualitatively maintain wild-type levels of activity with neddylated CRLs. Structural analysis showed that Cys21 and Cys107 are in close proximity, such that mutation of both residues to alanine may generate a destabilizing cavity at this site. Combining UBE2D2(C107A) with Cys21 mutated to isoleucine, leucine or valine to compensate for the reduced hydrophobic volume led to the identification of C21I(C107A) as a suitable version for testing all other possible replacements for Cys111. A similar approach was taken for UBE2D3. A total of 48 different versions of UBE2D were tested to identify the UBE2D(C21I/C107A/C111D) mutant for chemical trapping at the remaining active site cysteine. b, Top, schematic of pulse-chase assay testing intrinsic activation of thioester-linked UBE2D~Ub intermediates. Although this is often tested by monitoring RING-dependent discharge of ubiquitin from UBE2D to free lysine, RBX1 RING-dependent activity is limited in this assay owing to sequence constraints imposed by the requirements for binding to partners other than UBE2D39. Nonetheless, substrate-independent activation of UBE2D~Ub can be readily visualized using CUL1 complexed with a previously described hyperactive mutant RBX1(N98R)39, and high enzyme and lysine concentrations. UBE2D~Ub generated in a pulse reaction was mixed with NEDD8-modified CUL1–RBX1 (shown here with the N98R mutant) and free lysine, and ubiquitin discharge was monitored over time by Coomassie-stained SDS–PAGE (as shown by the representative gel at the bottom) demonstrating that standard serine or alanine mutations of noncatalytic cysteines compromised activity (shown for the mutant C21A/C107A/C111S), whereas the optimized mutant (C21I/C107A/C111D) retains activity similar to that of the wild type. c, Overview of the generation of our stable proxy for the phosphorylated IκBα substrate intermediate linked at a single atom, and comparison to the previous method used to visualize noncanonical Lys sumoylation63. d, Experiment validating our stable proxy for the UBE2D~Ub-phosphorylated IκBα substrate intermediate linked at a single atom, based on the hypothesis that its simultaneous occupation of the binding sites for the UBE2D~Ub intermediate and substrate should result in more potent inhibition of a neddylated CRL1β-TRCP-dependent substrate priming reaction compared to the individual constituents of the complex. e, Cryo-EM reconstruction of neddylated CRL1β-TRCP2 (with full-length, dimeric β-TRCP2) bound to a mimic of UBE2D2~Ub–IκBα generated by adapting the method used previously to visualize noncanonical lysine sumoylation63. Ubiquitin is isopeptide-bonded to the substituted residue of a UBE2D(L119K) mutant, and a cysteine residue that replaces the acceptor in the substrate is disulfide-bonded to the catalytic cysteine of UBE2D2. This electron microscopy map visualizes the catalytic architecture of dimeric CRL1β-TRCP2 in which the dimerization domain agrees well with the previous crystal structure27, and its linked NEDD8 (circled in yellow) is bound to the backside of UBE2D, but the donor ubiquitin (absent from the region circled in orange) was not visible—presumably owing to inadequacies of the method used to generate this mimic of the catalytic intermediate, in which the ubiquitin and substrate are not both simultaneously linked to the UBE2D catalytic cysteine. Variations between the two protomers of the dimer also exacerbated sample heterogeneity. f, Cryo-EM reconstruction of neddylated CRL1β-TRCP1∆D (with monomeric version of β-TRCP1, from residue 175 to the C terminus20) bound to our newly developed proxy for the UBE2D3~Ub–IκBα intermediate. The phospho-IκBα peptide-substrate-bound β-TRCP–SKP1–CUL1–RBX1–NEDD8–UBE2D portion of this map superimposes with the map for the dimeric complex shown in e, but here the entire complex is visible—including both the NEDD8 (circled in yellow) and donor ubiquitin (circled in orange). g, To further increase cryo-EM sample homogeneity, we considered that the RBX1 RING sequence represents a compromise to meet requirements for its many different catalytic activities achieved with neddylation E2s, various ubiquitin carrying enzymes, and regulators including the inhibitor GLMN72. Therefore, we introduced a second RBX1 linchpin residue via mutation (N98R), which has previously been shown to improve neddylated CRL and UBE2D-dependent substrate priming at the expense of other RBX1-dependent functions (for example, with UBE2M and UBE2R2)39. A Coomassie-stained SDS–PAGE gel from an assay for the intrinsic activity of UBE2D~Ub is shown, showing enhanced neddylated CRL-dependent activation of discharge to free lysine with the RBX1 N98R mutation. h, i, Cryo-EM reconstructions of neddylated CRL1β-TRCP1∆D with RBX1(N98R) bound to our newly developed proxies for the UBE2D3~Ub–IκBα and UBE2D2~Ub–IκBα intermediates, the latter of which was pursued for high-resolution electron microscopy (final reconstruction refined to 3.7 Å resolution, shown on right).
