Abstract
On infection of their host, temperate viruses that infect bacteria (bacteriophages; hereafter referred to as phages) enter either a lytic or a lysogenic cycle. The former results in lysis of bacterial cells and phage release (resulting in horizontal transmission), whereas lysogeny is characterized by the integration of the phage into the host genome, and dormancy (resulting in vertical transmission)1. Previous co-culture experiments using bacteria and mutants of temperate phages that are locked in the lytic cycle have shown that CRISPR–Cas systems can efficiently eliminate the invading phages2,3. Here we show that, when challenged with wild-type temperate phages (which can become lysogenic), type I CRISPR–Cas immune systems cannot eliminate the phages from the bacterial population. Furthermore, our data suggest that, in this context, CRISPR–Cas immune systems are maladaptive to the host, owing to the severe immunopathological effects that are brought about by imperfect matching of spacers to the integrated phage sequences (prophages). These fitness costs drive the loss of CRISPR–Cas from bacterial populations, unless the phage carries anti-CRISPR (acr) genes that suppress the immune system of the host. Using bioinformatics, we show that this imperfect targeting is likely to occur frequently in nature. These findings help to explain the patchy distribution of CRISPR–Cas immune systems within and between bacterial species, and highlight the strong selective benefits of phage-encoded acr genes for both the phage and the host under these circumstances.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Source Data associated with Figs. 1–4 and Extended Data Figs. 1–3, 5, 7–9 are provided with the paper. Sequencing data have been deposited in the European Nucleotide Archive under the study accession number PRJEB34503. The datasets analysed for the bioinformatic study are available on GitHub at https://github.com/davidchyou/Rollie-Chevallereau.
Code availability
Mathematical algorithms generated during this study are available in the Supplementary Information. Scripts generated for the bioinformatics analyses are available on GitHub at https://github.com/davidchyou/Rollie-Chevallereau.
Change history
03 March 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41586-020-2089-z
References
Stewart, F. M. & Levin, B. R. The population biology of bacterial viruses: why be temperate. Theor. Popul. Biol. 26, 93–117 (1984).
Westra, E. R. et al. Parasite exposure drives selective evolution of constitutive versus inducible defense. Curr. Biol. 25, 1043–1049 (2015).
van Houte, S. et al. The diversity-generating benefits of a prokaryotic adaptive immune system. Nature 532, 385–388 (2016).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Datsenko, K. A. et al. Molecular memory of prior infections activates the CRISPR/Cas adaptive bacterial immunity system. Nat. Commun. 3, 945 (2012).
Fineran, P. C. et al. Degenerate target sites mediate rapid primed CRISPR adaptation. Proc. Natl Acad. Sci. USA 111, E1629–E1638 (2014).
Howard-Varona, C., Hargreaves, K. R., Abedon, S. T. & Sullivan, M. B. Lysogeny in nature: mechanisms, impact and ecology of temperate phages. ISME J. 11, 1511–1520 (2017).
Heussler, G. E. et al. Clustered regularly interspaced short palindromic repeat-dependent, biofilm-specific death of Pseudomonas aeruginosa mediated by increased expression of phage-related genes. mBio 6, e00129-15 (2015).
Zegans, M. E. et al. Interaction between bacteriophage DMS3 and host CRISPR region inhibits group behaviors of Pseudomonas aeruginosa. J. Bacteriol. 191, 210–219 (2009).
Berngruber, T. W., Froissart, R., Choisy, M. & Gandon, S. Evolution of virulence in emerging epidemics. PLoS Pathog. 9, e1003209 (2013).
Trasanidou, D. et al. Keeping CRISPR in check: diverse mechanisms of phage-encoded anti-CRISPRS. FEMS Microbiol. Lett. 366, fnz098 (2019).
Bondy-Denomy, J. et al. Multiple mechanisms for CRISPR–Cas inhibition by anti-CRISPR proteins. Nature 526, 136–139 (2015).
Little, J. W. & Michalowski, C. B. Stability and instability in the lysogenic state of phage lambda. J. Bacteriol. 192, 6064–6076 (2010).
