Abstract
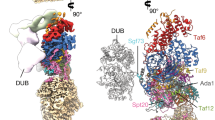
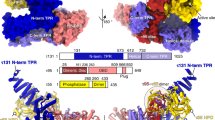
Gene transcription by RNA polymerase II is regulated by activator proteins that recruit the coactivator complexes SAGA (Spt–Ada–Gcn5–acetyltransferase)1,2 and transcription factor IID (TFIID)2,3,4. SAGA is required for all regulated transcription5 and is conserved among eukaryotes6. SAGA contains four modules7,8,9: the activator-binding Tra1 module, the core module, the histone acetyltransferase (HAT) module and the histone deubiquitination (DUB) module. Previous studies provided partial structures10,11,12,13,14, but the structure of the central core module is unknown. Here we present the cryo-electron microscopy structure of SAGA from the yeast Saccharomyces cerevisiae and resolve the core module at 3.3 Å resolution. The core module consists of subunits Taf5, Sgf73 and Spt20, and a histone octamer-like fold. The octamer-like fold comprises the heterodimers Taf6–Taf9, Taf10–Spt7 and Taf12–Ada1, and two histone-fold domains in Spt3. Spt3 and the adjacent subunit Spt8 interact with the TATA box-binding protein (TBP)2,7,15,16,17. The octamer-like fold and its TBP-interacting region are similar in TFIID, whereas Taf5 and the Taf6 HEAT domain adopt distinct conformations. Taf12 and Spt20 form flexible connections to the Tra1 module, whereas Sgf73 tethers the DUB module. Binding of a nucleosome to SAGA displaces the HAT and DUB modules from the core-module surface, allowing the DUB module to bind one face of an ubiquitinated nucleosome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The electron density reconstructions and models of the complete SAGA complex, the Tra1 module, the core module, the DUB module–nucleosome complex and the nucleosome-bound state of SAGA were deposited with the Electron Microscopy Data Bank (accession codes EMD-10412, EMD-10413, EMD-10414, EMD-10415 and EMD-10416 respectively) and with the Protein Data Bank (accession codes 6T9I, 6T9J, 6T9K, and 6T9L). All the other relevant data are included in the Supplementary Information or are available from the authors upon request.
References
Grant, P. A. et al. Yeast Gcn5 functions in two multisubunit complexes to acetylate nucleosomal histones: characterization of an Ada complex and the SAGA (Spt/Ada) complex. Genes Dev. 11, 1640–1650 (1997).
Hahn, S. & Young, E. T. Transcriptional regulation in Saccharomyces cerevisiae: transcription factor regulation and function, mechanisms of initiation, and roles of activators and coactivators. Genetics 189, 705–736 (2011).
Patel, A. B. et al. Structure of human TFIID and mechanism of TBP loading onto promoter DNA. Science 362, eaau8872 (2018).
Kolesnikova, O. et al. Molecular structure of promoter-bound yeast TFIID. Nat. Commun. 9, 4666 (2018).
Baptista, T. et al. SAGA is a general cofactor for RNA polymerase II transcription. Mol. Cell 70, 1163–1164 (2018).
Spedale, G., Timmers, H. T. & Pijnappel, W. W. ATAC-king the complexity of SAGA during evolution. Genes Dev. 26, 527–541 (2012).
Han, Y., Luo, J., Ranish, J. & Hahn, S. Architecture of the Saccharomyces cerevisiae SAGA transcription coactivator complex. EMBO J. 33, 2534–2546 (2014).
Sharov, G. et al. Structure of the transcription activator target Tra1 within the chromatin modifying complex SAGA. Nat. Commun. 8, 1556 (2017).
Liu, G. et al. Architecture of Saccharomyces cerevisiae SAGA complex. Cell Discov. 5, 25 (2019).
Köhler, A., Zimmerman, E., Schneider, M., Hurt, E. & Zheng, N. Structural basis for assembly and activation of the heterotetrameric SAGA histone H2B deubiquitinase module. Cell 141, 606–617 (2010).
Samara, N. L. et al. Structural insights into the assembly and function of the SAGA deubiquitinating module. Science 328, 1025–1029 (2010).
Morgan, M. T. et al. Structural basis for histone H2B deubiquitination by the SAGA DUB module. Science 351, 725–728 (2016).
