Abstract
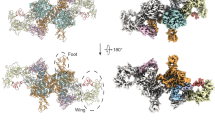
Thyroglobulin (TG) is the protein precursor of thyroid hormones, which are essential for growth, development and the control of metabolism in vertebrates1,2. Hormone synthesis from TG occurs in the thyroid gland via the iodination and coupling of pairs of tyrosines, and is completed by TG proteolysis3. Tyrosine proximity within TG is thought to enable the coupling reaction but hormonogenic tyrosines have not been clearly identified, and the lack of a three-dimensional structure of TG has prevented mechanistic understanding4. Here we present the structure of full-length human thyroglobulin at a resolution of approximately 3.5 Å, determined by cryo-electron microscopy. We identified all of the hormonogenic tyrosine pairs in the structure, and verified them using site-directed mutagenesis and in vitro hormone-production assays using human TG expressed in HEK293T cells. Our analysis revealed that the proximity, flexibility and solvent exposure of the tyrosines are the key characteristics of hormonogenic sites. We transferred the reaction sites from TG to an engineered tyrosine donor–acceptor pair in the unrelated bacterial maltose-binding protein (MBP), which yielded hormone production with an efficiency comparable to that of TG. Our study provides a framework to further understand the production and regulation of thyroid hormones.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Datasets generated during the current study are available from the RCSB Protein Data Bank (PDB) accession code 6SCJ, Electron Microscopy Data Bank (EMDB) accession code EMD-10141 and ProteomeXchange accession code PXD014821. All other data generated or analysed during this study are included in this published article and its Supplementary Information.
References
Di Jeso, B. & Arvan, P. Thyroglobulin from molecular and cellular biology to clinical endocrinology. Endocr. Rev. 37, 2–36 (2016).
Citterio, C. E., Targovnik, H. M. & Arvan, P. The role of thyroglobulin in thyroid hormonogenesis. Nat. Rev. Endocrinol. 15, 323–338 (2019).
Carvalho, D. P. & Dupuy, C. Thyroid hormone biosynthesis and release. Mol. Cell. Endocrinol. 458, 6–15 (2017).
Cahnmann, H. J., Pommier, J. & Nunez, J. Spatial requirement for coupling of iodotyrosine residues to form thyroid hormones. Proc. Natl Acad. Sci. USA 74, 5333–5335 (1977).
Holzer, G. et al. Thyroglobulin represents a novel molecular architecture of vertebrates. J. Biol. Chem. 291, 16553–16566 (2016).
Taylor, P. N. et al. Global epidemiology of hyperthyroidism and hypothyroidism. Nat. Rev. Endocrinol. 14, 301–316 (2018).
Sellitti, D. F. & Suzuki, K. Intrinsic regulation of thyroid function by thyroglobulin. Thyroid 24, 625–638 (2014).
Xiao, S., Dorris, M. L., Rawitch, A. B. & Taurog, A. Selectivity in tyrosyl iodination sites in human thyroglobulin. Arch. Biochem. Biophys. 334, 284–294 (1996).
Heidelberger, M. The molecular weight of thyroglobulin. Science 80, 414 (1934).
Gavaret, J. M., Cahnmann, H. J. & Nunez, J. Thyroid hormone synthesis in thyroglobulin. J. Biol. Chem. 256, 9167–9173 (1981).
Mondal, S., Raja, K., Schweizer, U. & Mugesh, G. Chemistry and biology in the biosynthesis and action of thyroid hormones. Angew. Chem. Int. Ed. 55, 7606–7630 (2016).
Gnidehou, S. et al. Iodotyrosine dehalogenase 1 (DEHAL1) is a transmembrane protein involved in the recycling of iodide close to the thyroglobulin iodination site. FASEB J. 18, 1574–1576 (2004).
Dedieu, A., Gaillard, J.-C., Pourcher, T., Darrouzet, E. & Armengaud, J. Revisiting iodination sites in thyroglobulin with an organ-oriented shotgun strategy. J. Biol. Chem. 286, 259–269 (2011).
Aricescu, A. R., Lu, W. & Jones, E. Y. A time- and cost-efficient system for high-level protein production in mammalian cells. Acta Crystallogr. D 62, 1243–1250 (2006).
Berg, G., Björkman, U. & Ekholm, R. The structure of newly synthesized intracellular thyroglobulin molecules. Mol. Cell. Endocrinol. 20, 87–98 (1980).
Gunčar, G., Pungerčič, G., Klemenčič, I., Turk, V. & Turk, D. Crystal structure of MHC class II-associated p41 Ii fragment bound to cathepsin L reveals the structural basis for differentiation between cathepsins L and S. EMBO J. 18, 793–803 (1999).