Goldberg, G. W., Jiang, W., Bikard, D. & Marraffini, L. A. Conditional tolerance of temperate phages via transcription-dependent CRISPR–Cas targeting. Nature 514, 633–637 (2014).
Samai, P. et al. Co-transcriptional DNA and RNA cleavage during type III CRISPR–Cas immunity. Cell 161, 1164–1174 (2015).
Landsberger, M. et al. Anti-CRISPR phages cooperate to overcome CRISPR–Cas immunity. Cell 174, 908–916 (2018).
Borges, A. L. et al. Bacteriophage cooperation suppresses CRISPR–Cas3 and Cas9 immunity. Cell 174, 917–925 (2018).
Poulsen, B. E. et al. Defining the core essential genome of Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 116, 10072–10080 (2019).
Mooij, M. J. et al. Characterization of the integrated filamentous phage Pf5 and its involvement in small-colony formation. Microbiology 153, 1790–1798 (2007).
Zhou, Y., Liang, Y., Lynch, K. H., Dennis, J. J. & Wishart, D. S. PHAST: a fast phage search tool. Nucleic Acids Res. 39, W347–W52 (2011).
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015).
Levy, A. et al. CRISPR adaptation biases explain preference for acquisition of foreign DNA. Nature 520, 505–510 (2015).
Stern, A., Keren, L., Wurtzel, O., Amitai, G. & Sorek, R. Self-targeting by CRISPR: gene regulation or autoimmunity? Trends Genet. 26, 335–340 (2010).
Vercoe, R. B. et al. Cytotoxic chromosomal targeting by CRISPR/Cas systems can reshape bacterial genomes and expel or remodel pathogenicity islands. PLoS Genet. 9, e1003454 (2013).
Jiang, W. et al. Dealing with the evolutionary downside of CRISPR immunity: bacteria and beneficial plasmids. PLoS Genet. 9, e1003844 (2013).
Goldberg, G. W. et al. Incomplete prophage tolerance by type III-A CRISPR–Cas systems reduces the fitness of lysogenic hosts. Nat. Commun. 9, 61 (2018).
Cady, K. C. & O’Toole, G. A. Non-identity-mediated CRISPR-bacteriophage interaction mediated via the Csy and Cas3 proteins. J. Bacteriol. 193, 3433–3445 (2011).
Liberati, N. T. et al. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc. Natl Acad. Sci. USA 103, 2833–2838 (2006).
Cady, K. C., Bondy-Denomy, J., Heussler, G. E., Davidson, A. R. & O’Toole, G. A. The CRISPR/Cas adaptive immune system of Pseudomonas aeruginosa mediates resistance to naturally occurring and engineered phages. J. Bacteriol. 194, 5728–5738 (2012).
Ceyssens, P.-J. et al. Comparative analysis of the widespread and conserved PB1-like viruses infecting Pseudomonas aeruginosa. Environ. Microbiol. 11, 2874–2883 (2009).
Chevallereau, A. et al. Exploitation of the cooperative behaviors of anti-CRISPR phages. Cell Host Microbe 27, 1–10 (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Arndt, D. et al. PHASTER: a better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44, W16– W21 (2016).
Acknowledgements
We thank A. R. Davidson for providing the mutant strains of PA14 ∆cas3, ∆cas7, ∆cas1 and ∆CRISPR2, G. A. O’Toole for the strain CRISPR2 ∆sp1–2 and J. Bondy-Denomy for the Tn::pilA (∆pilA) PA14 surface mutant. Genome sequencing was provided by MicrobesNG (http://www.microbesng.uk), which is supported by the BBSRC (grant number BB/L024209/1). This work was funded by a grant from the European Research Council (https://erc.europa.eu) (ERC-STG-2016-714478 - EVOIMMECH) and NERC Independent Research Fellowship (NE/M018350/1) awarded to E.R.W. A.C. has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No 834052. P.C.F. was supported by the Marsden Fund from the Royal Society of New Zealand.
