Abstract
Some patients infected with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) develop severe pneumonia and acute respiratory distress syndrome1 (ARDS). Distinct clinical features in these patients have led to speculation that the immune response to virus in the SARS-CoV-2-infected alveolus differs from that in other types of pneumonia2. Here we investigate SARS-CoV-2 pathobiology by characterizing the immune response in the alveoli of patients infected with the virus. We collected bronchoalveolar lavage fluid samples from 88 patients with SARS-CoV-2-induced respiratory failure and 211 patients with known or suspected pneumonia from other pathogens, and analysed them using flow cytometry and bulk transcriptomic profiling. We performed single-cell RNA sequencing on 10 bronchoalveolar lavage fluid samples collected from patients with severe coronavirus disease 2019 (COVID-19) within 48 h of intubation. In the majority of patients with SARS-CoV-2 infection, the alveolar space was persistently enriched in T cells and monocytes. Bulk and single-cell transcriptomic profiling suggested that SARS-CoV-2 infects alveolar macrophages, which in turn respond by producing T cell chemoattractants. These T cells produce interferon-γ to induce inflammatory cytokine release from alveolar macrophages and further promote T cell activation. Collectively, our results suggest that SARS-CoV-2 causes a slowly unfolding, spatially limited alveolitis in which alveolar macrophages containing SARS-CoV-2 and T cells form a positive feedback loop that drives persistent alveolar inflammation.
Similar content being viewed by others
Main
A minority of patients infected with SARS-CoV-2 develop severe pneumonia that requires mechanical ventilation, and these patients account for almost all of the morbidity and mortality associated with the infection1,3,4,5. The reported 20–40% mortality among patients with severe SARS-CoV-2 pneumonia, combined with a severe systemic inflammatory response in some patients, have led to speculation that the pathobiology of SARS-CoV-2 pneumonia is distinct from pneumonia caused by other respiratory viral and bacterial pathogens2.
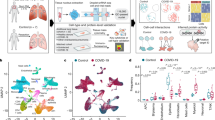
We obtained bronchoalveolar lavage (BAL) samples from 88 patients with respiratory failure secondary to severe SARS-CoV-2 pneumonia and compared them with BAL samples prospectively collected before and during the COVID-19 pandemic from 211 patients with pneumonia secondary to other pathogens and 42 intubated patients without pneumonia. For many patients, we were able to obtain samples within 48 h of intubation and sequentially over the course of the illness, allowing us to gain insights about the early pathogenesis and progression of COVID-19-induced respiratory failure. We profiled BAL samples using flow cytometry and performed bulk transcriptomic profiling of alveolar macrophages. Additionally, we performed single-cell RNA sequencing (RNA-seq) on BAL fluid collected less than 48 h after intubation from 10 patients with severe SARS-CoV-2 pneumonia. We used these data to develop a testable hypothesis to explain SARS-CoV-2 pathobiology (Fig. 1a).
a, Our model of the alveolus during infection with SARS-CoV-2, based on the main findings. (1) The normal alveolus contains ACE2-expressing alveolar type 1 and type 2 cells (AT1 and AT2, respectively) and TRAMs. (2) SARS-CoV-2 infects AT1 and AT2 cells and TRAMs. Infected TRAMs express T cell chemokines. (3) Cross-reactive or de novo-generated effector-memory T cells recognize SARS-CoV-2 antigens presented by TRAMs and produce IFNγ, further activating TRAM to produce cytokines and chemokines. (4) Activated T cells proliferate and continue to produce IFNγ, eventually leading to death of infected TRAMs and recruitment of monocytes, which rapidly differentiate into MoAMs. (5) Recruited MoAMs become infected with SARS-CoV-2, continuing to present antigens to T cells and maintaining the feedback loop until viral clearance is achieved. b, Timing of hospital admission, BAL fluid collection, duration of mechanical ventilation and duration of hospital stay (thin grey line) in patients with severe COVID-19, grouped by outcomes. Day 0 is defined as the day of the first intubation. c, Distribution of patient age. Differences not significant by pairwise t-test with false discovery rate (FDR) correction. d, Proportions of women (red) and men (blue) (pairwise χ2-tests of proportions with continuity and FDR correction). e, Self-reported ethnicity (pairwise χ2-tests of proportions with continuity and FDR correction). f, Body mass index (BMI) (t-test with FDR correction). g, SOFA score (pairwise Wilcoxon rank-sum tests with FDR correction). h, APS (pairwise Wilcoxon rank-sum tests with FDR correction). i, Length of stay in ICU (pairwise t-tests with FDR correction). j, Duration of mechanical ventilation (pairwise t-tests with FDR correction). k, Mortality in patients with COVID-19 was similar to patients in other groups (25% versus 35%, P = 0.10, χ2 = 2.63, χ2-tests of proportions).
Demographics of the cohort
Samples were collected as part of the Successful Clinical Response in Pneumonia Therapy (SCRIPT) Systems Biology Center, an observational study of patients with severe pneumonia (defined as those who require mechanical ventilation). During the initial phase of the pandemic, we prospectively enrolled 88 of the 179 patients with SARS-CoV-2-induced pneumonia and respiratory failure requiring mechanical ventilation in the intensive care unit (ICU) at Northwestern Memorial Hospital in Chicago (Fig. 1b, Extended Data Fig. 1a–c, Extended Data Table 1).
We compared patients with COVID-19 with 253 mechanically ventilated participants enrolled in the two years before and during the pandemic. These included patients intubated for reasons other than pneumonia (non-pneumonia control), patients diagnosed with severe non-SARS-CoV-2 viral pneumonia (other viral pneumonia) or patients with severe pneumonia secondary to bacterial or fungal pathogens (other pneumonia). Compared with these patients, patients with SARS-CoV-2 pneumonia had similar age, race and sex profiles, but had a significantly higher self-reported Hispanic or Latino ethnicity and body mass index (Fig. 1c–f, Extended Data Table 1).
Severity of illness estimated using the sequential organ failure assessment (SOFA) score and the acute physiology score (APS) was similar in patients with SARS-CoV-2 pneumonia compared with other pneumonia and was comparable to that observed in a recent study of ARDS6 (Fig. 1g, h, Extended Data Table 1). Patients with severe SARS-CoV-2 pneumonia had longer lengths of stay in the ICU and required longer periods of ventilation compared with all pneumonia and non-pneumonia controls (Fig. 1i, j). On the day that their first BAL samples were taken, patients with severe SARS-CoV-2 pneumonia had increased levels of C-reactive protein compared with patients with other pneumonias, whereas other biomarkers of inflammation were found at similar levels (Extended Data Fig. 1d–i). The BAL sampling rate per day among patients with COVID-19 was not higher than the sampling rate among patients with other pneumonias (Extended Data Fig. 1j–l). Mortality was not different between patients with SARS-CoV-2 pneumonia compared with the entire cohort (Fig. 1k).
BAL fluid composition in SARS-CoV-2 pneumonia
We began by performing flow cytometry on BAL samples collected within 48 h of intubation (Fig. 2a, b, Extended Data Fig. 2a–e). We found that, despite severe pneumonia requiring mechanical ventilation, only 31% of patients with severe COVID-19 exhibited neutrophilia (defined as a percentage of neutrophils over 50%) in BAL fluid (Fig. 2a, b). Instead, we found that in patients with severe SARS-CoV-2 pneumonia, the alveolar space was significantly enriched for CD4+ and CD8+ T cells and monocytes (Fig. 2a, b, Extended Data Fig. 3a).
a, Hierarchical clustering of flow cytometry data from BAL samples collected within 48 h of intubation. Column headers are colour-coded by the diagnosis and presence or absence of co-infection (infection status). Samples were clustered by Euclidean distance using Ward’s method. AM, alveolar macrophages. b, Proportions of cells detected within 48 h of intubation (q < 0.05, pairwise Wilcoxon rank-sum tests with FDR correction). Comparisons are not significant unless otherwise noted. c, k-means clustering of the 1,194 significantly variable genes (q < 0.05, likelihood-ratio test) across diagnoses. Columns represent each individual sample and are clustered using Ward’s method. Column headers are colour-coded by the diagnosis and presence or absence of a co-infecting pathogen (infection status). Representative genes and GO biological processes are shown for each cluster. In b, c, infection status refers only to the COVID-19 and ‘other viral pneumonia’ groups; blanks in these groups refer to samples for which microbiology data were incomplete and infectious status could not be determined; ‘viral infection only’ refers to viral pathogens as the only detected pathogen in a sample and ‘viral infection with bacterial or fungal co-infection’ refers to detection of a viral pathogen with one or more bacterial or fungal co-pathogens.
Distinct macrophage response in COVID-19
As macrophages respond to alterations in their microenvironment7, we reasoned that changes in the alveolar macrophage transcriptome may reflect unique features of the SARS-CoV-2-infected alveolus. We isolated alveolar macrophages from BAL fluid collected within the first 48 h after intubation from patients with severe SARS-CoV-2 pneumonia and compared them with alveolar macrophages from patients with pneumonia secondary to other pathogens, non-pneumonia controls and healthy volunteers. k-means clustering of the 1,194 significantly variable genes (q < 0.05, likelihood-ratio test) across diagnosis identified 5 clusters (Fig. 2c, Supplementary Data 1–3). Notably, the majority of patients with COVID-19 clustered together. Cluster 1 contained genes specifically upregulated in patients with COVID-19 and was characterized by genes involved in the response to interferon. Cluster 1 also included genes encoding the chemokines CCL7, CCL8 and CCL13, which drive recruitment of monocytes and T cells.
To detect viruses in alveolar macrophages, we aligned RNA-seq reads to a hybrid genome including the human, SARS-CoV-2 and influenza A/California/07/2009 reference genomes. An additional negative-strand SARS-CoV-2 transcript, which is transiently formed during its replication, was included to detect replicating virus8. We detected SARS-CoV-2 transcripts in alveolar macrophage transcriptomes from 67% of samples with PCR-confirmed SARS-CoV-2 infection. In 38% of these samples, we detected both positive- and negative-strand SARS-CoV-2 transcripts (Extended Data Fig. 3b, c).