Molina, F., Bouanani, M., Pau, B. & Granier, C. Characterization of the type-1 repeat from thyroglobulin, a cysteine-rich module found in proteins from different families. Eur. J. Biochem. 240, 125–133 (1996).
Lee, J. & Arvan, P. Repeat motif-containing regions within thyroglobulin. J. Biol. Chem. 286, 26327–26333 (2011).
Citterio, C. E., Morishita, Y., Dakka, N., Veluswamy, B. & Arvan, P. Relationship between the dimerization of thyroglobulin and its ability to form triiodothyronine. J. Biol. Chem. 293, 4860–4869 (2018).
Yang, S.-X., Pollock, H. G. & Rawitch, A. B. Glycosylation in human thyroglobulin: location of the N-linked oligosaccharide units and comparison with bovine thyroglobulin. Arch. Biochem. Biophys. 327, 61–70 (1996).
Lamas, L., Anderson, P. C., Foxy, J. W. & Dunn, J. T. Consensus sequences for early iodination and hormonogenesis in human thyroglobulin. J. Biol. Chem. 264, 13541–13545 (1989).
Pommier, J., Deme, D. & Nunez, J. Effect of iodide concentration on thyroxine synthesis catalysed by thyroid peroxidase. Eur. J. Biochem. 37, 406–414 (1973).
de Vijlder, J. J. & den Hartog, M. T. Anionic iodotyrosine residues are required for iodothyronine synthesis. Eur. J. Endocrinol. 138, 227–231 (1998).
Dunn, J. T., Kim, P. S. & Dunn, A. D. Favored sites for thyroid hormone formation on the peptide chains of human thyroglobulin. J. Biol. Chem. 257, 88–94 (1982).
Botta, R. et al. Sortilin is a putative postendocytic receptor of thyroglobulin. Endocrinology 150, 509–518 (2009).
Weber, J. et al. Interdependence of thyroglobulin processing and thyroid hormone export in the mouse thyroid gland. Eur. J. Cell Biol. 96, 440–456 (2017).
Rivolta, C. M. & Targovnik, H. M. Molecular advances in thyroglobulin disorders. Clin. Chim. Acta 374, 8–24 (2006).
Latrofa, F. et al. Thyroglobulin autoantibodies in patients with papillary thyroid carcinoma: comparison of different assays and evaluation of causes of discrepancies. J. Clin. Endocrinol. Metab. 97, 3974–3982 (2012).
Fiore, E., Latrofa, F. & Vitti, P. Iodine, thyroid autoimmunity and cancer. Eur. Thyroid J. 4, 26–35 (2015).
Guo, J., McLachlan, S. M., Hutchison, S. & Rapoport, B. The greater glycan content of recombinant human thyroid peroxidase of mammalian than of insect cell origin facilitates purification to homogeneity of enzymatically protein remaining soluble at high concentration. Endocrinology 139, 999–1005 (1998).
Bokori-Brown, M. et al. Cryo-EM structure of lysenin pore elucidates membrane insertion by an aerolysin family protein. Nat. Commun. 7, 11293 (2016).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Li, Y. et al. Mechanistic insights into caspase-9 activation by the structure of the apoptosome holoenzyme. Proc. Natl Acad. Sci. USA 114, 1542–1547 (2017).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Turk, D. MAIN software for density averaging, model building, structure refinement and validation. Acta Crystallogr. D 69, 1342–1357 (2013).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Söding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Mendes, M. L. et al. An integrated workflow for crosslinking mass spectrometry. Mol. Syst. Biol. 15, e8994 (2019).
Fischer, L. & Rappsilber, J. Quirks of error estimation in cross-linking/mass spectrometry. Anal. Chem. 89, 3829–3833 (2017).
Ledeţi, I. et al. Thermal stability of synthetic thyroid hormone l-thyroxine and l -thyroxine sodium salt hydrate both pure and in pharmaceutical formulations. J. Pharm. Biomed. Anal. 125, 33–40 (2016).
Acknowledgements
We thank C. Savva, G. Cannone and S. Chen for help with electron microscopes; S. Scheres, R. F. Leiro, J. Zivanov, P. Emsley, V. Chandrasekaran and S. Masilius for image processing and model building advice; R. Aricescu for help with the expression of TG in mammalian cells; C. Heroven and D. Laverty for help with mammalian tissue culture; F. Bürmann and G. Slodkowicz for help with data analysis; F. van den Ent and T. Nierhaus for advice on protein work; T. Darling and J. Grimmett for computing support; and D. Clare for electron microscopy data collection. We acknowledge the Diamond Light Source for access and support of the cryo-EM facilities at the UK’s National Electron Bio-imaging Centre (eBIC), funded by the Wellcome Trust, MRC and BBRSC. This work was funded by the Medical Research Council (U105184326 to J.L.), the Wellcome Trust (202754/Z/16/Z to J.L., 203149 to J.R.) and by the Slovenian Research Agency (ARRS; P1-0048, IO-0048 and J1-7479 to D.T.). This work was supported by the Wellcome Trust through a Senior Research Fellowship (103139 to J.R.) and by the DFG, German Research Foundation (329673113 and EXC 2008/1 – 390540038 to J.R.).