Díaz-Santín, L. M., Lukoyanova, N., Aciyan, E. & Cheung, A. C. Cryo-EM structure of the SAGA and NuA4 coactivator subunit Tra1 at 3.7 angstrom resolution. eLife 6, e28384 (2017).
Sun, J. et al. Structural basis for activation of SAGA histone acetyltransferase Gcn5 by partner subunit Ada2. Proc. Natl Acad. Sci. USA 115, 10010–10015 (2018).
Belotserkovskaya, R. et al. Inhibition of TATA-binding protein function by SAGA subunits Spt3 and Spt8 at Gcn4-activated promoters. Mol. Cell. Biol. 20, 634–647 (2000).
Warfield, L., Ranish, J. A. & Hahn, S. Positive and negative functions of the SAGA complex mediated through interaction of Spt8 with TBP and the N-terminal domain of TFIIA. Genes Dev. 18, 1022–1034 (2004).
Sermwittayawong, D. & Tan, S. SAGA binds TBP via its Spt8 subunit in competition with DNA: implications for TBP recruitment. EMBO J. 25, 3791–3800 (2006).
Helmlinger, D. & Tora, L. Sharing the SAGA. Trends Biochem. Sci. 42, 850–861 (2017).
Larschan, E. & Winston, F. The S. cerevisiae SAGA complex functions in vivo as a coactivator for transcriptional activation by Gal4. Genes Dev. 15, 1946–1956 (2001).
Brown, C. E. et al. Recruitment of HAT complexes by direct activator interactions with the ATM-related Tra1 subunit. Science 292, 2333–2337 (2001).
Dudley, A. M., Rougeulle, C. & Winston, F. The Spt components of SAGA facilitate TBP binding to a promoter at a post-activator-binding step in vivo. Genes Dev. 13, 2940–2945 (1999).
Daniel, J. A. et al. Deubiquitination of histone H2B by a yeast acetyltransferase complex regulates transcription. J. Biol. Chem. 279, 1867–1871 (2004).
Henry, K. W. et al. Transcriptional activation via sequential histone H2B ubiquitylation and deubiquitylation, mediated by SAGA-associated Ubp8. Genes Dev. 17, 2648–2663 (2003).
Wu, P. Y., Ruhlmann, C., Winston, F. & Schultz, P. Molecular architecture of the S. cerevisiae SAGA complex. Mol. Cell 15, 199–208 (2004).
Gangloff, Y. G. et al. The human TFIID components TAFII135 and TAFII20 and the yeast SAGA components ADA1 and TAFII68 heterodimerize to form histone-like pairs. Mol. Cell. Biol. 20, 340–351 (2000).
Birck, C. et al. Human TAFII28 and TAFII18 interact through a histone fold encoded by atypical evolutionary conserved motifs also found in the SPT3 family. Cell 94, 239–249 (1998).
Eisenmann, D. M., Arndt, K. M., Ricupero, S. L., Rooney, J. W. & Winston, F. SPT3 interacts with TFIID to allow normal transcription in Saccharomyces cerevisiae. Genes Dev. 6, 1319–1331 (1992).
Wu, P. Y. & Winston, F. Analysis of Spt7 function in the Saccharomyces cerevisiae SAGA coactivator complex. Mol. Cell. Biol. 22, 5367–5379 (2002).
Kamata, K. et al. C-terminus of the Sgf73 subunit of SAGA and SLIK is important for retention in the larger complex and for heterochromatin boundary function. Genes Cells 18, 823–837 (2013).
Sterner, D. E., Belotserkovskaya, R. & Berger, S. L. SALSA, a variant of yeast SAGA, contains truncated Spt7, which correlates with activated transcription. Proc. Natl Acad. Sci. USA 99, 11622–11627 (2002).
Pray-Grant, M. G. et al. The novel SLIK histone acetyltransferase complex functions in the yeast retrograde response pathway. Mol. Cell. Biol. 22, 8774–8786 (2002).
Bian, C. et al. Sgf29 binds histone H3K4me2/3 and is required for SAGA complex recruitment and histone H3 acetylation. EMBO J. 30, 2829–2842 (2011).