SARS-CoV-2 pneumonia persists over time
Unlike other types of pneumonia, SARS-CoV-2 pneumonia is characterized by a long duration between symptom onset and the development of respiratory failure (6–12 days) and a prolonged course of mechanical ventilation1,4 (Fig. 1b, i, j). To determine whether the unique cellular composition of the BAL fluid and the interferon-response signature in alveolar macrophages persists over the course of the disease, we analysed samples obtained early (less than 48 h) after intubation and samples obtained more than 48 h after intubation. Hierarchical clustering of BAL samples from all time points from serially sampled patients as well as direct pairwise comparison of early (less than 48 h after intubation) versus late (more than 48 h after intubation) samples between groups demonstrated that, in comparison with BAL samples from other pneumonia, samples from patients with COVID-19 were persistently enriched for T cells (Fig. 3a, b, Extended Data Fig. 4a, b). These findings persisted in the 40% of patients with severe SARS-CoV-2 pneumonia in whom a superinfecting pathogen was detected over the course of mechanical ventilation (Extended Data Table 2). Nevertheless, neutrophils were increased in BAL fluid samples collected after 48 h of mechanical ventilation in patients with COVID-19, probably attributable to the duration of mechanical ventilation and the attendant risk of bacterial superinfection (Extended Data Fig. 4c).
a, Hierarchical clustering of flow cytometry analysis of BAL samples from all time points from patients with serial sampling (n > 1) on the basis of their composition. Column headers are colour-coded by the diagnosis, patient, duration of mechanical ventilation, and presence or absence of superinfection (infection status). Samples were clustered by Euclidean distance using Ward’s method. b, Comparison of percentage of CD4+ and CD8+ T cells and neutrophils between early (<48 h after intubation) and late (>48 h of mechanical ventilation) samples (q < 0.05, pairwise Wilcoxon rank-sum tests with FDR correction). c, k-means clustering of the 2,323 significantly variable genes (q < 0.05, likelihood-ratio test) across diagnoses, columns represent each individual patient and clustered using Ward’s method. Representative genes and GO biological processes are shown for each cluster. Column headers are colour-coded by the diagnosis, duration of mechanical ventilation (white indicates chronically ventilated patients), and presence or absence of superinfection (infection status). In a, c, ‘infection status’ refers only to the COVID-19 and ‘other viral pneumonia’ groups; blanks in these groups refer to samples for which microbiology data were incomplete and infectious status could not be determined; ‘viral infection only’ refers to viral pathogens as the only detected pathogen in a sample and ‘viral infection with bacterial or fungal superinfection’ refers to detection of a viral pathogen with one or more bacterial or fungal co-pathogens.
We then performed RNA-seq on alveolar macrophages isolated early (less than 48 h) and later over the course of mechanical ventilation. k-means clustering of the 2,323 significantly variable genes (q < 0.05, likelihood-ratio test) across diagnoses identified five clusters (Fig. 3c, Extended Data Fig. 5a, Supplementary Data 1–3). Genes and related Gene Ontology (GO) biological processes associated with these clusters were notably similar to those identified in samples collected within 48 h after intubation (Fig. 2c). Specifically, samples from patients with COVID-19 continued to cluster together and were differentiated by increased expression of genes in cluster 1, which was enriched for interferon-response genes and T cell chemokines (Fig. 3c, Extended Data Fig. 5a, b).
To identify gene modules in alveolar macrophages that distinguish pneumonia type and outcome, we performed weighted gene co-expression network analysis (WGCNA) (Extended Data Fig. 5c, Supplementary Data 1). Module 15 was enriched for interferon-responsive genes and correlated with the detection of SARS-CoV-2 transcripts, levels of C-reactive protein, CD8+ T cell abundance and COVID-19 diagnosis (R = 0.28, 0.30, 0.28, 0.25, respectively). Notably, all SARS-CoV-2 genes included in this analysis were assigned to this module, further underscoring the relevance of the disease diagnosis category. Consistent with the results of k-means clustering, module 15 was enriched for type I and type II interferon-response genes (GO:0060337 and GO:0060333). We observed a significant negative correlation between interferon signalling and the duration of mechanical ventilation (Extended Data Fig. 5d–f). In addition, we identified modules related to macrophage maturation, including module 4, which exhibited a positive correlation with the percentage of CD206hi alveolar macrophages (R = 0.61), and module 12, which exhibited a negative correlation with percentage of CD206hi macrophages (R = −0.62) (Extended Data Fig. 5c). The unique features of SARS-CoV-2 pneumonia are further illustrated by a UMAP projection of all bulk RNA-seq samples, which separated largely by diagnosis, module 15 gene expression and abundance of T cells (Extended Data Fig. 5g).
We took advantage of the serial samples collected from the same patients with COVID-19 and explored the relationship between expression of canonical interferon-response genes and patient outcomes. Analysis of the serial samples from patients with COVID-19 who were discharged home (five patients) or to an inpatient facility (two patients) demonstrated decreased expression of interferon-response genes as the disease progressed, whereas in patients who were discharged to a long-term acute care facility (LTAC) (five patients) or who died (two patients), the expression of interferon-response genes was largely unchanged (Extended Data Fig. 5h). Consistent with these findings, we observed a significant negative correlation between abundance of SARS-CoV-2 transcripts in patients with confirmed COVID-19 and the time since intubation (ρ = −0.49, Spearman correlation) (Extended Data Fig. 5i).
Immune cell circuits in SARS-CoV-2 pneumonia
We performed single-cell RNA-seq on 10 patients with COVID-19 from whom BAL samples were collected within 48 h of intubation (Fig. 4a, Extended Data Fig. 6a–d). We included two additional patients, one with bacterial pneumonia and one non-pneumonia control (Extended Data Fig. 6e–h). Analysis of an integrated object resolved multiple clusters corresponding to macrophages and other cell types (Supplementary Data 4). Alveolar macrophages contained six clusters. Four clusters of monocyte-derived alveolar macrophages (MoAMs) were characterized by CCL2 expression and gradually increasing expression of genes associated with alveolar macrophage maturation. The other two clusters expressed markers of tissue-resident alveolar macrophages (TRAM) (Fig. 4b, Extended Data Fig. 6a). We did not detect expression of type I interferons in our single-cell dataset or in other publicly available single-cell RNA-seq datasets9,10 obtained from BAL fluid later in the clinical course of patients with COVID-19 (data not shown). By contrast, expression of type II interferon (IFNG) was detected in T cells from all ten patients with COVID-19 (Fig. 4c, Extended Data Fig. 6b). These results suggest that the interferon-response gene signature we observed in alveolar macrophages may be in response to IFNγ released from activated T cells.
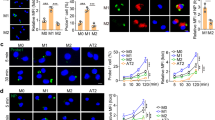
a, UMAP plot showing integrated analysis of 77,146 cells isolated from 10 patients with severe COVID-19 within 48 h after intubation. DC1, conventional dendritic cells type 1 (CLEC9A+); DC2, conventional dendritic cells type 2 (CD1C+); migratory DC, migratory dendritic cells (CCR7+); pDC, plasmacytoid dendritic cells (CLEC4C+); mixed myeloid, mixed cluster containing transitory dendritic cells and very immature monocyte-derived macrophages; iNKT cells, invariant natural killer T cells; Treg, regulatory T cells (FOXP3+). b, Heat map demonstrating expression of the selected genes of interest in two subsets of tissue-resident (TRAM1 and TRAM2) and four subsets of monocyte-derived (MoAM1, MoAM2, MoAM3 and MoAM4) alveolar macrophages. Each column represents a single patient with COVID-19. The MRC1 gene encodes CD206. c, Expression of IFNG is restricted to T cells. d, Detection of SARS-CoV-2 transcripts. Plot shows cumulative number of SARS-CoV-2 genes plus negative strand. e–g, Specific upregulation of CXCL10 (e), CCL4 (f) and IL1B (g) in TRAM2 cells. Density projection plots, expression averaged within hexagonal areas on UMAP. h, Heat map demonstrating selected differentially expressed genes between two subsets of tissue-resident alveolar macrophages (TRAM1 and TRAM2). i, SARS-CoV-2 infection is spatially restricted. Combined immunofluorescence microscopy for CD206, a marker of mature macrophages (red arrows) and smFISH (RNAscope) for positive- (yellow arrowheads) and negative-strand (cyan doublehead arrows) SARS-CoV-2 transcripts. Expanded regions show infected and non-infected CD206-positive alveolar macrophages in the adjacent alveoli.
As expected, positive- and negative-strand SARS-CoV-2 transcripts were detected in epithelial cells. We also detected SARS-CoV-2 transcripts in migratory CCR7+ dendritic cells, and MoAM2 and TRAM2 clusters, which do not express ACE2 (Fig. 4d, Extended Data Fig. 6c). Coronaviruses generate large numbers of positive-strand transcripts from a single negative-strand template8,11. Consistent with this known biology, we detected more transcripts for positive compared with negative strands in both our single-cell and bulk RNA-seq data (Fig. 4d, Extended Data Figs. 3c, 6d). These results suggest that alveolar macrophages contain SARS-CoV-2 and that they may support viral replication, as has been reported for SARS-CoV and Middle East respiratory syndrome coronavirus (MERS-CoV)12,13,14.
TRAM2 cells containing SARS-CoV-2 exhibited a distinct transcriptional program compared with uninfected TRAM1 (Fig. 4b, h, Extended Data Fig. 6g, h, Supplementary Data 5). In patients with COVID-19, genes distinguishing infected TRAM2 from non-infected TRAM1 cells included several chemokines and cytokines that are important for T cell and monocyte recruitment, such as CCL4, CCL20, CXCL10 and CXCL11. TRAM2 also expressed IL1B, TNFSF10 and DEFB1 (Fig. 4b, e–h, Supplementary Data 6). Finally, infected TRAM2 cells were marked by increased expression of interferon-response genes compared with non-infected TRAM1 cells (Fig. 4b, h).