Author information
Authors and Affiliations
Contributions
Conceptualization of the study was done by C.R., A.C. and E.R.W. Experimental design was carried out by C.R., A.C., B.N.J.W and E.R.W. Bacterial evolution, competition and growth experiments were done by C.R. and A.C. with assistance from O.F. and I.M. Virulent versus temperate phage competitions and prophage induction-rate experiments were performed by C.R. All experiments with acr phages and CRISPR 2 spacer 1 were done by A.C. B.N.J.W. and A.C. carried out Pf5 experiments. The experiment with superinfecting virulent phage was done by E.R.W. C.R., A.C., B.N.J.W., T.-y.C., C.M.B., P.C.F. and E.R.W. analysed the data. S.G. generated theoretical mathematical models. T.-y.C. conducted bioinformatic analyses supervised by C.M.B. and P.C.F. A.C. performed whole-genome sequencing analyses. C.R. wrote the original draft of the manuscript; A.C. wrote the revised version of the manuscript with contributions from C.R., B.N.J.W., T.-y.C., S.G., C.M.B. and P.C.F. E.R.W. supervised the project and provided comments on all versions of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Eugene Koonin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Infection with a 50:50 mix of temperate:virulent phages.
a, b, Bacterial (a) and phage (b) titres during a co-culture experiment of wild-type PA14 (red) or ∆cas7 mutant (blue), and a 50:50 mix of DMS3 and DMS3vir. c, d, Resistance phenotypes at day 3 (c) or day 7 (d) of the co-culture experiment, based on 24 random clones per replicate experiment. Data are the mean of six biological replicates per treatment. Error bars represent 95% confidence intervals.
Extended Data Fig. 2 The suppression of lysogeny and immunopathological effects are due to spacer 1 of CRISPR array 2.
a, b, Phage (a) and bacterial (b) titres during co-culture of phage DMS3 and P. aeruginosa PA14 ΔCRISPR2, expressing a non-targeting spacer from a plasmid (ΔCRISPR2-NT) or the original CRISPR2 spacer 1 (ΔCRISPR2-sp1). c, d,The proportion of lysogens (c) and the frequency of loss of CRISPR–Cas immune systems (d) at 1 and 3 days post-infection, based on PCR analyses of 24 random clones per replicate experiment. a–d, Data are the mean of three biological replicates. Error bars represent 95% confidence intervals. e–g, Growth of three independent lysogen clones isolated at three days post-infection, as determined by OD600 nm measurements. ΔCRISPR2-NT (e) and ΔCRISPR2-sp1 (f) lysogen clones carry the ancestral ΔCRISPR2 CRISPR–Cas immune system, whereas the ΔCRISPR2-sp1 (g) lysogen clones have evolved to lose CRISPR–Cas.
Extended Data Fig. 3 Prophage induction rates are increased in hosts with active CRISPR–Cas.
a, The percentage of lysogens formed upon infection of wild-type host with DMS3 phages engineered to produce AcrIF1 or AcrIF4 anti-CRISPR proteins. b, c, Optical density (b) and phage titres (c) during growth of lysogens of DMS3, DMS3 acrIFI or DMS3 acrIF4 phages in a wild-type PA14 or ∆cas7 genetic background. d, Relative fitness of the DMS3 phage during competition with the virulent mutant DMS3vir in the presence of varying fractions of sensitive (∆cas7) host and resistant hosts with CRISPR-based immunity (BIM) or surface-based immunity (sm2) against these phages. Data show mean fitness at 8 h after infection. All panels show the mean of six biological replicates and error bars represent 95% confidence intervals.
