Abstract
The hippocampus is an important part of the limbic system in the human brain that has essential roles in spatial navigation and the consolidation of information from short-term memory to long-term memory1,2. Here we use single-cell RNA sequencing and assay for transposase-accessible chromatin using sequencing (ATAC–seq) analysis to illustrate the cell types, cell linage, molecular features and transcriptional regulation of the developing human hippocampus. Using the transcriptomes of 30,416 cells from the human hippocampus at gestational weeks 16–27, we identify 47 cell subtypes and their developmental trajectories. We also identify the migrating paths and cell lineages of PAX6+ and HOPX+ hippocampal progenitors, and regional markers of CA1, CA3 and dentate gyrus neurons. Multiomic data have uncovered transcriptional regulatory networks of the dentate gyrus marker PROX1. We also illustrate spatially specific gene expression in the developing human prefrontal cortex and hippocampus. The molecular features of the human hippocampus at gestational weeks 16–20 are similar to those of the mouse at postnatal days 0–5 and reveal gene expression differences between the two species. Transient expression of the primate-specific gene NBPF1 leads to a marked increase in PROX1+ cells in the mouse hippocampus. These data provides a blueprint for understanding human hippocampal development and a tool for investigating related diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The scRNA-seq data and ATAC-seq data used in this study have been deposited in the Gene Expression Omnibus (GEO) under accession number GSE131258. Raw image files used in the figures that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Bird, C. M. & Burgess, N. The hippocampus and memory: insights from spatial processing. Nat. Rev. Neurosci. 9, 182–194 (2008).
Miller, A. M., Vedder, L. C., Law, L. M. & Smith, D. M. Cues, context, and long-term memory: the role of the retrosplenial cortex in spatial cognition. Front. Hum. Neurosci. 8, 586 (2014).
Lavado, A., Lagutin, O. V., Chow, L. M., Baker, S. J. & Oliver, G. Prox1 is required for granule cell maturation and intermediate progenitor maintenance during brain neurogenesis. PLoS Biol. 8, e1000460 (2010).
Sugiyama, T., Osumi, N. & Katsuyama, Y. The germinal matrices in the developing dentate gyrus are composed of neuronal progenitors at distinct differentiation stages. Dev. Dyn. 242, 1442–1453 (2013).
Galceran, J., Miyashita-Lin, E. M., Devaney, E., Rubenstein, J. L. R. & Grosschedl, R. Hippocampus development and generation of dentate gyrus granule cells is regulated by LEF1. Development 127, 469–482 (2000).
Lee, S. M. K., Tole, S., Grove, E. & McMahon, A. P. A local Wnt-3a signal is required for development of the mammalian hippocampus. Development 127, 457–467 (2000).
Zhong, S. J. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Mu, L. et al. SoxC transcription factors are required for neuronal differentiation in adult hippocampal neurogenesis. J. Neurosci. 32, 3067–3080 (2012).
Wang, Y., Lin, L., Lai, H., Parada, L. F. & Lei, L. Transcription factor Sox11 is essential for both embryonic and adult neurogenesis. Dev. Dyn. 242, 638–653 (2013).
Bandler, R. C., Mayer, C. & Fishell, G. Cortical interneuron specification: the juncture of genes, time and geometry. Curr. Opin. Neurobiol. 42, 17–24 (2017).
Pollen, A. A. et al. Molecular identity of human outer radial glia during cortical development. Cell 163, 55–67 (2015).
Cooper, A. et al. Trisomy of the G protein-coupled K+ channel gene, Kcnj6, affects reward mechanisms, cognitive functions, and synaptic plasticity in mice. Proc. Natl Acad. Sci. USA 109, 2642–2647 (2012).
Gomez Perdiguero, E., Schulz, C. & Geissmann, F. Development and homeostasis of “resident” myeloid cells: the case of the microglia. Glia 61, 112–120 (2013).
Hochgerner, H., Zeisel, A., Lönnerberg, P. & Linnarsson, S. Conserved properties of dentate gyrus neurogenesis across postnatal development revealed by single-cell RNA sequencing. Nat. Neurosci. 21, 290–299 (2018).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Dumas, L. J. et al. DUF1220-domain copy number implicated in human brain-size pathology and evolution. Am. J. Hum. Genet. 91, 444–454 (2012).
Mangale, V. S. et al. Lhx2 selector activity specifies cortical identity and suppresses hippocampal organizer fate. Science 319, 304–309 (2008).
Grove, E. A., Tole, S., Limon, J., Yip, L. & Ragsdale, C. W. The hem of the embryonic cerebral cortex is defined by the expression of multiple Wnt genes and is compromised in Gli3-deficient mice. Development 125, 2315–2325 (1998).
Yoshida, M., Assimacopoulos, S., Jones, K. R. & Grove, E. A. Massive loss of Cajal-Retzius cells does not disrupt neocortical layer order. Development 133, 537–545 (2006).
