Abstract
Xist represents a paradigm for the function of long non-coding RNA in epigenetic regulation, although how it mediates X-chromosome inactivation (XCI) remains largely unexplained. Several proteins that bind to Xist RNA have recently been identified, including the transcriptional repressor SPEN1,2,3, the loss of which has been associated with deficient XCI at multiple loci2,3,4,5,6. Here we show in mice that SPEN is a key orchestrator of XCI in vivo and we elucidate its mechanism of action. We show that SPEN is essential for initiating gene silencing on the X chromosome in preimplantation mouse embryos and in embryonic stem cells. SPEN is dispensable for maintenance of XCI in neural progenitors, although it significantly decreases the expression of genes that escape XCI. We show that SPEN is immediately recruited to the X chromosome upon the upregulation of Xist, and is targeted to enhancers and promoters of active genes. SPEN rapidly disengages from chromatin upon gene silencing, suggesting that active transcription is required to tether SPEN to chromatin. We define the SPOC domain as a major effector of the gene-silencing function of SPEN, and show that tethering SPOC to Xist RNA is sufficient to mediate gene silencing. We identify the protein partners of SPOC, including NCoR/SMRT, the m6A RNA methylation machinery, the NuRD complex, RNA polymerase II and factors involved in the regulation of transcription initiation and elongation. We propose that SPEN acts as a molecular integrator for the initiation of XCI, bridging Xist RNA with the transcription machinery—as well as with nucleosome remodellers and histone deacetylases—at active enhancers and promoters.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
References
Minajigi, A. et al. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, aab2276 (2015).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Chu, C. et al. Systematic discovery of Xist RNA binding proteins. Cell 161, 404–416 (2015).
Monfort, A. et al. Identification of Spen as a crucial factor for Xist function through forward genetic screening in haploid embryonic stem cells. Cell Rep. 12, 554–561 (2015).
Moindrot, B. et al. A pooled shRNA screen identifies Rbm15, Spen, and Wtap as factors required for Xist RNA-mediated silencing. Cell Rep. 12, 562–572 (2015).
Nesterova, T. B. et al. Systematic allelic analysis defines the interplay of key pathways in X chromosome inactivation. Nat. Commun. 10, 3129 (2019).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Schulz, E. G. et al. The two active X chromosomes in female ESCs block exit from the pluripotent state by modulating the ESC signaling network. Cell Stem Cell 14, 203–216 (2014).
Yabe, D. et al. Generation of a conditional knockout allele for mammalian Spen protein Mint/SHARP. Genesis 45, 300–306 (2007).
Borensztein, M. et al. Xist-dependent imprinted X inactivation and the early developmental consequences of its failure. Nat. Struct. Mol. Biol. 24, 226–233 (2017).
Grimm, J. B. et al. A general method to improve fluorophores for live-cell and single-molecule microscopy. Nat. Methods 12, 244–250 (2015).
Masui, O., Heard, E. & Koseki, H. in X-Chromosome Inactivation (ed. Sado, T.) Methods Mol. Biol. Vol. 1861, 67–72 (Humana, 2018).
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016).
Deng, X. et al. Bipartite structure of the inactive mouse X chromosome. Genome Biol. 16, 152 (2015).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Lu, Z. et al. RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 165, 1267–1279 (2016).
Wutz, A., Rasmussen, T. P. & Jaenisch, R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nat. Genet. 30, 167–174 (2002).
Shi, Y. et al. Sharp, an inducible cofactor that integrates nuclear receptor repression and activation. Genes Dev. 15, 1140–1151 (2001).
Oswald, F. et al. RBP-Jκ/SHARP recruits CtIP/CtBP corepressors to silence Notch target genes. Mol. Cell. Biol. 25, 10379–10390 (2005).
Ha, N. et al. Live-cell imaging and functional dissection of Xist RNA reveal mechanisms of X chromosome inactivation and reactivation. iScience 8, 1–14 (2018).
Ariyoshi, M. & Schwabe, J. W. R. A conserved structural motif reveals the essential transcriptional repression function of Spen proteins and their role in developmental signaling. Genes Dev. 17, 1909–1920 (2003).
Oswald, F. et al. A phospho-dependent mechanism involving NCoR and KMT2D controls a permissive chromatin state at Notch target genes. Nucleic Acids Res. 44, 4703–4720 (2016).