Extended Data Fig. 4 Lysogens lose their CRISPR–Cas immune systems.
a, PCR amplification of the c-repressor gene of the prophage (c-rep, 611 bp), the fimV gene (located about 1 Mb from the CRISPR loci and used as a positive control for the PCR, 116 bp) and CRISPR loci 1 (349 bp) and 2 (206 bp) on the host genome. PCRs were performed on 6 independent DMS3 lysogens in wild-type, Δcas1 and Δcas7 backgrounds isolated at 1 or 7 days post-infection, as well as on 6 independent lysogens of DMS3, DMS3 acrIF1 or DMS3 acrIF4 (wild-type background) isolated at 6 or 120 h after infection. Red frames indicate a failure to amplify a product. PCR amplifications were performed on clones isolated from three biological replicate experiments and produced similar results. For gel source data, see Supplementary Fig. 1. b, Schematic of the CRISPR–Cas locus of wild-type PA14, which spans a region of around 11 kb. Primers used to amplify regions of CRISPR arrays 1 or 2 are shown as red arrows. c–e, Whole-genome sequencing of DMS3 lysogens that lost their CRISPR–Cas system (red frames in a) in wild-type PA14 (c), ∆cas1 (d) or ∆cas7 (e) backgrounds. Graphs show the read coverage of the region encompassing positions 2.70–2.97 Mb of the wild-type PA14 genome. The CRISPR–Cas locus is indicated by a green box on the x axis. A genome map depicting coding sequences (yellow arrows) is shown above the graphs. The region comprising 2.84–2.88 Mb includes sequences that are repeated elsewhere on the PA14 genome, explaining why reads that map to these positions are still detected in some of the deletion mutants. The high peak at the 3′ end of the CRISPR locus corresponds to the coverage of spacer 20 of CRISPR2 by reads that derive from DMS3 prophage (5′ and 3′ extremities of these reads map to the phage genome). Spacer 20 of CRISPR2 has 100% identity to DMS3 but is not immunogenic because there is no consensus protospacer-adjacent motif.
Extended Data Fig. 5 Expression of Pf5 priming spacer in P. aeruginosa PA14.
a, Growth of ∆cas7 (dashed line) or wild-type (solid line) clones carrying an expression plasmid encoding a non-targeting spacer (pNT) or a spacer targeting the PA14 natural prophage Pf5 with one mismatch (pPf5-MS), as determined by OD600 nm measurements. Graphs show mean curves from 6 biological replicates, and shaded areas correspond to 95% confidence intervals. b, Relative fitness of wild-type pNT or wild-type pPf5-MS during competition with ∆cas7 pNT. Data are the mean of six biological replicates per treatment. Error bars represent 95% confidence intervals.
Extended Data Fig. 6 Simulations of population and evolutionary dynamics of bacteria–phage interactions, when virulent and temperate phages compete on bacteria with a CRISPR–Cas system.
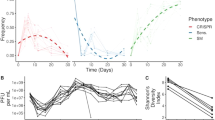
a–c, e–g, Graphs show densities of susceptible hosts, CRISPR-resistant bacteria and lysogens (a, e) or free viruses over time (b, f), as well as the proportion of temperate phages in a population composed of both temperate and virulent types (c, g). Temperate phages can transmit both horizontally and vertically, whereas virulent phages can transmit only horizontally and cannot superinfect lysogens. d, h, Frequency of evolutionary loss of CRISPR–Cas system in the lysogen population over time. The simulations shown in a–d reflect a situation in which both virulent and temperate phages lack acr genes, whereas those in e–h reflect a scenario in which the temperate type carries an acr gene.
Extended Data Fig. 7 Matches between spacers and temperate phages are widespread.
a, Total matches between non-redundant spacers (n = 1,239,973) from 171,361 RefSeq and GenBank complete genomes and a non-redundant set of temperate phages (n = 19,996)21. The counts of perfect (0) or mismatched (1–5) targets are shown. As a control, the temperate phages were shuffled ten times, while retaining the hexanucleotide content (control). b, Counts of spacers matching temperate phages from all genera with over 500 spacer–prophage matches. The total number (n) of spacer–prophage matches is shown for each genus in parentheses. Counts of matches are shown (0, green; 1–5 mismatches, red). The number of temperate phages analysed is plotted (prophages in purple) as are the matches to shuffled prophages. The control is shown in blue, but is not visible because it had only 0 to 10 counts. c, The percentage of prophages within each genus that were targeted by self-priming spacers (1–5 mismatches). d, Heat map of the distribution of mismatches (0–5). Genera are as in b and data are shown as log(count) for each genus, as the number of matches varied widely between genera.