IL-6 induces the transcription of clotting factors in the liver and tissue factor in the endothelium to promote thrombosis, and elevated levels of IL-6 predict mortality in patients with COVID-1915,16,17. Because single-cell RNA-seq analyses of peripheral blood from patients with COVID-19 failed to identify circulating cells producing IL-618,19, it has been suggested that IL-6 is produced by inflammatory cells in the alveolus17. In our dataset, the overall expression of IL6 was low and primarily restricted to very immature MoAM1 cells originating from two patients (Extended Data Fig. 7a). Furthermore, the expression levels of IL6 in BAL fluid cells from patients with COVID-19 was similar to previously reported IL6 expression levels in alveolar macrophages from patients with later-stage COVID-199 and lower than the level of IL6 expression observed in stromal and endothelial cells from a published single-cell RNA-seq dataset describing the healthy or fibrotic lung20. Consistently, in our bulk RNA-seq data from flow cytometry-sorted alveolar macrophages, IL6 expression was not different between patients with pneumonia secondary to SARS-CoV-2 compared with other respiratory pathogens, although it was higher than in healthy controls, in whom IL6 transcripts were never detected (Extended Data Fig. 5b). Although we did not sample neutrophils, there was no increased IL6 expression in neutrophils in single-cell RNA-seq datasets from BAL fluid from patients with SARS-CoV-2 pneumonia9,10.
In mouse models of influenza A viral pneumonia, IFNγ drives apoptosis of tissue-resident alveolar macrophages21. We therefore investigated whether the CD206hi alveolar macrophages in patients with severe SARS-CoV-2 pneumonia were truly tissue-resident alveolar macrophages or maturing MoAMs. We performed cell-type deconvolution of our bulk RNA-seq data and found that in patients with SARS-CoV-2 pneumonia, only a small fraction of alveolar macrophages were tissue-resident alveolar macrophages. Instead, the majority of alveolar macrophages were MoAM3 cells (Extended Data Fig. 7b, c).
The BAL procedure samples cells from a lung segment, which includes many thousands of alveoli. Therefore, the detection of infected and non-infected alveolar macrophages in the same BAL fluid sample in our single-cell RNA-seq data suggests that infected and uninfected alveoli co-exist within a single lung segment. To test this prediction, we examined lung tissue from a patient who died from SARS-CoV-2 pneumonia while forgoing treatments, except for comfort measures. Using single-molecule fluorescence in situ hybridization (smFISH), we detected positive- and negative-strand SARS-CoV-2 transcripts in the lung epithelium and alveolar macrophages (Fig. 4i, Supplementary Data 7, Supplementary Video 1). Positive-strand SARS-CoV-2 transcripts exhibited diffuse cytoplasmic staining, and the number of punctae exceeded that of negative transcripts. As predicted from our model of lung infection, cells containing SARS-CoV-2 transcripts were also detected in a spatially restricted manner, as regions of the lung containing the virus were adjacent to the regions with nearly normal architecture where viral particles were absent.
Discussion
A minority of patients with severe SARS-CoV-2 pneumonia develop respiratory failure, but it is these patients who account for almost all of the morbidity, mortality and socioeconomic cost associated with the COVID-19 pandemic. We systematically sampled the alveolar space in patients with new-onset respiratory failure secondary to SARS-CoV-2 pneumonia and compared these samples with those from a large comparison cohort of patients with pneumonia from other respiratory pathogens collected before and during the pandemic. We used these data to develop a model for the unique pathobiology of SARS-CoV-2 pneumonia (Fig. 1a). Our model proposes that SARS-CoV-2 initially infects and replicates in epithelial cells in the nasopharynx, which express relatively high levels of ACE2 compared with epithelial cells in lower airways or the distal lung22,23. Whether by progressive movement distally in the tracheobronchial tree or via aspiration of nasopharyngeal contents, some virus gains access to the distal alveolar space. In the alveolar space, SARS-CoV-2 infects alveolar epithelial cells and tissue-resident alveolar macrophages24. A transcriptional program that promotes the recruitment of memory T cells to the alveolar space is activated within infected tissue-resident alveolar macrophages. There, memory T cells become activated, releasing IFNγ, which activates tissue-resident alveolar macrophages containing SARS-CoV-2. Eventually, these tissue-resident alveolar macrophages die and monocyte-derived alveolar macrophages are recruited, which in turn become infected with SARS-CoV-2 to sustain the inflammatory signalling loop with T cells. These infected alveolar macrophages may act similarly to a ‘Trojan horse’, transferring the virus to adjacent lung regions, slowly propagating SARS-CoV-2 infection across the lung.
Our model is informed by a wealth of causal data generated in cell and animal models of SARS-CoV-2 and the related coronaviruses SARS-CoV and MERS-CoV. SARS-CoV-2 proteins have been shown to suppress type I interferon responses24,25,26, and we did not detect ongoing expression of type I interferons in our bulk or single-cell RNA-seq data25,27. Our data suggesting that SARS-CoV-2 replicates in alveolar macrophages are consistent with reports that SARS-CoV, MERS-CoV, and SARS-CoV-2 can infect macrophages in vitro12,13,14,24,28, and the detection of SARS-CoV-2 in alveolar macrophages in autopsy studies of patients with SARS-CoV-2 pneumonia29.
Alveolar macrophage infection might result from phagocytosis of infected alveolar epithelial cells followed by viral escape from the lysosome. Alternatively, alveolar macrophages might be directly infected, as was shown for SARS-CoV and MERS-CoV12,14. Finally, antibody-dependent enhancement has been suggested from cell and animal models of SARS-CoV and MERS-CoV infection30,31.
Activation of memory T cells leads to IFNγ production, local proliferation of activated memory T cells, mounting of inflammatory responses and recruitment of monocytes and T cells. We observed enrichment of CD4+ and CD8+ T cells in the alveolar space of patients with SARS-CoV-2 pneumonia relative to pneumonia secondary to other pathogens. Furthermore, our single-cell RNA-seq data confirmed production of IFNγ by both CD4+ and CD8+ T cells. These results raise the question of how T cells in the alveolar space become activated following SARS-CoV-2 infection. Although tissue-resident alveolar macrophages are poor antigen-presenting cells and do not convert naive T cells into effector T cells32, a low level of antigen presentation by alveolar macrophages might be sufficient to activate pre-existing memory T cells that cross-react with SARS-CoV-2. Existence of such cross-reactive memory T cells has been reported for SARS-CoV33 and for SARS-CoV-234,35,36,37,38. A larger number of SARS-CoV-2 cross-reactive memory T cells was observed independently in elderly people and in patients with severe COVID-19, compared with those with mild COVID-1938. These cross-reactive T cells exhibited lower avidity and reduced antiviral responses in response to stimulation with SARS-CoV-2 peptides compared with T cells from patients who recovered from COVID-19. This mechanism might explain the epidemiology of severe SARS-CoV-2 pneumonia, which disproportionately affects elderly individuals.
Single-cell RNA-seq atlases and smFISH studies of the normal human lung show that only a small number of alveolar epithelial cells express ACE2, the gene encoding the receptor for SARS-CoV-2 entry22,23. By contrast, the sialic acid residues that serve as receptors for influenza A virus are abundantly expressed in alveolar type 2 cells22. Thus, whereas influenza A infects large numbers of cells leading to rapid viral replication, widespread injury, robust antiviral responses and death of infected epithelial cells, infection by SARS-CoV-2 is likely to lead to spatially localized areas of infection. This could explain the localized areas of ground glass infiltrates observed in chest computed tomography in minimally symptomatic patients with COVID-1939. We speculate that alveolar macrophages containing SARS-CoV-2 might spread virus between alveoli. For example, tissue-resident alveolar macrophages, once thought to be sessile, were recently shown to travel between alveoli through pores of Kohn, particularly during viral infection40. In each new area of infection, positive feedback loops between alveolar macrophages containing the virus and activated T cells could promote persistent injury and inflammation.
Our model of SARS-CoV-2 pneumonia as a slowly progressive, spatially restricted infection explains some of the unusual clinical features of COVID-19. Most notably, the clinical course of severe SARS-CoV-2 pneumonia is much longer than that of other respiratory viruses. The time from the onset of symptoms to respiratory failure in patients with SARS-CoV-2 infection is 6–12 days, compared with 1–3 days or even less in patients with influenza A virus infection4,41. Furthermore, in our cohort the duration of mechanical ventilation and ICU stay were much longer in patients with severe SARS-CoV-2 pneumonia compared with other respiratory pathogens causing pneumonia, despite a similar severity of illness and mortality. This longer clinical course might also explain some of the systemic complications of the disease. Although sparsely sampled, the levels of inflammatory biomarkers in the blood were similar in patients with COVID-19 and those with other aetiologies of pneumonia in our cohort, confirming more recent systematic reports42,43. This observation raises the possibility that the increased number of systemic complications observed in patients with SARS-CoV-2 pneumonia are related to prolonged, rather than higher level, exposure to circulating inflammatory cytokines.
In summary, we present a dataset that supports a testable model in which alveolar macrophages containing SARS-CoV-2 form positive feedback loops with IFNγ-secreting T cells to promote alveolitis in patients with severe COVID-19.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Human participants
All research involving human participants was approved by the Northwestern University Institutional Review Board. Samples from patients with COVID-19, viral pneumonia and other pneumonia, and non-pneumonia controls were collected from participants enrolled in the Successful Clinical Response In Pneumonia Therapy (SCRIPT) study STU00204868. Alveolar macrophages from healthy volunteers were obtained under study STU00206783. Autopsy tissues were obtained by the usual protocols and analysed under study STU00079445. All study participants or their surrogates provided informed consent.
Individuals of at least 18 years of age with suspicion of pneumonia based on clinical criteria (including but not limited to fever, radiographic infiltrate and respiratory secretions) were screened for enrolment into the SCRIPT study. Inability to safely perform BAL or non-bronchoscopic bronchoalveolar lavage (NBBAL) were considered exclusion criteria44. In our centre, patients with respiratory failure are intubated on the basis of the judgement of bedside clinicians for worsening hypoxaemia, hypercapnia or work of breathing refractory to high-flow oxygen or non-invasive ventilation modes. Extubation occurs on the basis of the judgement of bedside clinicians following a protocol-driven trial of spontaneous breathing in patients demonstrating physiological improvement in their cardiorespiratory status during their period of mechanical ventilation.