Author information
Authors and Affiliations
Contributions
F.C. performed TG mammalian expression, cryo-EM, TG biochemistry, ELISA design and data analysis. F.C., J.L. and D.T. built and refined the TG model. A.T.-V. and I.B. performed TPO biochemistry. V.T.C. performed expression of TG in mammalian cells with F.C. T.I. collected initial negative staining data on eTG. M.R. developed initial eTG purifications. L.S., F.J.O. and J.R. performed crosslinking mass-spectrometry analysis. F.C. and J.L. wrote the manuscript with assistance of other authors. J.L., F.C., D.T. and A.T.-V. were responsible for project strategy and data interpretation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Ulrich Schweizer, Janet Vonck and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The iodine cycle in the thyroid gland and the chemistry of thyroid-hormone formation.
a, Iodide is extracted from the blood vessels and into the thyroid cells via the Na+ and I− symporter. TSH binds TSH receptor (TSHR) to induce the expression of TG. TG is secreted into the extracellular lumen of follicular cells (also called colloid). DUOX and TPO catalyse the iodination of TG; therefore, T4 (or T3) hormones are formed on the TG polypeptide chain. After hormonogenesis, TG is reimported and proteolysed in lysosomes to release T4/T3 into the blood. DEHAL1 de-iodinates iodotyrosines to recycle iodide in thyroid cells. b, T4 (or T3) synthesis from TG in the thyroid gland.
Extended Data Fig. 2 Cryo-EM reconstruction of eTG and rTG.
a, SDS–PAGE of eTG from extracts from goitrous thyroid, and rTG expressed in HEK293T cells. b, Cryo-EM micrograph of eTG with calculated reference-free 2D class averages below. Scale bar, 200 Å. c, Cryo-EM micrograph of recombinant rTG with 2D class averages, showing the two proteins to be structurally identical at this level of analysis. Scale bar, 200 Å. d, Schematic illustrating the C2 symmetry expansion and recentring procedure, which was used to enhance TG map quality in peripheral regions. For a detailed procedure see ‘Cryo-EM image processing’ in Methods. e, Local resolution of the C2 and symmetry-expanded and recentred eTG maps. f, Flexibility of NTD results in varying map quality and occupancy of this region in a number of 3D class averages (calculated in RELION). g, FSC between RELION ‘gold standard’ half-maps and between the final eTG and rTG maps, showing their strong similarity.
Extended Data Fig. 3 Local properties of the atomic TG model.
a, Per-residue atomic B-factor and cross-correlation with the rTG map, plotted per residue number. b, Local B-factor colour-coded onto the surface of the TG structure. c, FSC between the map and model calculated for rTG. FSC 0.5 is indicated.
Extended Data Fig. 4 Validation of the 3D architecture of TG by mass spectrometry crosslinking.
a, Size-exclusion chromatograms of rTG before and after BS3 crosslinking and subsequent SDS–PAGE (Coomassie-stained). b, Negative-staining micrograph of crosslinked rTG, showing the absence of higher-order structures caused by unwanted inter-dimer crosslinks. c, Plot representing experimental crosslinks (circles) overlapping with predicted crosslinks, calculated from the structure determined here. d–f, Detail of key TG interfaces confirmed by the crosslinking.
Extended Data Fig. 5 Details of TG cryo-EM map.
a, All disulfide bonds in TG included in the model (yellow spheres). b, Glycans detected in the cryo-EM maps and included in the TG atomic model (green spheres). c, Close-up view of a typical α-helix in the TG cryo-EM map (part of the core region). d, Close-up of a β-sheet in TG (part of the ChEL domain). e, Close-up of the disulfide bond C900–C921 (core region). f, Map details of N2013 and N-linked GlcNAc between two TG subunits. g, Close-up views showing conformational disorder within the hormonogenic sites (making precise side-chain placements difficult), but the backbone positions are resolved (rTG map).