Extended Data Fig. 4 Flow chart showing the stages of cryo-EM image processing.
a, Cryo-EM image-processing flow chart. Ultimately, reconstruction of the data yielded a focused refinement at 3.46 Å resolution and a global refinement at 3.7 Å resolution that superimposes well with lower-resolution maps that were obtained during attempts to visualize substrate priming with neddylated wild-type dimeric CRL1β-TRCP. b, Two-dimensional classes representing particles used for final reconstructions. c, Angular distribution of final reconstruction. d, Gold-standard Fourier shell correlation (FSC) curve showing overall resolution at 3.72 Å at an FSC of 0.143. e, Electron microscopy density map coloured by local resolution. NEDD8, circled in yellow, is the entity displaying the highest local resolution in the map.
Extended Data Fig. 5 Extraordinary cullin–RING conformational changes in catalytic architecture juxtaposing the substrate and the active site of ubiquitylation.
a, Side-by-side comparison of relative RING-domain locations in different CRL complexes after superposition of the C/R domains from the original CUL1–RBX1 structure (PDB: 1LDJ, ‘pre-neddylation’—which data herein show is dynamic—although the crystal structure probably captured the conformation that enabled CAND1 binding and substrate receptor exchange)7, the structure representing the neddylation reaction (PDB: 4P5O)39, and a structure of a neddylated CUL5–RBX1 domain (PDB: 3DQV, labelled ‘post-neddylation’, which revealed the potential for conformational changes in the neddylated CUL WHB- and RBX1 RING-domains3, and data herein shows is dynamic), and the structure presented here showing how the neddylated CUL1 WHB domain and RBX1 RING domain are harnessed in a catalytic architecture for ‘active ubiquitylation’. Trp35 of RBX1 is highlighted to show how it serves as a multifunctional platform for either the RING domain in different orientations, or for the E2-linked NEDD8 during neddylation39. b, Superposition of the structures shown in a, highlighting different relative positions of the RING domain. c, Comparison of the relative locations of the CUL WHB domain in different structures after superimposing their C/R domains (not shown). d, Cryo-EM density from the neddylated CRL1β-TRCP–UBE2D~Ub–substrate intermediate complex, showing patchiness of the region corresponding to CUL1 ‘helix-29’7. This CUL1 region connecting the C/R and WHB domains is visible only as patchy density, whereas in previous cullin crystals this forms the rod-like helix-29 continuing into the WHB domain7. It seems that helix-29 of CUL1 dissolves into a flexible tether, which rationalizes the previously observed proteolytic sensitivity of this region in a neddylated CUL1–RBX1 complex3, and enables the displacement and rotation required for placing the ensuing WHB domain and its linked NEDD8 at the centre of the ubiquitylation complex.
Extended Data Fig. 6 Geometry between phosphodegron and acceptor in structure, substrates and ubiquitylation.
a, Cryo-EM density highlighting the relative placement of the substrate degron and the UBE2D~Ub active site. The approximately 22 Å distance between the UBE2D~Ub active site and the phosphodegron of β-TRCP-bound substrate requires at least 6 intervening residues in a substrate. b, Alignments for several reported β-TRCP substrates32, highlighting the degron sequence (yellow) and nearby lysines (red). Also shown are sequences of peptide substrates with a single acceptor Lys that were used in kinetics analyses. The peptide sequences were derived from IκBα, and from β-catenin with varying spacers between phosphodegron and acceptor Lys: wild-type β-catenin peptide, medium β-catenin peptide with lysine corresponding to IκBα, and short β-catenin peptide with a lysine five residues upstream of the N-terminal phosphoserine in the degron, which would be too short to bridge the structurally observed distance between the phosphodegron binding site on β-TRCP and the UBE2D catalytic cysteine in the ubiquitylation active site. c, Representative autoradiogram (n = 2) of SDS–PAGE gel showing products from indicated time points of ubiquitylation reactions under multiturnover conditions with either neddylated or unneddylated CRL1β-TRCP and radiolabelled short β-catenin peptide substrate. The amount of short β-catenin peptide modified by neddylated CRL1β-TRCP and UBE2D is too low in the single-encounter ubiquitylation reaction to enable quantification of kinetic parameters; however, product formation is apparent under multi-turnover conditions and shows that most products are heavily ubiquitylated. d, Plots fitting consumption of unmodified short β-catenin peptide substrate (S0) compared to formation of polyubiquitin chains with five or more ubiquitins (S5+) from reactions as in c. The symbols show the data from independent experiments (n = 2 technical replicates).