Extended Data Fig. 8 Self-targeting genomes are enriched for acr gene(s).
a, b, The number of P. aeruginosa genomes with complete CRISPR–Cas systems that contain (+) or lack (−) genes encoding known Acr proteins. For these strains, the total number of strains with perfect (0) or mismatched (1–5) self-targeting (ST) spacers to anywhere in the genome (a) or to prophages (b) are shown. For complete P. aeruginosa genomes, all self-targeting events were analysed for matches to prophages using PHASTER34. The number of genomes with acr genes (acr +) and self-targeting (ST +) spacers is significantly greater than the number of genomes with acr genes and without self-targeting spacers (P = 8.14 × 10−5, two-sided Fisher’s exact test, n = 71).
Extended Data Fig. 9 Presence of a superinfecting virulent phage does not alter immunopathological effects.
a–c, Bacterial (a) and phage titres upon individual (b) or mixed (c) infection of wild-type PA14 with phage DMS3 and virulent phage LMA2. d, e, Resistance phenotypes evolved by bacteria against DMS3 upon individual (d) or mixed (e) infection. f, Frequency of loss of CRISPR–Cas immune systems upon infection with phage DMS3 or with both the phages DMS3 and LMA2, based on 24 random clones per replicate experiment. g, Relative fitness of wild-type PA14 during competition with PA14 Δcas7 in the presence or absence of phages DMS3 and LMA2. a–g, Data are the means of six biological replicates. Error bars indicate 95% confidence intervals. h–o, Simulations of population and evolutionary dynamics during infection of bacteria carrying CRISPR–Cas systems with a mixed population of unrelated virulent and temperate phages. Graphs show densities of susceptible hosts, CRISPR-resistant bacteria and lysogens (h, i) and free viruses over time (j, k), as well as the frequencies of temperate phages in a population composed of both temperate and virulent types (l, m). Temperate phage can transmit both horizontally and vertically, whereas virulent phage can transmit only horizontally and can superinfect the lysogens (because temperate and virulent phages are unrelated). n, o, Frequencies of evolutionary loss of CRISPR–Cas system in the lysogen population over time. The simulations shown in h, j, l, n reflect a scenario in which bacteria can evolve CRISPR-based resistance against both phages, whereas those shown in i, k, m, o reflect a situation in which CRISPR-based resistance does not evolve against the virulent phage, and bacteria instead evolve costly surface-based resistance (as it is the case in our experiments). A detailed description of the simulations is provided in the Supplementary Information.
Supplementary information
Supplementary Information
These files contain Supplementary Methods: Description of epidemiological modelling of phage dynamics (mathematical algorithms) and of bioinformatic analysis of widespread priming off temperate phages. Supplementary Table 1: Parameters of the mathematical model with default values. Supplementary Figure 1: Source data images for PCR amplification.
Source data
Rights and permissions
About this article
Cite this article
Rollie, C., Chevallereau, A., Watson, B.N.J. et al. Targeting of temperate phages drives loss of type I CRISPR–Cas systems. Nature 578, 149–153 (2020). https://doi.org/10.1038/s41586-020-1936-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-1936-2
This article is cited by
-
Arbitrium communication controls phage lysogeny through non-lethal modulation of a host toxin–antitoxin defence system
Nature Microbiology (2024)
-
Slow growing bacteria survive bacteriophage in isolation
ISME Communications (2023)
-
Analysis of CRISPR-Cas Loci and their Targets in Levilactobacillus brevis
Interdisciplinary Sciences: Computational Life Sciences (2023)
-
Interactions between bacterial and phage communities in natural environments
Nature Reviews Microbiology (2022)
-
High-pressure processing-induced transcriptome response during recovery of Listeria monocytogenes
BMC Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.