We collected data and samples from patients enrolled in SCRIPT from 15 June 2018 to 6 July 2020 in the ICU at Northwestern Memorial Hospital in Chicago. We prospectively enrolled 88 of the 179 patients with SARS-CoV-2-induced pneumonia and respiratory failure requiring mechanical ventilation in the ICU (Fig. 1b, Extended Data Fig. 1a, b; Extended Data Table 1), all but one of whom had been discharged at the time of our submission. Management of patients with COVID-19 was guided by protocols published and updated on the Northwestern Medicine intranet as new information became available over the pandemic. Clinical laboratory testing including studies ordered on BAL fluid was at the discretion of the care team; however, quantitative cultures, multiplex PCR (BioFire Film Array Respiratory 2 panel), and automated cell count and differential were recommended by local ICU protocols. Most patients also underwent urinary antigen testing for Streptococcus pneumoniae and Legionella pneumophilia serogroup 1 on admission. Clinicians were encouraged to manage all patients, including those with COVID-19, according to ARDSNetwork protocols including the use of a higher PEEP/lower FiO2 strategy for those with severe hypoxaemia45,46. Prone positioning (16 h per day) was performed in all patients with a PaO2/FiO2 <150 who did not have contraindications47. In those who had a response to prone positioning evidenced by improved oxygenation, prone positioning was repeated. Oesophageal balloon catheters (Cooper Surgical) were placed at the discretion of the care team to estimate transpulmonary pressure and optimize PEEP, particularly in patients with a higher-than-normal BMI.
Autopsy specimen used for smFISH (RNAscope) (Fig. 4i): an 81-year-old woman with end-stage renal disease and cirrhosis was admitted for a fever of 38.5 °C. She had a positive nasopharyngeal swab for SARS-CoV-2. The patient developed increased O2 requirements and was subsequently transferred to the COVID ICU. In the ICU, the decedent developed hypotension to about 60/40 mm Hg and after discussion with the clinical team, the decedent’s family elected to focus on comfort care. The patient died 8 days after admission.
NBBAL and BAL procedures
Consent was obtained from patients or their surrogates for bronchoscopic procedures. Bronchoscopic BAL was performed in intubated ICU patients with flexible, single-use Ambu aScope (Ambu) devices. Patients were given sedation and topical anaesthetic at the physician proceduralist’s discretion. Vital signs were monitored continuously throughout the procedure. The bronchoscope was wedged in the segment of interest based on available chest imaging or intra-procedure observations, aliquots of 30 ml of normal saline at a time, generally 90–120 ml total, were instilled and aspirated back. The fluid returned following the first aliquot was routinely discarded. Samples were split (if sufficient return volume was available) and sent for clinical studies and an aliquot was reserved for research. A similar procedure was applied to NBBAL; however, NBBAL was performed with directional (lateral) but not visual guidance, and as usual procedural care by a respiratory therapist rather than a pulmonologist44.
For bronchoscopies performed in patients with COVID-19, additional precautions were taken to minimize the risk to healthcare workers including only having essential providers present in the room, clamping of the endotracheal tube, transient disconnection of the inspiratory limb from the ventilator, and preloading of the bronchoscope through the adaptor48. Sedation and neuromuscular blockade to prevent cough were administered for these procedures at the physician’s discretion. In most cases of early bronchoscopy, the procedure was performed immediately after intubation, taking advantage of neuromuscular blockade administered for the intubation procedure.
For all patients with COVID-19, samples were collected from regions of greatest chest radiograph abnormality by a critical care physician using a disposable bronchoscope. The majority of samples before the pandemic were collected by respiratory therapists using a NBBAL catheter that is the same diameter as a standard bronchoscope with the catheter directed to the most radiographically affected lung. For both bronchoscopic and NBBAL, the recommended instillate volume was 120 ml and the initial aliquot was discarded if adequate return was obtained44.
Pneumonia adjudication
Five critical care physicians (J.M.K., C.O.P., B.D.S., J.M.W. and R.G.W.) retrospectively adjudicated patients as COVID-19 pneumonia, non-COVID-19 viral pneumonia, pneumonia secondary to other pathogens, or non-pneumonia controls (intubated for reasons other than pneumonia), according to a standardized adjudication procedure (the adjudication protocol can be found in Supplementary Data 8). Non-pneumonia controls were defined as patients who underwent BAL to exclude pneumonia but had negative quantitative cultures, a negative multiplex PCR for viral and bacterial pathogens (when available) and negative urine antigens for S. pneumoniae and L. pneumophilia serogroup 1, as well as an alternative diagnosis. The treating clinician’s impression was considered, but concordance was not required. Subsequent course and the entirety of the clinical record was used for adjudication. Some of the patients adjudicated as non-pneumonia controls developed ventilator-associated pneumonia later in their clinical course. Viral pneumonia was diagnosed on the basis of detection of a respiratory viral pathogen from either a nasopharyngeal swab or BAL fluid in the appropriate clinical setting. Bacterial pneumonia was defined as positive quantitative cultures with more than 100 colony-forming units per ml, detection of a bacterial pathogen by PCR analysis of BAL fluid or a positive urine antigen. Over the course of the study, BAL fluid was analysed using a methicillin-resistant Staphylococcus aureus (MRSA) PCR (MRSA/SA SSTI) and the BioFire FilmArray Respiratory 2 (RP2) panel and Pneumonia panels. SARS-CoV-2 was detected with a variety of assay platforms including the Cepheid Gene Expert, Abbott ID NOW, Becton Dickinson, and a locally developed and validated PCR. For some patients without COVID-19, the diagnosis of pneumonia was made on the basis of clinical suspicion, radiographic findings and response to antimicrobial therapy in the absence of an identified pathogen. Ventilator-associated pneumonia was diagnosed as detection of a new respiratory pathogen using quantitative culture or PCR more than 48 h after intubation, the detection of a new respiratory pathogen on serial BAL samples, or the reappearance of a respiratory pathogen after a negative BAL on a subsequent study. Clinical laboratory data were obtained from the Northwestern Medicine Enterprise Data Warehouse using Structured Query Language (SQL). APS and SOFA scores were generated from the Electronic Health Record using previously validated programming.
Flow cytometry and cell sorting
NBBAL and BAL samples were filtered through a 70-μm cell strainer, pelleted by centrifugation at 300g for 10 min at 4 °C, followed by hypotonic lysis of red blood cells with 2 ml BD PharmLyse reagent for 2 min. Lysis was stopped by adding 13 ml MACS buffer (Miltenyi Biotech). Cells were pelleted again and resuspended in 100 μl of 1:10 Fc-Block (Human TruStain FcX, Biolegend) in MACS buffer, and a 10-μl aliquot was taken for counting using K2 Cellometer (Nexcelom) with AO/PI reagent. The volume of Fc-Block was adjusted so the concentration of cells was always less than 5 × 107 cells per ml and the fluorophore-conjugated antibody cocktail was added in 1:1 ratio (Extended Data Table 3). After incubation at 4 °C for 30 min, cells were washed with 5 ml MACS buffer, pelleted by centrifugation, and resuspended in 500 μl MACS buffer with 2 μl SYTOX Green viability dye (ThermoFisher). Cells were sorted on a FACS Aria III SORP instrument using a 100-μm nozzle at 20 psi. Cells were sorted into 300 μl of MACS buffer for bulk RNA-seq or 300 μl of 2% BSA in PBS for single-cell RNA-seq. Sample processing was performed in BSL-2 facility using BSL-3 practices. Analysis of the flow cytometry data was performed using FlowJo 10.6.2. using a uniform sequential gating strategy (Extended Data Fig. 2a, b) reviewed by three investigators (S.S., B.D.S. and A.V.M.). Immune populations were defined using canonical markers44,49 as shown in Extended Data Fig. 2a, b. Alveolar macrophages were defined by the expression of CD206 (mannose receptor). We further subdivided alveolar macrophages into CD206lo alveolar macrophages, which represent differentiating monocyte-derived alveolar macrophages, and CD206hi alveolar macrophages, which include both tissue-resident alveolar macrophages, present in the lung before the onset of pneumonia, and mature monocyte-derived alveolar macrophages. Because CD206hi alveolar macrophages can be found in BAL fluid from both healthy volunteers and patients across different types of pneumonia, and the presence of CD206lo alveolar macrophages varies across the patients with pneumonia49,50,51, we focused our transcriptomic analysis on CD206hi alveolar macrophages. Abundance of specific populations in individual BAL fluid samples can be found in Supplementary Data 9.
Bulk RNA-seq of flow cytometry-sorted alveolar macrophages
Immediately after sorting, cells were pelleted by centrifugation and lysed in 350 μl RLT Plus lysis buffer (Qiagen) supplemented with 2-mercaptoethanol. Lysates were stored at −80 °C until RNA isolation using the AllPrep DNA/RNA Micro kit according to the manufacturer’s protocol (Qiagen). RNA quality and quantity were assessed using TapeStation 4200 High Sensitivity RNA tapes (Agilent), and RNA-seq libraries were prepared from 250 pg of total RNA using SMARTer Stranded Total RNA-seq Kit v2 (Takara Bio). Libraries were pooled using dual indexing and sequenced on a NextSeq 500 instrument (Illumina), 75 cycles, single-end, to an average sequencing depth of 19.55 million reads.
FASTQ files were generated using bcl2fastq (Illumina). To enable detection of viral RNA, a custom hybrid genome was prepared by joining FASTA, GFF and GTF files for GRCh37.87, SARS-CoV-2 (NC_045512.2), and Influenza A/California/07/2009 (GCF_001343785.1), which was the dominant strain of influenza throughout BAL fluid collection at our hospital52. An additional negative strand transcript spanning the entirety of the SARS-CoV-2 genome was then added to the GTF and GFF files to enable detection of SARS-CoV-2 replication. Normalized counts tables later revealed high enrichment of SARS-CoV-2 transcripts in patients diagnosed with COVID-19, and enrichment of influenza A virus genes in patients marked as other viral pneumonia. Of note, as our alveolar macrophage sorting strategy for bulk RNA-seq (Extended Data Fig. 2a, b) focused only on CD206hi cells, our bulk RNA-seq data probably underestimate infection of alveolar macrophages infected with SARS-CoV-2 (Fig. 2d).