Extended Data Fig. 7 Interactions shaping the catalytic architecture of neddylated CRL1β-TRCP–UBE2D~Ub–IκBα substrate intermediate.
a, NEDD8 and the catalytic module from the structure representing the neddylated CRL1β-TRCP–UBE2D~Ub–IκBα intermediate, highlighting distinctive interactions between NEDD8 (yellow) and donor ubiquitin (orange) with UBE2D. b, Catalytic module from the neddylated CRL1β-TRCP–UBE2D~Ub–IκBα intermediate, highlighting the covalently linked proxy for the IκBα substrate’s acceptor in the active site relative to a superimposed representative previous crystal structure of an isolated RING–UBE2D~Ub complex (grey, PDB: 4AP4)28,29. In the inset, the density for the covalently linked proxy for IκBα substrate’s acceptor is shown in red in the active site. The chemical trap superimposes with consensus acceptors visualized in active sites of sumoylation63 and neddylation39 intermediates, where aromatic side chains guide the lysine targets (blue and green, respectively)39,73. However, the myriad substrates of UBE2D neither conform to a specific motif nor do they or UBE2D display specific side chains that guide lysine acceptors into the catalytic centre. Instead, in the neddylated CRL1β-TRCP–UBE2D~Ub–substrate complex, density from backbone atoms preceding the chemical proxy for the acceptor lysine corresponds to the aromatic guides in sumoylation and neddylation intermediates. c, Overview of assays for activation of intrinsic reactivity of the UBE2D~Ub intermediate. Top, schematic of pulse-chase assay for testing the effects of UBE2D mutations on activation, monitoring UBE2D~Ub discharge to free lysine activated by neddylated CUL–RBX1 compared to unneddylated or RING-like UBE4B controls. Bottom, sites of mutations shown as spheres on the structure of UBE2D from the cryo-EM structure of neddylated CRL1β-TRCP–UBE2D~Ub–substrate complex. The colours of spheres reflect both the locations and the effects on UBE2D~Ub discharge to free lysine. Sites of mutations with marginal or no effect are shown in cyan, whereas those with major effects are otherwise coloured. Mutations that cause major defects map to the RBX1 RING-binding site (blue), the interaction surface with the donor ubiquitin (orange), and the interaction surface with NEDD8 (yellow). d, Representative Coomassie-stained SDS–PAGE gels (of two independent experiments) shown for reactions monitoring substrate-independent discharge of UBE2D~Ub to free lysine, in the presence of CUL1–RBX1(N98R) that was either neddylated or unneddylated (K720R), with either wild-type or the indicated UBE2D3 mutants at binding sites for backside-bound NEDD8(S22R), the RBX1 RING(F62A), and the covalently linked donor ubiquitin in the closed conformation (S108L). e, As in d, except testing the effect of NEDD8(Q40E), which would disrupt the activation module. f, Reactions performed as in d, except with indicated variants of UBE2D2, in reactions with CUL1–RBX1(N98R) that was either neddylated or unneddylated (K720R), or with the optimized RING-like U-box domain from UBE4B as a reference58. For mutations reporting on the catalytic conformation (G24K, T36K, M38K, A96K and D112K), representative gels are shown for two experiments. All other experiments were performed once. g, Comparison of β1/β2-loop conformations after superimposing the indicated structures of NEDD8 and ubiquitin. The comparison suggests that whereas NEDD8 and ubiquitin can adopt both loop-in and loop-out conformations, donors linked to E2 active sites in RING-activated complexes adopt the loop-in conformation, and those bound to the UBE2D backside adopt loop-out conformations. h, Only a loop-out conformation is compatible with the neddylated CRL activation module structure, because loop-in conformation from the structures shown in g would prevent noncovalent interactions with the WHB domain of CUL1 (green). i, Only a loop-out conformation is compatible for the CUL1-linked NEDD8 to bind the catalytic module, because loop-in conformations in the structures shown in g would prevent noncovalent interactions with the backside of UBE2D (cyan).