Simon, M. D. et al. The site-specific installation of methyl-lysine analogs into recombinant histones. Cell 128, 1003–1012 (2007).
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol. 375, 23–44 (2004).
Tegunov, D. & Cramer, P. Real-time cryo-electron microscopy data preprocessing with Warp. Nat. Methods 16, 1146–1152 (2019).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Yang, B. et al. Identification of cross-linked peptides from complex samples. Nat. Methods 9, 904–906 (2012).
Chen, Z. L. et al. A high-speed search engine pLink 2 with systematic evaluation for proteome-scale identification of cross-linked peptides. Nat. Commun. 10, 3404 (2019).
Combe, C. W., Fischer, L. & Rappsilber, J. xiNET: cross-link network maps with residue resolution. Mol. Cell. Proteomics 14, 1137–1147 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Yang, J. et al. The I-TASSER suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Elías-Villalobos, A. et al. Chaperone-mediated ordered assembly of the SAGA and NuA4 transcription co-activator complexes in yeast. Nat. Commun. 10, 5237 (2019).
Acknowledgements
We thank M. Ninov for help with mass spectrometry and T. Schulz for yeast fermentation. H.W. was supported by an EMBO long-term fellowship (ALTF 650-2017). H.U. was supported by the Deutsche Forschungsgemeinschaft (SFB860). A.C.M.C. was supported by Wellcome (102535/Z/13/Z). P.C. was supported by the Deutsche Forschungsgemeinschaft (SFB860, SPP1935, EXC 2067/1-390729940), the European Research Council (advanced investigator grant TRANSREGULON, grant agreement no. 693023) and the Volkswagen Foundation.
Author information
Authors and Affiliations
Contributions
H.W. carried out all experiments and data analysis except mass spectrometry analysis. C.D. assisted with cryo-EM data collection. A.C.M.C. contributed to developing the purification protocol and assisted with model building. A.S. and H.U. carried out mass spectrometry analysis. P.C. supervised research. H.W. and P.C. interpreted the data and wrote the manuscript, with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Steve Hahn and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cryo-EM structure determination and analysis of SAGA.
Related to data shown in Fig. 1. a, Purification of endogenous SAGA from S. cerevisiae. SDS–PAGE of peak fraction used for cryo-EM grid preparation. Identity of the bands was confirmed by mass spectrometry. For gel source data, see Supplementary Fig. 1. b, Exemplary cryo-EM micrograph of data collection. The micrograph is shown before (left) and after (right) denoising using Warp35. c, The 2D class averages. d, Sorting and classification tree used to reconstruct SAGA. e, FSC between half maps of the final reconstructions of the complete SAGA complex and the SAGA modules Tra1 and core. Resolutions for the gold-standard FSC 0.143 criterion are listed. f, Angular distribution plot for all particles in the final reconstructions of the SAGA core (top) and Tra1 (bottom) modules. Colour shading from blue to yellow correlates with the number of particles at a specific orientation as indicated.
Extended Data Fig. 2 Quality of the SAGA structure.
Relates to data in Figs. 1, 2. a, SAGA reconstruction coloured according to local resolution43. Model–map FSC curves calculated between the refined atomic models and maps are shown below. b, Electron density (grey transparent surface) for various SAGA regions as indicated. c, Overview of the cross-linking data. Circular plot of high-confidence lysine–lysine intersubunit (green) and intrasubunit (purple) cross-links obtained by mass spectrometry for the SAGA complex. The mass spectrometry measurement was repeated twice independently with similar results. Totals of 396 unique intersubunit cross-links and 514 intrasubunit cross-links were obtained. d, Validated cross-links mapped onto the SAGA structure. Out of 396 unique intersubunit cross-links, 120 could be mapped onto the core-module structure, and 109 were located within the 30 Å distance limit for the BS3 cross-linker. Blue lines depict the cross-links with cross-linked sites within the 30 Å distance permitted by BS3, whereas red lines depict cross-links over more than 30 Å.
Extended Data Fig. 3 Comparison of the histone-like fold in SAGA with the histone octamer, details of Taf5–Spt20 interactions, and model of the SAGA–TBP complex.