Kempermann, G., Song, H. & Gage, F. H. Neurogenesis in the adult hippocampus. Cold Spring Harb. Perspect. Biol. 7, a018812 (2015).
Ortiz-Matamoros, A., Salcedo-Tello, P., Avila-Muñoz, E., Zepeda, A. & Arias, C. Role of wnt signaling in the control of adult hippocampal functioning in health and disease: therapeutic implications. Curr. Neuropharmacol. 11, 465–476 (2013).
Varela-Nallar, L. & Inestrosa, N. C. Wnt signaling in the regulation of adult hippocampal neurogenesis. Front. Cell. Neurosci. 7, 100 (2013).
Berg, D. A. et al. A common embryonic origin of stem cells drives developmental and adult neurogenesis. Cell 177, 654–668.e15 (2019).
Davis, J. M. et al. DUF1220 dosage is linearly associated with increasing severity of the three primary symptoms of autism. PLOS Genet. 10, e1004241 (2014).
Keeney, J. G., Dumas, L. & Sikela, J. M. The case for DUF1220 domain dosage as a primary contributor to anthropoid brain expansion. Front. Hum. Neurosci. 8, 427 (2014).
Popesco, M. C. et al. Human lineage-specific amplification, selection, and neuronal expression of DUF1220 domains. Science 313, 1304–1307 (2006).
Huang, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Tripathi, S. et al. Meta- and orthogonal integration of influenza “OMICs” data defines a role for UBR4 in virus budding. Cell Host Microbe 18, 723–735 (2015).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Langfelder, P. & Horvath, S. Fast R functions for robust correlations and hierarchical clustering. J. Stat. Softw. 46, i11 (2012).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: A method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21–29 (2015).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Palop, J. J., Roberson, E. D. & Cobos, I. Step-by-step in situ hybridization method for localizing gene expression changes in the brain. Methods Mol. Biol. 670, 207–230 (2011).
Wang, X., Tsai, J. W., LaMonica, B. & Kriegstein, A. R. A new subtype of progenitor cell in the mouse embryonic neocortex. Nat. Neurosci. 14, 555–561 (2011).
Acknowledgements
This work was supported by the National Key R&D Program of China (2019YFA0110100), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDA16020601, XDB32010100), the National Basic Research Program of China (2017YFA0102601, 2017YFA0103303), the National Natural Science Foundation of China (NSFC) (91732301, 31671072, 31771140, 81891001), the Grants of Shanghai Brain-Intelligence Project from STCSM (16JC1420500), the Grants of Beijing Brain Initiative of Beijing Municipal Science & Technology Commission (Z181100001518004).
Author information
Authors and Affiliations
Contributions
Q.W. and X.W. conceived the project, designed the experiments and wrote the manuscript. S. Zhong and Y.L. performed the scRNA-seq experiment. W.D. performed the ATAC-seq and animal surgery. S. Zhong, Y.L., X.F., S. Zhang and F.T. analysed the RNA-seq data. S. Zhong and H.D. analysed the ATAC-seq data. L.S. and R.C. collected single cells by patch-clamping. Z.L. and S. Zhong carried out qRT–PCR. Q.M. prepared the samples. S. Zhong, Q.W. and W.D. performed immunostaining, in situ hybridization and imaging. All authors edited and proofread the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Joseph Loturco, Christopher Walsh and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
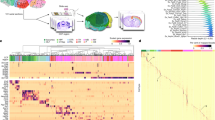
Extended Data Fig. 1 Single-cell RNA-seq information and molecular diversity of single cells.
a, Scheme of bioinformatic analysis. b, Expression of known markers shown using the same layout as in Fig. 1b. Grey, no expression; red, relative expression. c, Heat map showing the expression level and identity of genes in all cells in the developing hippocampus. Sample sizes: astrocytes, 703; Cajal–Retzius cells, 101; endothelial cells, 540; non-DG ExN, 8,199; MGE-derived InN, 6,377; microglia, 2,660; oligodendrocytes, 209; OPCs, 1,250; progenitors, 2,486; DG ExN, 2,516; CGE-derived InN, 5,375. d, t-SNE plots of cells in the hippocampus. Two repetitions of GW22 are labelled in different shapes, and no obvious distribution differences are observed among the different batches from the same embryo stages. Each cell colour represents the gestational week. Sample size: GW16, 4,411 cells; GW18, 4,035 cells; GW20, 10,101 cells; GW22#01, 1,617 cells; GW22#02, 2,485 cells; GW25, 2,824 cells; GW27, 4,943 cells.