Extended Data Fig. 8 Qualitative validation of the mechanistic principles that underlie substrate priming by neddylated CRLs and UBE2D.
Gel panels in this figure are representative of two independent experiments; n = 2 technical replicates. a, Schematic of a qualitative substrate priming assay for testing effects of mutations in neddylated CRL1β-TRCP or UBE2D on substrate priming, monitoring fluorescent ubiquitin transfer from UBE2D3 to the phosphorylated IκBα substrate. b, Scan of SDS–PAGE detecting fluorescent ubiquitin transferred to the IκBα-derived substrate in a qualitative assay for NEDD8 activation of substrate priming. c, As in b, showing the effect on substrate priming of disrupting the activation module with the Q40E mutation of NEDD8. d, As in b, showing the effect on substrate priming of disrupting interactions between the activation and catalytic modules with the NEDD8(I44A) or UBE2D(S22R) mutants. e, As in b, showing the effect on substrate priming of disrupting interactions between the activation and substrate-scaffolding modules, though CUL1 modification by a ‘ubiquitylized’ NEDD8 mutant with six residues swapped for their ubiquitin counterparts (L2Q/K4F/E14T/D16E/G63K/G64E). f, As in b, showing the effect of the H32A in UBE2D at the interface between the catalytic and substrate-scaffolding modules. g, Scheme of pulse-chase assay for testing the effects of mutations in neddylated CRL1FBW7 or UBE2D on substrate priming. The assay monitors the transfer of fluorescent ubiquitin from UBE2D to peptide substrate derived from phosphorylated cyclin E (pCyE). h, Fluorescent scan detecting ubiquitin transferred to the pCyE substrate by neddylated CRL1FBW7 and the indicated mutants of UBE2D. i, Fluorescent scan detecting ubiquitin transferred to the pCyE substrate by UBE2D and indicated variants of neddylated (or ubiquitylated) CRL1FBW7. Experiment with unneddylated CRL1FBW7 used the K720R variant of CUL1 to prevent artefactual ubiquitylation. j, Scheme of pulse-chase assay for testing effects of mutations in neddylated CRL4CRBN or UBE2D on substrate priming, monitoring fluorescent ubiquitin transfer from UBE2D to the IKZF1/3 ZF2 substrate in the presence of the immunomodulatory drug pomalidomide. k, Fluorescent scan of assay validation, showing dependence on pomalidomide. l, Fluorescent scan detecting ubiquitin transferred to the IKZF substrate by CRL4CRBN, pomalidomide and the indicated variants of UBE2D. m–o, Fluorescent scan detecting ubiquitin transferred to the IKZF substrate by UBE2D and the indicated variants of neddylated (or ubiquitylated) CRL4CRBN with pomalidomide. Experiments with unneddylated CRL4CRBN used the K705R variant of CUL4A to prevent artefactual ubiquitylation.
Supplementary information
Supplementary Information
This file contains uncropped raw gel images.
Rights and permissions
About this article
Cite this article
Baek, K., Krist, D.T., Prabu, J.R. et al. NEDD8 nucleates a multivalent cullin–RING–UBE2D ubiquitin ligation assembly. Nature 578, 461–466 (2020). https://doi.org/10.1038/s41586-020-2000-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2000-y
This article is cited by
-
Mechanism of millisecond Lys48-linked poly-ubiquitin chain formation by cullin-RING ligases
Nature Structural & Molecular Biology (2024)
-
Structural snapshots along K48-linked ubiquitin chain formation by the HECT E3 UBR5
Nature Chemical Biology (2024)
-
Structural mechanisms of autoinhibition and substrate recognition by the ubiquitin ligase HACE1
Nature Structural & Molecular Biology (2024)
-
Dynamic molecular architecture and substrate recruitment of cullin3–RING E3 ligase CRL3KBTBD2
Nature Structural & Molecular Biology (2024)
-
Protein neddylation and its role in health and diseases
Signal Transduction and Targeted Therapy (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.