To facilitate reproducible analysis, samples were processed using the publicly available nf-core/RNA-seq pipeline version 1.4.2 implemented in Nextflow 19.10.0 using Singularity 3.2.1-1 with the minimal command nextflow run nf-core/rnaseq -r 1.4.2 –singleEnd -profile singularity –reverseStranded –three_prime_clip_r2 353,54,55. In brief, lane-level reads were trimmed using trimGalore! 0.6.4 and aligned to the hybrid genome described above using STAR 2.6.1d56. Gene-level assignment was then performed using featureCounts 1.6.457. Putative sample swaps were identified first by comparing known patient sex with sex determined by levels of XIST and RPS4Y1 expression, followed by single-nucleotide polymorphism analysis with NGSCheckMate v.1.0.0 in FASTQ mode using default settings58. Samples exhibiting unexpected correlation were excluded from analysis.
Bulk differential expression analysis
All analysis was performed using custom scripts in R version 3.6.3 using the DESeq2 v.1.26.0 framework59. Correspondence between lanes was first confirmed by principal component analysis (PCA) before merging counts using the command collapseReplicates(). One outlier sample from the ‘other pneumonia’ group with low RIN score and exhibiting extreme deviation on PCA and poor alignment and assignment metrics was excluded from downstream analysis. For differential expression analysis (DEA), both proportion of alveolar macrophage from flow cytometry data and diagnosis were used as explanatory factors. A ‘local’ model of gene dispersion was used, as this better fit dispersion trends without obvious overfitting, and gene outlier replacement was disabled; otherwise default settings were used (see code for details).
k-means clustering of bulk samples
A custom-built function was used (available at https://github.com/NUPulmonary/utils/blob/master/R/k_means_Figure.R) for k-means clustering. In brief, variable genes were identified using a likelihood-ratio test with local estimates of gene dispersion in DESeq2 with diagnosis as the full model as well as a reduced model corresponding to intercept alone. Genes with q ≥0.05 were discarded. Extant genes were then clustered using the Hartigan–Wong method with 25 random sets and a maximum of 1,000 iterations using the kmeans function in R stats 3.6.3. Samples were then clustered using Ward’s method and plotted using pheatmap version 1.0.12. GO term enrichment was then determined using Fisher’s exact test in topGO version 2.38.1, with org.Hs.eg.db version 3.10.0 and GO.db version 3.10.0 as references.
Weighted gene coexpression network analysis
WGCNA was performed manually using WGCNA v.1.69 with default settings unless otherwise noted60. Genes with counts >5 and detection in at least 10% of samples were included in the analysis. To best capture patterns of co-regulation, a signed network was used. Using the pickSoftThreshold function, we empirically determined a soft threshold of 7 to best fit the network structure. A minimum module size of 30 was chosen to isolate relatively large gene modules. Module eigengenes were then related back to patient and sample metadata using biweight midcorrelation. Module GO enrichment was then determined as above using Fisher’s exact test in topGO v.2.38.1, with org.Hs.eg.db v.3.10.0 as a reference. UMAP plotting was performed using uwot version 0.1.8 using the first 20 principal components of the same genes used in WGCNA analysis after z-scaling and centering, with a minimum distance of 0.2 (ref. 61). Default parameters were otherwise used.
Single-cell RNA-seq of flow cytometry-sorted BAL cells
For patients with COVID-19 we limited our analysis to samples in which flow cytometry identified distinct populations of CD206hi and CD206lo macrophages (patients 1, 2, 3, 4, 5, 7, 8, 9, A and B). We included two additional patients, one with bacterial pneumonia secondary to infection with Pseudomonas aeruginosa and Acinetobacter baumannii (assigned as ‘other pneumonia’, patient 6) and one intubated for airway protection to facilitate endoscopy for severe gastrointestinal bleeding without pneumonia (assigned as a ‘non-pneumonia control’, patient C) to examine nonspecific effects of inflammation and mechanical ventilation, respectively, on transcriptomic signatures.
Samples were enriched by flow cytometry sorting for live cells, excluding granulocytes. Cells were sorted into 2% BSA in Dulbecco’s PBS (DPBS), pelleted by centrifugation at 400g for 5 min at 4 °C, resuspended in 0.1% BSA in DPBS to about 1,000 cells μl−1. Concentration was confirmed using K2 Cellometer (Nexcelom) with AO/PI reagent and cells were loaded on 10x Genomics Chip A with Chromium Single Cell 5′ gel beads and reagents (10x Genomics) aiming to capture around 5,000–10,000 cells per library. Libraries were prepared according to the manufacturer’s protocol (10x Genomics, CG000086_RevM). After quality checks, single-cell RNA-seq libraries were pooled and sequenced on a NovaSeq 6000 instrument.
Data were processed using the Cell Ranger 3.1.0 pipeline (10x Genomics). To enable detection of viral RNA, reads were aligned to a custom hybrid genome containing GRCh38.93 and SARS-CoV-2 (NC_045512.2). An additional negative-strand transcript spanning the entirety of the SARS-CoV-2 genome was then added to the GTF and GFF files to enable detection of SARS-CoV-2 replication. Data were processed using Scanpy v.1.5.162, doublets were detected with scrublet v.0.2.163 and removed, ambient RNA was corrected with FastCAR (https://github.com/LungCellAtlas/FastCAR), and multisample integration was performed with BBKNN v.1.3.1264. Only human transcripts were used during integration, selection of highly variable genes and clustering, SARS-CoV-2 transcripts did not influence clustering. Gene set enrichment analysis was performed with signatures retrieved from the gsea-msigdb.org website65 using following terms: HALLMARK_INTERFERON_GAMMA_RESPONSE M5913, HALLMARK_INTERFERON_ALPHA_RESPONSE M5911. Computations were automated with snakemake v.5.5.466.
With the exclusion of patients A and B, single-cell RNA-seq was performed without multiplexing, using cells from a single patient per single 10x Genomics chip channel. Cells from patients A and B were split into three 10x Genomics chip channels: sample 14 contained cells from patient A, sample 15 contained cells from patient B and sample 16 contained cells from patients A and B multiplexed together. To assign cells from this sample to patients, we used souporcell v.2.067 (commit 34eade2ad3a361f045a31f53fee58c2e0c49423f) with the list of common variants for GRCh38 genome, provided on the souporcell page. We ran souporcell for samples 14, 15 and 16 with the number of clusters k = 2. We computed Pearson correlation between integer-coded single-nucleotide polymorphisms in genotypic clusters in sample pairs 14–16 and 15–16 to determine which genotypic clusters came from the same patients. Genotypic doublets and unassigned cells were discarded (see code for details).
Deconvolution of bulk RNA-seq alveolar macrophage signatures
Deconvolution of bulk RNA-seq alveolar macrophage signatures was performed using AutoGeneS v.1.0.368 and signatures derived from the integrated single-cell RNA-seq object. We used an integrated single-cell RNA-seq object containing the first 6 subjects included into analysis (patients 1–6) to train the AutoGeneS model. Signatures were automatically identified from 4,500 highly variable genes with function optimize(ngen = 200, seed = 0, nfeatures = 2000, mode = ”fixed”) (the code is available at https://github.com/NUPulmonary/2020_Grant). The model was then applied to bulk RNA-seq data to estimate the proportion of specific cell types using regression. We used bulk RNA-seq samples from healthy volunteers (which contain only tissue-resident alveolar macrophages and do not contain inflammatory monocyte-derived alveolar macrophages) to validate and optimize selection of the cluster-specific genes and deconvolution results.
RNAscope of paraffin lung slices
RNAscope Multiplex V2 manual assay from ACDbio was performed on paraffin-embedded 5-μm slices of lung tissue using mild digest times according to manufacturer instructions as we have described. Probes used were RNAscope Probe-V-nCoV2019-S-C3 (catalogue (cat.) no. 848561) with Akoya Bio Opal Dye 520 using the 488-nm laser line and RNAscope Probe-nCoV2019-orf1ab-sense-C2 (cat. no. 859151) with Opal Dye 690 using the 640-nm laser line. After the RNAscope assay was complete, slides were washed twice in TBST (1× TBS pH 7.6 with 0.1% Tween-20) for 2 min with agitation. Slides were incubated in the dark at room temperature for 30 min with 10% normal goat serum in 1× TBS with 1% BSA. The blocking solution was removed from slides via gentle flicking. Slides were then incubated in primary antibody solution containing anti-CD206 antibody (clone C-10) conjugated to AF546 (Santa Cruz Biotechnology sc-376232, RRID:AB_10989352) at 1:100 dilution in TBS with 1% BSA for 1 h at room temperature in the dark. Slides were rinsed twice with TBST for 5 min with agitation, rinsed twice in TBST buffer for 5 min and then mounted and dried overnight. Images were acquired at the Center for Advanced Microscopy at Northwestern University Feinberg School of Medicine using the Nikon W1-Spinning Disk Confocal microscope. Nucleus was added to the images using machine-based learning network trained on one patient using DAPI and brightfield images in Nikon Elements. Final images were rendered using Fiji.
Statistical analysis
Statistical analysis was performed using base R v.3.6.3 with tidyverse version 1.3.069 and Python 3.6. For all comparisons, normality was first assessed using a Shapiro–Wilk test and manual examination of distributions. For parameters that exhibited a clear lack of normality, nonparametric tests were used. In cases of multiple testing, P values were corrected using FDR correction. In Python, we used the mannwhitneyu function from scipy package v.1.3.170 for nonparametric tests, and corrected for multiple testing with the statsmodels package v.0.10.171. Adjusted P values <0.05 were considered significant. Two-sided statistical tests were performed in all cases.