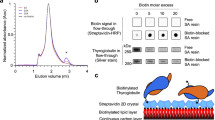
Extended Data Fig. 6 Quantitative thyroid-hormone ELISA assays.
a, Schematic summarizing in vitro thyroid-hormone synthesis and quantification via ELISA assays. b, T4 assay calibration curves with added T4, under manufacturer-recommended and modified (T4 synthesis as performed here) conditions. c, Validation of the T4 ELISA assay. eTG presumably contains already-reacted tyrosine side chains. rTG produces T4. Addition of iodide is required for the reaction to occur. Lactoperoxidase is as active as TPO, taking the reduced 20% haem content in our TPO into account. Lysozyme (some tyrosines), FtsZ from S. aureus (no tyrosines) and T3 produce no T4 signal. d, Mutating residues in hormonogenic site D in a version of TG that is active only in site D shows that a conserved lysine residue is not important for the reaction. Adding an extra Ser-Asp before Y1310 has no effect, but the mutation D1309S abolished activity. e, Synthesis of T4 from tyrosine copolymers as measured by the T4 ELISA assay. Only a polymer in which tyrosines are spaced apart and preceded by Lys-Asp produces some T4. Activity is lower than in a single site of TG (or MBP, compare with Fig. 3). f, T3 assay calibration curves with added T3 under recommended and modified (as for T4) conditions. g, No substantial T3 production was detected from iodinated rTG or eTG from goitre. In c–e, g, bar plot and error bars indicate mean and s.d., n = 3.
Extended Data Fig. 7 Tyrosine-pair proximity plots for TG and MBP.
a, b, Proximity plots of tyrosine residues closer than 15 Å to each other, calculated from TG (a) and MBP (b) atomic models (TG, this study; MBP, PDB accession code 1ANF). The coordinates of each point in the plot represent a tyrosine-pair position (residue number). For the TG dimer, the distance between tyrosines from the same or the other subunit in the dimer are shown in grey or black, respectively. In TG, there are no more than five pairs that are exposed and in <15 Å proximity at the same time, predicting the absence of other sites important for hormonogenesis. In MBP, only one pair closer than 15 Å is sufficiently exposed to be a candidate for hormonogenesis. c, d, Ribbon diagram of TG and MBP in which tyrosine residues are represented as spheres and coloured by B-factor, which largely indicates solvent-exposed residues.
Supplementary information
Supplementary Information
Supplementary information 1: Amino acid sequences of recombinant proteins generated and used in this study. Supplementary information 2: Images of the uncropped gels presented in this study. A) Figure 3b. B) Extended Data Figure 5a.
Supplementary Table
Supplementary Table 1: List of BS3 crosslinks detected by mass spectrometry.
Supplementary Video 1
The architecture of human thyroglobulin (TG) and its T4 hormonogenic sites. TG is a convoluted dimer and several glycans are proposed to be important for dimer stability since they are found at the interface between two subunits. The arrangement of TG's domains produces an intertwined structure that serves as a scaffold for flexible regions that contain the reactive tyrosine pairs. Highlighted in yellow are the hormonogenic tyrosine pairs in Sites A-D that we identified via our structural and mutational analysis. Donor and acceptor tyrosines in flexible and exposed areas are readily iodinated and undergo the aromatic ring rearrangement that leads to T4 synthesis.
Supplementary Video 2
Engineering tyrosine pairs in maltose binding protein (MBP) to produce thyroid hormones.The analysis of the B-factors of all tyrosines in MBP highlights the presence of exposed and hence more flexible residues, which we utilised to make TG-like hormonogenic sites, to form T4. Y171 and Y176, present in the unmodified sequence, showed only marginal activity, and engineering the tyrosine pair Y341/R367Y did not produce additional activity, presumably because they are too rigidly attached to the backbone of the proteins. Only after creating a flexible and long insertion at the C-terminus (C-ins2) did tyrosine coupling occur as efficiently as in a single TG site, as detected by T4 ELISA.
Rights and permissions
About this article
Cite this article
Coscia, F., Taler-Verčič, A., Chang, V.T. et al. The structure of human thyroglobulin. Nature 578, 627–630 (2020). https://doi.org/10.1038/s41586-020-1995-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-1995-4
This article is cited by
-
Comparing the efficacy of thyroglobulin and thyroglobulin/ thyroid-stimulating hormone ratio models in predicting a successful response to radioactive iodine therapy
BMC Endocrine Disorders (2023)
-
Epigenetics of the far northern Yakutian population
Clinical Epigenetics (2023)
-
Radiomics features from whole thyroid gland tissue for prediction of cervical lymph node metastasis in the patients with papillary thyroid carcinoma
Journal of Cancer Research and Clinical Oncology (2023)
-
Form factor determination of biological molecules with X-ray free electron laser small-angle scattering (XFEL-SAS)
Communications Biology (2023)
-
Comparative analysis of human and bovine thyroglobulin structures
Journal of Analytical Science and Technology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.