Guenther, M. G., Barak, O. & Lazar, M. A. The SMRT and N-CoR corepressors are activating cofactors for histone deacetylase 3. Mol. Cell. Biol. 21, 6091–6101 (2001).
Żylicz, J. J. et al. The implication of early chromatin changes in X chromosome inactivation. Cell 176, 182–197 (2019).
Patil, D. P. et al. m6A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537, 369–373 (2016).
Bornelöv, S. et al. The nucleosome remodeling and deacetylation complex modulates chromatin structure at sites of active transcription to fine-tune gene expression. Mol. Cell 71, 56–72 (2018).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013).
Knuckles, P. et al. Zc3h13/Flacc is required for adenosine methylation by bridging the mRNA-binding factor Rbm15/Spenito to the m6A machinery component Wtap/Fl(2)d. Genes Dev. 32, 415–429 (2018).
Hatchell, E. C. et al. SLIRP, a small SRA binding protein, is a nuclear receptor corepressor. Mol. Cell 22, 657–668 (2006).
de Vries, W. N. et al. Expression of Cre recombinase in mouse oocytes: a means to study maternal effect genes. Genesis 26, 110–112 (2000).
Zylicz, J. J. et al. G9a regulates temporal preimplantation developmental program and lineage segregation in blastocyst. eLife 7, e33361 (2018).
Tang, F. et al. RNA-seq analysis to capture the transcriptome landscape of a single cell. Nat. Protoc. 5, 516–535 (2010).
Huang, Y. et al. Stella modulates transcriptional and endogenous retrovirus programs during maternal-to-zygotic transition. eLife 6, e22345 (2017).
Belaghzal, H., Dekker, J. & Gibcus, J. H. Hi-C 2.0: An optimized Hi-C procedure for high-resolution genome-wide mapping of chromosome conformation. Methods 123, 56–65 (2017).
Chen, J. et al. High efficiency of HIV-1 genomic RNA packaging and heterozygote formation revealed by single virion analysis. Proc. Natl Acad. Sci. USA 106, 13535–13540 (2009).
Barau, J. et al. The DNA methyltransferase DNMT3C protects male germ cells from transposon activity. Science 354, 909–912 (2016).
Poullet, P., Carpentier, S. & Barillot, E. myProMS, a web server for management and validation of mass spectrometry-based proteomic data. Proteomics 7, 2553–2556 (2007).
Valot, B., Langella, O., Nano, E. & Zivy, M. MassChroQ: a versatile tool for mass spectrometry quantification. Proteomics 11, 3572–3577 (2011).
Acknowledgements
We thank K. Ancelin for support throughout the project and help with in vivo experiments; J. Barau, D. Holoch and R. Margueron for support and help with the project; M. Carrara for critical reading of the manuscript; and members of the Heard laboratory for discussions. We thank E. Nora for sharing the OsTIR1 and AID-targeting plasmids; T. Honjo for sharing the Spenflox mouse line; O. Masui for sharing the Xist–Bgl stem–loop targeting constructs and L. Lavis for sharing Halo-JF646 and JF549 with us. We also thank the imaging platform (A. Dauphin and PICT-IBiSA (UMR3215/U934)) and the protein purification and sequencing platforms of Institut Curie as well as L. Villacorta, J. Provaznik and V. Benes of GeneCore at EMBL. This work was funded by a Boehringer Ingelheim doctoral fellowship (20017-2019 to F. Dossin), an ERC Advanced Investigator award (ERC-ADG-2014 671027 to E.H.), Labellisation La Ligue (to E.H.), ANR (DoseX 2017: ANR-17-CE12-0029, Labex DEEP: ANR-11- LBX-0044, ABS4NGS: ANR-11-BINF-0001, and part of the IDEX PSL: ANR-10-IDEX-0001-02 PSL to E.H.), a Sir Henry Wellcome Postdoctoral Fellowship (201369/Z/16/Z to J.J.Z.), ‘Région Ile-de-France’ and Fondation pour la Recherche Médicale grants (to D.L.), EMBO long-term fellowships (ALTF 549-2014 to I.P., ALTF 301-2015 to T.C.), and Fondation pour la Recherche Medicale (SPF 20140129387 to I.P.) and NIH (HG003143 to J.D.) grants. J.D. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
F. Dossin and E.H. conceived the experiments and F. Dossin performed them unless stated otherwise. T.M. provided the Spen-knockout mice and J.J.Z. performed the embryo experiments. E.H. and I.P. supervised the work. J.R. established the Xist stem–loop cell line and acquired live-cell images. F. Dossin, J.R. and V.K. analysed live-cell imaging experiments. V.K. wrote the image analysis software. F. Dossin and S.C. processed and analysed the data. A.L.S. cloned the SPEN cDNA truncation constructs. T.C. helped with some experiments and M.A. derived NPC clones. Y.Z. prepared Hi-C libraries in the laboratory of J.D. F. Dingli carried out the mass spectrometry experimental work and D.L. supervised mass spectrometry experiments and data analysis. F. Dossin and E.H. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 SPEN mediates gene silencing across the entire X chromosome in vitro and in vivo.