Visualization
Plotting was performed in Figs. 1–3 and Extended Data Figs. 1–3 using ggplot2 v.3.3.1 unless otherwise noted. Comparisons for these figures were added using ggsignif v.0.6.0. Heat maps in Figs. 2, 3 were generated using pheatmap v.1.0.12. Sankey or Alluvial plots in Extended Data Fig. 1 were generated using ggalluvial v.0.12.072. Figure layouts for Figs. 1–4 and Extended Figs. 1, 3 were generated using patchwork v.1.01 and edited in Adobe Illustrator 2021. Figure 4 and Extended Data Figs. 4, 5 were generated with matplotlib v.3.2.173. In all box plots, box limits represent the interquartile range (IQR) with a centre line at the median. Whiskers represent the largest point within 1.5× IQR. All points are overlaid.
Study limitations
First, this is an observational study, and in the absence of a specific intervention targeting a necessary component of our model, our data are hypothesis-generating. Moreover, our observational cohorts are heterogeneous with respect to treatments received and other processes of care. Second, although we made every attempt to standardize BAL fluid volumes, the number of alveoli sampled and the return volume during both bronchoscopic and non-bronchoscopic BAL are variable. Our observations are therefore relative rather than quantitative. Third, our flow cytometry panels, although rigorous, were limited by the number of antibodies that could be used for high-volume characterization of clinical samples. Fourth, our BAL samples were collected as part of clinical care; therefore, sicker patients were more likely to be sampled. Finally, although we made every effort to standardize care for patients with SARS-CoV-2 pneumonia in our ICU, some of our patients were enrolled in clinical trials of remdesivir or sarilumab, many patients received unproven off-label therapies—including hydroxychloroquine and tocilizumab—and our study largely preceded reports on the use of steroids in patients with severe SARS-CoV-2 pneumonia (Extended Data Table 1). Despite these limitations, our systems approach to understand SARS-CoV-2 pathobiology provides a model with testable predictions that can serve as a template for the design of targeted interventions in patients with severe disease.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Bulk RNA-seq counts tables and metadata are included as Supplementary Data 2 and 3. Single-cell RNA-seq counts tables and integrated objects are available through the Gene Expression Omnibus with accession number GSE155249. Raw data are available through the Sequence Read Archive SRA/dbGaP phs002300.v1.p1.
Code availability
All code used for analysis is available at https://github.com/NUPulmonary/2020_Grant. High-level bulk and single-cell RNA-seq data can be explored via our in-house data browsers at https://www.nupulmonary.org/covid-19/.
References
Yang, X. et al. Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study. Lancet Respir. Med. 8, 475–481 (2020).
Gattinoni, L. et al. COVID-19 does not lead to a “typical” acute respiratory distress syndrome. Am. J. Respir. Crit. Care Med. 201, 1299–1300 (2020).
Baud, D. et al. Real estimates of mortality following COVID-19 infection. Lancet Infect. Dis. 20, 773 (2020).
Zhou, F. et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 395, 1054–1062 (2020).
RECOVERY Collaborative Group. Dexamethasone in hospitalized patients with Covid-19—preliminary report. N. Engl. J. Med. NEJMoa2021436 (2020).
National Heart, Lung, and Blood Institute PETAL Clinical Trials Network. Early neuromuscular blockade in the acute respiratory distress syndrome. N. Engl. J. Med. 380, 1997–2008 (2019).
Watanabe, S., Alexander, M., Misharin, A. V. & Budinger, G. R. S. The role of macrophages in the resolution of inflammation. J. Clin. Invest. 129, 2619–2628 (2019).
Kim, D. et al. The architecture of SARS-CoV-2 transcriptome. Cell 181, 914–921 (2020).
Liao, M. et al. Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat. Med. 26, 842–844 (2020).
Chua, R. L. et al. COVID-19 severity correlates with airway epithelium-immune cell interactions identified by single-cell analysis. Nat. Biotechnol. 38, 970–979 (2020).
Sola, I., Almazán, F., Zúñiga, S. & Enjuanes, L. Continuous and discontinuous RNA synthesis in coronaviruses. Annu. Rev. Virol. 2, 265–288 (2015).
Zhou, J. et al. Active replication of Middle East respiratory syndrome coronavirus and aberrant induction of inflammatory cytokines and chemokines in human macrophages: implications for pathogenesis. J. Infect. Dis. 209, 1331–1342 (2014).
Yip, M. S. et al. Antibody-dependent infection of human macrophages by severe acute respiratory syndrome coronavirus. Virol. J. 11, 82 (2014).
Cheung, C. Y. et al. Cytokine responses in severe acute respiratory syndrome coronavirus-infected macrophages in vitro: possible relevance to pathogenesis. J. Virol. 79, 7819–7826 (2005).
Soberanes, S. et al. Metformin targets mitochondrial electron transport to reduce air-pollution-induced thrombosis. Cell Metab. 29, 503 (2019).
Price, L. C., McCabe, C., Garfield, B. & Wort, S. J. Thrombosis and COVID-19 pneumonia: the clot thickens! Eur. Respir. J. 56, 2001608 (2020).
Vabret, N. et al. Immunology of COVID-19: current state of the science. Immunity 52, 910–941 (2020).
Wilk, A. J. et al. A single-cell atlas of the peripheral immune response in patients with severe COVID-19. Nat. Med. 26, 1070–1076 (2020).
Schulte-Schrepping, J. et al. Severe COVID-19 is marked by a dysregulated myeloid cell compartment. Cell 182, 1419–1440 (2020).
Habermann, A. C. et al. Single-cell RNA sequencing reveals profibrotic roles of distinct epithelial and mesenchymal lineages in pulmonary fibrosis. Sci. Adv. 6, eaba1972 (2020).
Califano, D., Furuya, Y. & Metzger, D. W. Effects of influenza on alveolar macrophage viability are dependent on mouse genetic strain. J. Immunol. 201, 134–144 (2018).
Sungnak, W. et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 26, 681–687 (2020).
Hou, Y. J. et al. SARS-CoV-2 reverse genetics reveals a variable infection gradient in the respiratory tract. Cell 182, 429–446 (2020).
Chu, H. et al. Comparative replication and immune activation profiles of SARS-CoV-2 and SARS-CoV in human lungs: an ex vivo study with implications for the pathogenesis of COVID-19. Clin. Infect. Dis. 71, 1400–1409 (2020).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045 (2020).
Konno, Y. et al. SARS-CoV-2 ORF3b is a potent interferon antagonist whose activity is increased by a naturally occurring elongation variant. Cell Rep. 32, 108185 (2020).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Jaume, M. et al. Anti-severe acute respiratory syndrome coronavirus spike antibodies trigger infection of human immune cells via a pH- and cysteine protease-independent FcγR pathway. J. Virol. 85, 10582–10597 (2011).
Martines, R. B. et al. Pathology and pathogenesis of SARS-CoV-2 associated with fatal coronavirus disease, United States. Emerg. Infect. Dis. 26, 2005–2015 (2020).
Sariol, A. & Perlman, S. Lessons for COVID-19 immunity from other coronavirus infections. Immunity 53, 248–263 (2020).
Liu, L. et al. Anti-spike IgG causes severe acute lung injury by skewing macrophage responses during acute SARS-CoV infection. JCI Insight 4, e123158 (2019).
Guilliams, M., Lambrecht, B. N. & Hammad, H. Division of labor between lung dendritic cells and macrophages in the defense against pulmonary infections. Mucosal Immunol. 6, 464–473 (2013).
Gioia, C. et al. T-Cell response profiling to biological threat agents including the SARS coronavirus. Int. J. Immunopathol. Pharmacol. 18, 525–530 (2005).
Grifoni, A. et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell 181, 1489–1501 (2020).
Le Bert, N. et al. SARS-CoV-2-specific T cell immunity in cases of COVID-19 and SARS, and uninfected controls. Nature 584, 457–462 (2020).
Mateus, J. et al. Selective and cross-reactive SARS-CoV-2 T cell epitopes in unexposed humans. Science 370, 89–94 (2020).
Braun, J. et al. SARS-CoV-2-reactive T cells in healthy donors and patients with COVID-19. Nature 587, 270–274 (2020).
Bacher, P. et al. Low avidity CD4+ T cell responses to SARS-CoV-2 in unexposed individuals and humans with severe COVID-19. Immunity 53, 1258–1271 (2020).
Oran, D. P. & Topol, E. J. Prevalence of asymptomatic SARS-CoV-2 infection: a narrative review. Ann. Intern. Med. 173, 362–367 (2020).
Neupane, A. S. et al. Patrolling alveolar macrophages conceal bacteria from the immune system to maintain homeostasis. Cell 183, 110–125 (2020).
Louie, J. K. et al. Factors associated with death or hospitalization due to pandemic 2009 influenza A(H1N1) infection in California. J. Am. Med. Assoc. 302, 1896–1902 (2009).
Leisman, D. E. et al. Cytokine elevation in severe and critical COVID-19: a rapid systematic review, meta-analysis, and comparison with other inflammatory syndromes. Lancet Respir. Med. 8, 1233–1244 (2020).
Kox, M., Waalders, N. J. B., Kooistra, E. J., Gerretsen, J. & Pickkers, P. Cytokine levels in critically ill patients with COVID-19 and other conditions. J. Am. Med. Assoc. (2020).
Walter, J. M., Helmin, K. A., Abdala-Valencia, H., Wunderink, R. G. & Singer, B. D. Multidimensional assessment of alveolar T cells in critically ill patients. JCI Insight 3, e123287 (2018).
Acute Respiratory Distress Syndrome Network. Ventilation with lower tidal volumes as compared with traditional tidal volumes for acute lung injury and the acute respiratory distress syndrome. N. Engl. J. Med. 342, 1301–1308 (2000).
Brower, R. G. et al. Higher versus lower positive end-expiratory pressures in patients with the acute respiratory distress syndrome. N. Engl. J. Med. 351, 327–336 (2004).
Guérin, C. et al. Prone positioning in severe acute respiratory distress syndrome. N. Engl. J. Med. 368, 2159–2168 (2013).