Relates to data in Figs. 1–3. a, Comparison of the SAGA core module histone octamer-like structure with the canonical histone octamer core (PDB: 1AOI). The canonical octamer core is rendered as the colour for the SAGA octamer-like fold. b, Details of Taf5–Spt20 wedge interactions. Residues involved in the interactions are shown in sticks and coloured as indicated. c, Details of interactions between the Taf5 LisH domain and Spt20 SEP domain. Residues involved in the interactions are shown in sticks and coloured as indicated. d, Model of the SAGA–TBP complex. The model was generated by superposing the TBP-containing TFIID lobe A onto the SAGA core structure. A homology model for Spt8 was generated by the I-TASSER server44.
Extended Data Fig. 4 Details of intermodule interactions.
Relates to data in Figs. 1, 2. a, Binding interface between core and Tra1 modules. The Tra1 FAT domain (grey) is shown as a surface representation. The TIRs of Taf12 (green) and Spt20 (yellow) are shown in cartoon representation. b, Details of the interactions depicted in a. c, Sgf73 (turquoise) tethers the DUB module to the core module. Residues involved in the interactions are shown in sticks and coloured as indicated. d, Sequence alignment of SAGA subunit regions involved in intermodule interactions. Conserved residues are highlighted in blue. Key residues are labelled with asterisks45. Sc, S. cerevisiae; Pp, Pichia pastoris; Sp, Schizosaccharomyces pombe.
Extended Data Fig. 5 Cryo-EM structure determination and analysis of the SAGA–nucleosome complex.
Relates to data in Fig. 4. a, Exemplary cryo-EM micrograph of data collection. The micrograph is shown before (left) and after (right) denoising using Warp35. b, The 2D class averages for the SAGA–nucleosome complex. c, The 2D class averages for the DUB module–nucleosome subcomplex. d, Sorting and classification tree used to reconstruct the DUB module–nucleosome complex at 3.7 Å resolution. e, FSC between half maps of the final reconstructions of the SAGA module, Tra1 and the DUB module–nucleosome complex from SAGA–nucleosome complex data. Resolutions for the gold-standard FSC 0.143 criterion are listed. f, Angular distribution plot for all particles in the final reconstruction of the SAGA DUB module–nucleosome complex. Colour shading from blue to yellow correlates with the number of particles at a specific orientation as indicated. g, Superposition of the crystal structure of DUB-ubiquitinated nucleosome (4ZUX)12 onto the cryo-EM structure presented here. Structures are shown in cartoon and coloured as indicated. h, Comparison of the low-pass-filtered overall cryo-EM maps of SAGA and the SAGA–nucleosome complex. Densities for the HAT and DUB modules are lost on nucleosome binding to SAGA.
Supplementary information
Supplementary Figures
Supplementary Figure 1: This file contains an uncropped scan with size marker indication (cf. Extended Data Fig. 1a).
Supplementary Table
List of BS3 crosslinks within SAGA: List of intra- and inter-subunit lysine-lysine crosslinks as identified by LC-MS analyses and subsequent database search using pLink 138 and pLink 239. The respective scores of cross-link identification are listed as well as the number of CSMs (cross-linked spectra matches).
Video 1
Overview of SAGA structure The video shows a vertical rotation of the SAGA complex structure. It first depicts the overall shape of SAGA based on a low pass-filtered cryo-EM density map. It then shows the high-resolution cryo-EM reconstructions for the Tra1 and core modules, respectively. Finally, it shows the Tra1, core, and DUB module structures fitted to the densities.
Rights and permissions
About this article
Cite this article
Wang, H., Dienemann, C., Stützer, A. et al. Structure of the transcription coactivator SAGA. Nature 577, 717–720 (2020). https://doi.org/10.1038/s41586-020-1933-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-1933-5
This article is cited by
-
The SAGA histone acetyltransferase module targets SMC5/6 to specific genes
Epigenetics & Chromatin (2023)
-
The NuA4 histone acetyltransferase: variations on a theme of SAGA
Nature Structural & Molecular Biology (2023)
-
Regulation of the RNA polymerase II pre-initiation complex by its associated coactivators
Nature Reviews Genetics (2023)
-
SGF29 nuclear condensates reinforce cellular aging
Cell Discovery (2023)
-
The structure of the NuA4–Tip60 complex reveals the mechanism and importance of long-range chromatin modification
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.