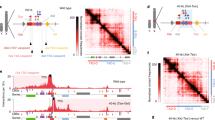
a, Schematic representation of the SPEN-degron genotype with AID-HaloTag insertions in frame with the C terminus of endogenous SPEN. Targeted homozygous insertion of V5-tagged OsTIR1 at the TIGRE locus (top left) results in its constitutive protein expression as assessed by western blot (bottom left). Right, Sanger sequencing results for a PCR amplicon specific to AID-HaloTag insertions and covering a SNP outside of the recombined left homology arm. Detection of both alleles in the amplicon confirms homozygous AID knock-in. b, Fixed-cell imaging of HaloTag in wild-type cells (left), in SPEN-degron mouse ES cells (middle) and in SPEN-degron mouse ES cells exposed to auxin for 4 h (right). Cells were labelled with Halo-JF646 before fixation. SPEN–Halo is properly localized to the nucleus, and is depleted upon auxin treatment. This experiment was repeated at least twice with similar results. c, Bar graph showing the proportion of cells displaying Xist RNA clouds (quantified using RNA FISH) before and after degradation of SPEN (n, number of cells counted; χ2 test). d, Violin plot showing the distribution of X-chromosomal transcript allelic ratios (obtained by RNA-seq) after 0 h DOX, 24 h DOX or 24 h DOX + auxin treatment in wild-type SPEN-degron mouse ES cells. Horizontal lines denote the median, box limits correspond to upper and lower quartiles, averages of two independent clones shown, n = 434 genes, two-sided Student’s t-test. e, RNA FISH experiments for Xist (red) and two X-linked genes: Atrx (grey) and Huwe1 (green), in SPEN-degron mouse ES cells treated with DOX only, or DOX in combination with auxin for 24 h. The proportion of Atrx/Huwe1 monoallelic and biallelic expression among Xist-expressing cells is shown (n, number of cells counted; χ2 test). f, Illustration of the control hybrid mouse crossbreeding scheme for the experiment shown in Fig. 1g, h. g, Quantitative PCR (qPCR) analysis of Spen and Xist transcripts in wild-type (n = 7) and maternal-zygotic Spen-knockout (n = 5) E3.5 embryos. h, Pyrosequencing assay of three X-linked transcripts in maternal-zygotic Spen-knockout (n = 5) and wild-type (n = 7) E3.5 embryos (two-sided Student’s t-test). In g, h, bars show the mean value and individual data points are shown as dots.
Extended Data Fig. 2 SPEN localizes to the X chromosome immediately upon Xist upregulation and throughout the stages of XCI, but is dispensable for maintenance of X-linked gene silencing.
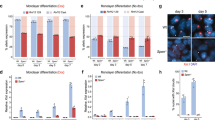
a, Scheme of the strategy for live-cell imaging of SPEN protein and Xist RNA. b, Live-cell snapshot after 16 h of Xist induction in the cell line shown in a. This experiment was repeated at least twice with similar results. c, d, Kinetics of total intensity (c) and area (d) of Xist (red) and SPEN (green) domains over time during Xist induction. The data in c, d are the averages of 27 tracked cells. Error bars indicate standard deviation. Images were acquired every 10 min. Time point 1 is defined as the earliest time at which a SPEN or Xist domain is detected in each cell. Intensity and area values were respectively normalized to the maximum value reached for each signal (SPEN and Xist). e, Hi-C map of the inactive (top) and active (bottom) X chromosomes (resolution, 1.024 Mb) in NPCs after 0 h or 48 h of auxin-mediated SPEN depletion. f, Heat map of the average contact enrichment on scaled topologically associating domains containing escapees in NPCs after 0 h or 48 h of auxin-mediated SPEN depletion. g, Quantification of the allelic ratio (inactive/active X chromosome) of the Hi-C signal within topologically associating domains (n = 37) shown in f, after 0 h or 48 h of auxin-mediated SPEN depletion. Horizontal lines denote the median, box limits correspond to upper and lower quartiles, two-sided Wilcoxon rank-sum test. In e, f, averages of two independent clones are shown.