Gao, C. A. et al. Bronchoscopy on intubated COVID-19 patients is associated with low infectious risk to operators. Ann. Am. Thorac. Soc. https://doi.org/10.1513/AnnalsATS.202009-1225RL (2020).
Walter, J. M. et al. Multidimensional assessment of the host response in mechanically ventilated patients with suspected pneumonia. Am. J. Respir. Crit. Care Med. 199, 1225–1237 (2019).
Yu, Y.-R. A. et al. Flow cytometric analysis of myeloid cells in human blood, bronchoalveolar lavage, and lung tissues. Am. J. Respir. Cell Mol. Biol. 54, 13–24 (2016).
Bharat, A. et al. Flow cytometry reveals similarities between lung macrophages in humans and mice. Am. J. Respir. Cell Mol. Biol. 54, 147–149 (2016).
Charbonneau, D. H. & James, L. N. FluView and FluNet: tools for influenza activity and surveillance. Med. Ref. Serv. Q. 38, 358–368 (2019).
Di Tommaso, P. et al. Nextflow enables reproducible computational workflows. Nat. Biotechnol. 35, 316–319 (2017).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Kurtzer, G. M., Sochat, V. & Bauer, M. W. Singularity: scientific containers for mobility of compute. PLoS ONE 12, e0177459 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Lee, S. et al. NGSCheckMate: software for validating sample identity in next-generation sequencing studies within and across data types. Nucleic Acids Res. 45, e103 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Polański, K. et al. BBKNN: fast batch alignment of single cell transcriptomes. Bioinformatics 36, 964–965 (2020).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Köster, J. & Rahmann, S. Snakemake-a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Heaton, H. et al. Souporcell: robust clustering of single-cell RNA-seq data by genotype without reference genotypes. Nat. Methods 17, 615–620 (2020).
Aliee, H. & Theis, F. AutoGeneS: Automatic gene selection using multi-objective optimization for RNA-seq deconvolution. Preprint at https://doi.org/10.1101/2020.02.21.940650 (2020).
Wickham, H. et al. Welcome to the tidyverse. J. Open Source Softw. 4, 1686 (2019).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Seabold, S. & Perktold, J. Statsmodels: econometric and statistical modeling with Python. in Proc. 9th Python in Science Conference Vol. 57 (eds van der Walt, S. & Millman, J.) 92–96 (2010).
Brunson, J. ggalluvial: layered grammar for alluvial plots. J. Open Source Softw. 5, 2017 (2020).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Acknowledgements
Northwestern University Flow Cytometry Core Facility is supported by NCI Cancer Center Support Grant P30 CA060553 awarded to the Robert H. Lurie Comprehensive Cancer Center. Cell sorting was performed on a BD FACSAria SORP cell sorter purchased through the support of NIH 1S10OD011996-01. Imaging work was performed at the Northwestern University Center for Advanced Microscopy supported by NCI Cancer Center Support Grant P30 CA060553 awarded to the Robert H. Lurie Comprehensive Cancer Center. This research was supported in part through the computational resources and staff contributions provided by the Genomics Compute Cluster, which is jointly supported by the Feinberg School of Medicine, the Center for Genetic Medicine, and Feinberg’s Department of Biochemistry and Molecular Genetics, the Office of the Provost, the Office for Research and Northwestern Information Technology. The Genomics Compute Cluster is part of Quest, Northwestern University’s high-performance computing facility, with the purpose to advance research in genomics. We thank the Office for Research Safety and Institutional Biosafety Committee at Northwestern University for their help and guidance in setting up protocols and workflows for safe processing of samples from patients with COVID-19. Single-cell RNA-seq was performed with support from Simpsons Querrey Institute for Epigenetics. R.G. was supported by NIH grant T32AG020506-18. L.M.-N. was supported by T32HL076139 and F32HL151127. A.B. was supported by NIH grants HL145478, HL147290 and HL147575. C.J.G. was supported by NIH grants GM129312 and HL134800. G.R.S.B. was supported by NIH grants U19AI135964, P01AG049665, P01AG04966506S1, R01HL147575 and Veterans Affairs grant I01CX001777. A.V.M. was supported by NIH grants U19AI135964, P01AG049665, R56HL135124, R01HL153312 and NUCATS COVID-19 Rapid Response Grant. B.D.S. was supported by NIH awards K08HL128867, U19AI135964, R01HL149883 and P01AG049665. R.G.W. was supported by NIH grant U19AI135964 and a GlaxoSmithKline Distinguished Scholar in Respiratory Health grant from the CHEST Foundation.
Author information
Authors and Affiliations
Consortia
Contributions
G.R.S.B., A.V.M., B.D.S. and R.G.W. conceived and designed the project and provided funding. R.A.G., L.M.-N., N.S.M., G.R.S.B., A.V.M., B.D.S. and R.G.W. curated and analysed the data and wrote the manuscript. C.O.P., J.M.K., J.M.W., B.D.S. and R.G.W. performed adjudication of clinical outcomes. A.D., H.K.D., I.A.G., N.B. and Z.M.K. recruited patients to the study and coordinated interactions between clinical and research teams. A.C.A., A.B., B.D.S., C.O.P., E.S.M., G.R.S.B., J.M.K., J.M.W., L.M.-N., R.G.W. and S.B.S. performed BAL procedures and sample collection. A.S. contributed equipment. C.Q. performed clinical microbiological assessment of BAL fluid. J.W.L. performed autopsies on patients who died from COVID-19 and provided biological materials. M.Q. performed RNAscope assay, imaging and analysis. A.E.P., D.S., M.K. and P.N. performed programmatic extraction of clinical data and data curation. C.J.G. and E.S.M. coordinated and performed sample collection from healthy volunteers and provided biological materials. D.A.A., E.R.G., A.V.M. and S.S. processed BAL fluid samples and performed flow cytometry and cell sorting. D.A.A., H.K., L.S., R.A.G. and Z.L. performed post-sorting sample processing including RNA isolation for RNA-seq. B.D.S., A.V.M. and S.S. analysed flow cytometry data. A.V.M. and Z.L. performed single-cell RNA-seq. H.A.-V. and Y.P. performed bulk RNA-seq. All authors provided edits and feedback on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Overview of the study and biomarkers.
We compared BAL fluid obtained sequentially from 88 patients with severe SARS-CoV-2 pneumonia requiring mechanical ventilation with that from 38 patients with confirmed pneumonia secondary to other respiratory viruses (other viral pneumonia), 173 patients with non-viral pneumonia (other pneumonia) and 42 mechanically ventilated patients without pneumonia undergoing BAL (non-pneumonia controls). a, Sankey diagram illustrating steps in analysis performed for at least one BAL sample for participants with COVID-19, other viral pneumonia, non-viral pneumonia (other pneumonia), non-pneumonia controls and healthy controls. This includes samples from patients (1) enrolled in the SCRIPT study (346 patients), (2) analysed via flow cytometry (241 patients), (3) for whom bulk RNA-seq was performed on flow cytometry-sorted alveolar macrophages (181 patients) and (4) for whom single-cell RNA-seq was performed on cells from BAL fluid (10 patients with SARS-CoV-2 pneumonia, 1 patient with bacterial pneumonia and 1 patient intubated for reasons other than pneumonia (gastrointestinal bleeding requiring endoscopy, a non-pneumonia control)). Some samples were cryopreserved and sorted post-cryorecovery. Because cryopreservation affects the number of neutrophils, these samples were not included in flow cytometric analysis but were used for bulk RNA-seq profiling of flow cytometry-sorted alveolar macrophages. Samples for which flow or bulk RNA-seq analysis was skipped are represented by alluvia flowing over the grouping bars. b, Sankey diagram illustrating steps in analysis performed for all BAL samples from participants with COVID-19, other viral pneumonia, non-viral pneumonia (other pneumonia) and non-pneumonia controls. This includes samples from patients (1) enrolled in the SCRIPT study (564 samples), (2) analysed via flow cytometry (352 samples), (3) for whom bulk RNA-seq was performed on flow cytometry-sorted alveolar macrophages (232 samples) and (4) for whom single-cell RNA-seq was performed on cells from BAL fluid (12 samples). c, Self-reported smoking status. Significantly fewer active smokers were observed in the COVID-19 cohort as compared with all control groups (q < 0.05, pairwise χ2-tests of proportions with continuity and FDR correction). d–i, Biomarkers: C-reactive protein (CRP) (d), D-dimer (e), ferritin (f), procalcitonin (g), creatine phosphokinase (CPK) (h) and lactate dehydrogenase (LDH) (i). The green-shaded area indicates the normal range. j, Number of patients remaining on mechanical ventilation. k, Number of BAL samples taken per day of mechanical ventilation. l, The BAL sampling rate per day among patients with COVID-19 was not higher than the sampling rate among patients with other pneumonias.
Extended Data Fig. 2 Representative gating strategy to identify immune cell subsets in BAL samples.
a, We developed a gating strategy that enabled us to quantify immune cell populations including monocytes, alveolar macrophage subsets and T cell subsets. We defined alveolar macrophages by their expression of CD206, subdividing them into early monocyte-derived alveolar macrophages (CD206lo) and more mature (CD206hi) alveolar macrophages. T cells were identified as CD3-positive and further subdivided into CD4+ and CD8+ T cells. Treg cells were identified as CD3+CD4+CD25+CD127−. Neutrophils were identified as CD15+ cells. Monocytes were identified as HLA-DR+CD4+CD206− cells. Of note, only CD206hi alveolar macrophages were flow cytometry-sorted for bulk RNA-seq analysis (Figs. 2, 3); hence, early MoAMs (MoAM1 and MoAM2 in our single-cell RNA-seq data (Fig. 4a–h)) were not captured in bulk RNA-seq. A representative sample from a patient without neutrophilia is shown. Solid red arrows indicate direct sequential gating, dashed blue arrows indicate Boolean ‘not’ gates. Numbers on plots indicate the percentage of the parent population. Axis labels indicate laser line (UV, 355 nm; V, 405 nm; B, 488 nm; YG, 552 nm; and R, 640 nm), bandpass filter, fluorochrome and antigen/dye. b, Representative sample from a patient with neutrophilia illustrates loss of CD206hi alveolar macrophages and influx of monocyte-derived CD206lo alveolar macrophages. c, d, Contour plot and histogram overlays illustrating forward (FSC) and side scatter (SSC) properties of the CD3+ T cells (CD3), CD15+ neutrophils (CD15), monocytes, CD206lo alveolar macrophages (CD206lo) and CD206hi alveolar macrophages (CD206hi) in the representative sample from a patient without neutrophilia (c) and with neutrophilia (d). Note that neutrophils have higher side scatter than monocytes. e, Representative contour plots illustrating a sample with two distinct populations of CD206hi alveolar macrophages. Single-cell RNA-seq analysis (Fig. 4a–h) suggests that CD206hi alveolar macrophages (double-headed arrow) are bona fide tissue-resident alveolar macrophages.