Extended Data Fig. 3 The SPOC domain of SPEN mediates gene silencing and interacts with multiple molecular pathways.
a, Scheme of complementation strategy. b, Western blot detection of overexpressed 3×Flag-tagged SPEN protein rescue fragments. c, Scheme showing endogenous deletion of SPOC. d, Sub-nuclear localization of endogenous SPEN lacking its SPOC domain upon Xist RNA induction. The inactive X chromosome is identified using immunofluorescence detection of H2AK119ub1. e, Bar graph showing the proportion of cells with Xist RNA clouds (assayed by RNA FISH) in wild-type cells and three independent SPOC-deletion clones after induction of Xist for 24 h (n, number of counted cells). f, RNA FISH for Xist (red) and Huwe1 (green) in SPOC-deletion and wild-type cells treated with DOX for 24 h. g, Violin plot showing the distribution of X-chromosomal transcript allelic ratios (measured by RNA-seq) after 0 h or 24 h DOX treatment in wild-type and SPOC-deletion mouse ES cells. Horizontal lines denote the median, box limits correspond to upper and lower quartiles, averages of three independent clones shown, n = 469 genes, two-sided Student’s t-test. h, Bar graph of transcript allelic ratios (obtained from pyrosequencing) for four X-linked genes in SPOC-deletion (blue) or wild-type (grey) cells. Bars show mean values for three independent SPOC-deletion clones (*P < 10−4, two-sided Student’s t-test). i, Bar graph showing the proportion of cells expressing Huwe1 monoallelically (white) or biallelically (grey), assayed by RNA FISH, in wild-type cells and in three independent SPOC-deletion clones after induction of Xist for 24 h (n, number of counted cells). j, Density plot showing the distribution of gene silencing defects (see Methods) observed across the X chromosome in RNA-seq data from HDAC3-knockout24 SPEN-degron and SPOC-deletion (this study) ES cells after 24 h of Xist induction. k, Bar graph of normalized allelic ratios (obtained from pyrosequencing) for four X-linked genes in HDAC3-knockout (brown), SPOC-deletion (blue) and wild-type (grey) cells after 24 h of Xist induction. Bars show mean values for two independent HDAC3 clones and three independent SPOC deletion clones; individual data points are shown. l, Volcano plot of fold changes in GFP-pull-down (BglG–GFP–SPOC compared with BglG–GFP) and their adjusted P values (Benjamini–Hochberg procedure, see Methods for statistical analysis). Quantitative label-free mass spectrometric analysis was performed on four independent biological replicates. In b, d, f, experiments were repeated at least twice with similar results.
Extended Data Fig. 4 SPEN is recruited by Xist to active gene promoters and enhancers where it silences transcription and subsequently disengages from chromatin.
a, Bar graph showing the number of SPEN peaks on each chromosome after 0 h, 4 h, 8 h and 24 h of Xist induction in mouse ES cells. b, Annotation of SPEN peaks on autosomes. c, Heat map showing allelic ratios at SPEN peaks during XCI among different X-linked genomic features. d, Violin plot showing expression (RPKM) of genes accumulating SPEN (n = 259) or not accumulating SPEN (n = 689) at their promoters. Genes showing 0 RPKM were excluded from this plot. e, Box plots showing SPEN enrichment after 4 h of Xist induction within promoter windows of genes grouped on the basis of their level of dependency on SPEN for gene silencing (see Fig. 1e). f, Box plots showing SPEN enrichment after 4 h of Xist induction within promoter windows of genes grouped on the basis of whether or not they are silenced at 24 h of Xist induction (see Methods). In d–f, data were analysed using the two-sided Wilcoxon rank-sum test, horizontal lines denote the median, box limits correspond to upper and lower quartiles. g, UCSC Genome Browser allele-specific track showing SPEN binding around Kdm6a, an escaping gene (blue, Cast-Xa; red, B6-Xi; all tracks are scaled identically). h, Bar graphs showing overlap between SPEN-binding sites and the binding sites of four different factors at X-linked enhancers and promoters. i, j, Heat maps showing normalized SPEN enrichment (log2) at promoters (both replicates are shown) (i) and gene silencing kinetics (allelic ratio) during XCI (j) within three groups of X-linked genes showing different dynamics of SPEN accumulation and loss. k, Schematic of the function of SPEN in XCI. In a–f, h–j, data are from two biological replicates.
Extended Data Fig. 5 UCSC Genome Browser allelic tracks of SPEN binding and transcript expression at X-linked genes.
a–n, Top, Genome Browser allelic tracks of SPEN binding (from CUT&RUN) at silenced genes (a–g) and non-silenced genes (h–n) during a time course of Xist induction in mouse ES cells (blue, Cast-Xa; red, B6-Xi; scaled identically within each panel). Bottom, allelic tracks of transcript expression (from RNA-seq) at 0 h and 24 h of Xist induction in mouse ES cells (light grey, Cast-Xa; black, B6-Xi; scaled identically within each panel). The relative position of each gene along the X chromosome is shown at the top of the figure.
Supplementary information
Supplementary Figures
Supplementary Figure 1: Uncropped images of Western blot gels.
Supplementary Table
Supplementary Table 1: SPEN is essential for XCI in mouse embryonic stem cells. Allelic ratios for X-linked genes in untreated, dox treated, and dox+auxin treated SPEN-degron mouse embryonic stem cells. Data represents mean values for two independent clones.
Supplementary Table
Supplementary Table 2: SPEN is essential for imprinted XCI in vivo. Allelic ratios for X-linked genes in WT, maternal-only and maternal-zygotic Spen KO E3.5 embryos.
Supplementary Table
Supplementary Table 3: SPEN is dispensable for maintenance of XCI in NPCs. Allelic ratios for X-linked genes in untreated, 24h auxin, and 48h auxin treated SPEN-degron NPCs. Data represents mean values for two independent clones.
Supplementary Table
Supplementary Table 4: List of proteins identified in SPOC-immunoprecipitation followed by mass spectrometry. Data is representative of 4 independent biological replicates, and p-values were adjusted using the Benjamini-Hochberg procedure. See Methods for statistical analysis.
Supplementary Table
Supplementary Table 5: CUT&RUN profiles at promoters. Normalised SPEN enrichment at promoters and corresponding transcript allelic-ratios during Xist induction. Data is representative of 2 independent biological replicates.
Video 1
SPEN is recruited to the X chromosome immediately upon Xist RNA upregulation. Live cell imaging of SPEN-GFP (left panel, green in the right panel) and Xist (middle panel, red in the right panel) in mouse embryonic stem cells during the earliest stage of Xist RNA upregulation. Xist RNA is visualized through expression of a BglG-mCherry fusion protein binding an array of BglSL stem loops on the endogenous Xist RNA. Images were acquired every 10 minutes for more than 4 hours. Scalebar represents 2um. This experiment was repeated independently with more than 20 cells, yielding simila.
Source data
Rights and permissions
About this article
Cite this article
Dossin, F., Pinheiro, I., Żylicz, J.J. et al. SPEN integrates transcriptional and epigenetic control of X-inactivation. Nature 578, 455–460 (2020). https://doi.org/10.1038/s41586-020-1974-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-1974-9
This article is cited by
-
Identification of the RSX interactome in a marsupial shows functional coherence with the Xist interactome during X inactivation
Genome Biology (2024)
-
Transcription regulation by long non-coding RNAs: mechanisms and disease relevance
Nature Reviews Molecular Cell Biology (2024)
-
Unraveling the role of Xist in X chromosome inactivation: insights from rabbit model and deletion analysis of exons and repeat A
Cellular and Molecular Life Sciences (2024)
-
The chromatin-associated RNAs in gene regulation and cancer
Molecular Cancer (2023)
-
RNA polymerase II depletion from the inactive X chromosome territory is not mediated by physical compartmentalization
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.