Extended Data Fig. 3 At the time of intubation, the alveolar space in patients with severe SARS-CoV-2 pneumonia is enriched for T cells and monocytes and contains alveolar macrophages containing SARS-CoV-2 RNA.
a, Proportions of cells detected within 48 h of intubation (q < 0.05, pairwise Wilcoxon rank-sum tests with FDR correction). Comparisons are not significant unless otherwise noted. b, Hierarchical clustering of viral reads for SARS-CoV-2 and influenza A/California/07/2009 virus using Ward’s method; log10(DESeq2-normalized counts) are shown. c, Cumulative coverage plot of RNA-seq reads from flow cytometry-sorted alveolar macrophages aligned to the SARS-CoV-2 genome.
Extended Data Fig. 4 The BAL fluid from patients with SARS-CoV-2 pneumonia is persistently enriched for T cells irrespective of superinfection status.
a, Heat map of flow cytometry data demonstrating composition of BAL samples from all time points, grouped by diagnosis and ordered by the duration of mechanical ventilation. Column headers are colour-coded by the diagnosis, duration of mechanical ventilation (white indicates chronically ventilated patients), and presence or absence of superinfection (infection status). ‘Infection status’ refers only to the COVID-19 and ‘other viral pneumonia’ groups; blanks in these groups refer to samples for which microbiology data were incomplete and infectious status could not be determined. ‘Viral infection only’ refers to viral pathogens as the only detected pathogen in a sample, and ‘viral infection with bacterial/fungal superinfection’ refers to detection of a viral pathogen with one or more bacterial or fungal co-pathogens. b, Comparison of percentage of CD206lo and CD206hi alveolar macrophages between early (<48 h after intubation) and late (>48 h of mechanical ventilation) samples (q < 0.05, pairwise Wilcoxon rank-sum tests with FDR correction). c, Comparison of CD4+ and CD8+ T cell and neutrophil abundance in the COVID-19 group, with and without superinfection in early and late sampling. Superinfection is represented by lighter bars. Differences between groups are not significant after FDR correction.
Extended Data Fig. 5 SARS-CoV-2 pneumonia is characterized by a persistent interferon-response signature in alveolar macrophages.
a, k-means clustering of the 2,323 significantly variable genes (q < 0.05, likelihood-ratio test) across diagnosis, columns represent each individual patient, grouped by diagnosis and ordered by day from first intubation. ‘Infection status’ refers only to the COVID-19 and ‘other viral pneumonia’ groups; blanks in these groups refer to samples for which microbiology data were incomplete and infectious status could not be determined. ‘Viral infection only’ refers to viral pathogens as the only detected pathogen in a sample and ‘viral infection with bacterial/fungal superinfection’ refers to detection of a viral pathogen with one or more bacterial or fungal co-pathogens. Representative genes and GO biological processes are shown for each cluster. Column headers are colour-coded by diagnosis and duration of mechanical ventilation (white indicates chronically ventilated patients). b, Expression of selected genes between the groups. Expression of IL6 is not increased in any group. All significant comparisons are shown (q < 0.05, Wald test with FDR correction in DESeq2). c, WGCNA. d–f, Interferon-response signatures in alveolar macrophages from patients with COVID-19 gradually decrease over the course of disease. Correlation between average expression of genes from GO:0060337 type I interferon signalling pathway (R = −0.51, P = 5.7 × 10−6, Pearson correlation) (d), GO:0060333 interferon-γ-mediated signalling pathway (R = −0.22, P = 0.06, Pearson correlation) (e), module 15 of WGCNA (R = −0.40, P = 5.5 × 10−4, Pearson correlation) (f) and time on mechanical ventilation. Grey boundaries represent 95% confidence intervals. g, UMAP projections of all bulk RNA-seq samples. Average expression of WGCNA module 15 (left) and percent of CD3+ T cells in BAL (right) are shown by point area. h, Heat map demonstrating time-dependent changes in gene expression of the canonical interferon-response genes from module 15 from patients with positive outcomes (discharged home or inpatient facility), poor outcomes (discharged to a LTAC or deceased). i, Correlation between detection of SARS-CoV-2 reads and disease progression (ρ = −0.49, P = 8.3 × 10−4, Spearman correlation).
Extended Data Fig. 6 Single-cell RNA-seq identifies a positive feedback loop between IFNγ-producing T cells and SARS-CoV-2-infected alveolar macrophages.
a, Subsets of alveolar macrophages and T cells are represented by the cells from all 10 patients with COVID-19. b, IFNG expression is detected in T cells from all 10 patients with COVID-19, T cells with at least one count of IFNG were used for analysis. c, Detection of the SARS-CoV-2 negative strand. Density projection plot, with expression averaged within hexagonal areas on UMAP. d, Coverage plot of single-cell RNA-seq reads aligned to the SARS-CoV-2 genome. Cumulative data from ten individuals. Reads were aligned to genes on the positive strand or to the entire negative strand. e. UMAP plot showing integrative analysis of 105,715 cells isolated from 10 patients with severe COVID-19 within 48 h after intubation (Fig. 4a–h), one intubated patient with bacterial pneumonia and one intubated non-pneumonia control patient. f, Cells from non-pneumonia control (patient 6) and a patient with bacterial pneumonia (patient C) primarily contribute to the TRAM1 cluster and have limited contribution to MoAM clusters. g, UMAP plot showing cells from non-pneumonia control (patient 6) and a patient with bacterial pneumonia (patient C) from the integrative analysis in Extended Data Fig. 5e. h, Presence of co-infection does not affect clustering. UMAP plot showing cells from patients with and without co-infection from the integrative analysis in Fig. 4.
Extended Data Fig. 7 Deconvolution of bulk RNA-seq demonstrates loss of tissue-resident alveolar macrophages and persistence of mature monocyte-derived alveolar macrophages in patients with severe COVID-19.
a, Dot plot showing IL6 expression across cell types. Dot size is proportional to the number of cells expressing IL6 in the corresponding cluster. Data from the present study (G) are presented per patient, data from ref. 9 (L) and ref. 20 (H.) are averaged by condition. b, Heat map demonstrating proportion of alveolar macrophage subsets predicted from deconvolution analysis. Data are grouped by condition and ordered by proportion of CD206hi alveolar macrophages. c, Proportion of alveolar macrophage cell types in patients with COVID-19 in comparison to other types of pneumonia obtained from deconvolution analysis (pairwise Wilcoxon rank-sum tests with FDR correction; *q < 0.05, **q < 0.01, ***q < 0.001).
Supplementary information
Supplemental Data File 1
Data related to k-means clustering and differential expression analysis (DEA) of bulk RNA-seq samples: gene cluster assignments, cluster GO enrichment, pairwise DEA across diagnosis, and WGCNA module assignments. Excel workbook, multiple tabs.
Supplemental Data File 2
Raw bulk RNA-seq gene counts. CSV file.
Supplemental Data File 3
Bulk RNA-seq metadata. CSV file.
Supplemental Data File 4
Marker genes for integrated scRNA-seq object containing cells from 10 patients with COVID-19. CSV file.
Supplemental Data File 5
Marker genes for integrated scRNA-seq object containing cells from 10 patients with COVID-19, one patient with bacterial pneumonia and one intubated non-pneumonia control. CSV file.
Supplemental Data File 6
Differentially expressed genes between TRAM1 and TRAM2 clusters. CSV file.
Supplemental Data File 7
SARS-CoV-2 infection is spatially restricted. Combined immunofluorescence microscopy for CD206, a marker of mature macrophages (red), and smFISH (RNAscope) for positive- (yellow) and negative-strand (cyan) SARS-CoV-2 transcripts.
Supplemental Data File 8
Standardized tool for pneumonia category adjudication.
Supplemental Data File 9
Cell-type percentages from flow cytometry analysis, including metadata for flow cytometry, bulk RNA-seq, and single-cell RNA-seq data.
Supplemental Data Video 1
Detection of SARS-CoV-2 positive and negative strand transcripts in a CD206-positive alveolar macrophage. Combined immunofluorescence microscopy for CD206, a marker of mature macrophages (red), and smFISH (RNAscope) for positive- (yellow) and negative-strand (cyan) SARS-CoV-2 transcripts.
Rights and permissions
About this article
Cite this article
Grant, R.A., Morales-Nebreda, L., Markov, N.S. et al. Circuits between infected macrophages and T cells in SARS-CoV-2 pneumonia. Nature 590, 635–641 (2021). https://doi.org/10.1038/s41586-020-03148-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-03148-w
This article is cited by
-
Exosomal secreted SCIMP regulates communication between macrophages and neutrophils in pneumonia
Nature Communications (2024)
-
Asialoglycoprotein receptor 1 promotes SARS-CoV-2 infection of human normal hepatocytes
Signal Transduction and Targeted Therapy (2024)
-
Alveolar macrophage-expressed Plet1 is a driver of lung epithelial repair after viral pneumonia
Nature Communications (2024)
-
ACE2-dependent and -independent SARS-CoV-2 entries dictate viral replication and inflammatory response during infection
Nature Cell Biology (2024)
-
CircRNA: a rising therapeutic strategy for lung injury induced by pulmonary toxicants
Archives of Toxicology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.