Extended Data Fig. 2 Molecular diversity of subgroups of cells.
a–c, Heat maps show the subclasses of inhibitory neurons (a), astrocytes (b) and oligodendrocytes (c). The genes are organized into clusters. The bar chart on the top shows the gestational week. Specific genes related to each subtype are highlighted on the right with enriched GO terms. Interneurons: 3,189, 2,334, 909, 670, 1,765, 1,073, 1,019, 793 cells; astrocytes: 275, 95, 141, 134, 58 cells; oligodendrocytes: 103, 227, 131, 282, 257, 459 cells. d, The enriched GO terms show the cell properties of the hippocampus in different cell types. Progenitors, 2,486 cells; excitatory neurons, 10,715 cells. e, Immunostaining for oligodendrocyte markers at GW16 showing the position ands morphology of oligodendrocytes in human prefrontal cortex. Scale bar, 500 μm. The experiment was repeated three times independently with similar results.
Extended Data Fig. 3 Molecular diversity of progenitors in the hippocampus.
a, Heat map showing the expression levels and identities of genes in the progenitor subclasses. Known gene expression in each type and GO enrichments are shown to the right. The graph above shows the distribution of each subclass by gestational week, and the graph below shows the subclusters of progenitors. Clusters 1–8: 204, 159, 483, 730, 397, 300, 139 and 74 cells. b, Dot plot for known markers of subtypes of progenitors in Fig. 2b. The size of each dot represents the percentage of cells in each cluster. Grey-to-blue gradient shows low-to-high gene expression. Progenitors: 204, 159, 483, 730, 397, 300, 139 and 74 cells. c, Dot plot for novel markers of subtypes of progenitors. The size of each dot represents the percentage of cells in each cluster. Grey-to-red gradient shows low-to-high gene expression. Progenitors: 204, 159, 483, 730, 397, 300, 139 and 74 cells. d, Abstracted graph shows the connection on the transcriptome of different subtypes in the developing human hippocampus. Each dot represents a single cell, and cell colour represents the cell type. e, Abstracted graph shows the connection on the transcriptome of all subtypes in the developing human hippocampus. Each dot represents a single cell, and cell colour represents the cell type. f, Abstracted graph shows the connection on the transcriptome of different weeks in the developing human hippocampus. Each dot represents a single cell, and cell colour represents the week. g, h, Visualization of eight subtypes of progenitors in the developing human hippocampus using t-SNE (g), and expression of known markers using the same layout (h). Grey, no expression; red, relative expression.
Extended Data Fig. 4 Immunostaining of progenitors in the developing hippocampus.
a, Immunofluorescence images of PAX6 and SOX2 at GW11. Scale bar, 2,000 μm. b, Immunofluorescence images of HOPX and SOX2 at GW11. Scale bar, 2,000 μm. c, Immunofluorescence images of PROX1, PAX6, HOPX and SOX2 at GW14. Scale bar, 1,000 μm. d–i, Immunofluorescence images of PROX1, PAX6, HOPX and SOX2 in GW16 (d–f) and GW22 (g–i). Scale bar, 500 μm. The experiment was repeated three times independently with similar results.
Extended Data Fig. 5 Immunostaining of developing hippocampus.
a, b, Immunofluorescence images of PAX6, HOPX and MKI67 at GW25. Scale bar, 500 μm. c, d, Cell cycle analysis of PAX6+ (c) or HOPX+ (d) progenitors. e, Immunofluorescence images of PROX1 in GW25 to show granule cell layer. Scale bars, 500 μm (left); 100 μm (right, panels 1–3). f–i, Immunofluorescence images of PAX6, HOPX, NEUROD1 and GFAP at GW25. Scale bar, 500 μm. The experiment was repeated three times independently with similar results. j, k, The maturation scores of PAX6+ (j) and HOPX+ (k) progenitors.
Extended Data Fig. 6 Molecular diversity of excitatory neurons in the hippocampus.
a, b, Expression of known markers (a) and new markers (b) shown using the same layout as in Fig. 3a. Grey, no expression; red, relative expression. c, Cluster dendrogram showing the modules selected to calculate the gene network in Fig. 3h. d, The cluster trees and heat map show the correlation of different gene modules in excitatory neurons.
Extended Data Fig. 7 Molecular diversity of inhibitory neurons in the hippocampus.
a, GO analysis of modules created by clustering the two main branches from the lineage tree. The analysis reflects cell fate commitment. In this heat map, the middle represents the start of pseudo-time. From this point, one lineage moves to the CA and the other moves to the DG. Rows are GO terms correlated into different modules. Sample size: 12,115 cells. b, c, Expression of known markers shown using the same layout as in Fig. 3k. d, Expression of novel markers of MGE-derived inhibitory neurons shown using the same layout as in Fig. 3k. Grey, no expression; red, relative expression. e, Expression of novel markers of CGE-derived inhibitory neurons shown using the same layout as in Fig. 3k.
Extended Data Fig. 8 Molecular diversity of microglia in the human hippocampus.
a, Heat map showing the expression levels and identities of genes in the microglia subclasses. The graph above shows the distribution of each subclass by gestational week. b, Visualization of ten subtypes of microglia in the developing human hippocampus using t-SNE. Each dot represents a single cell, and cells are laid out to show similarities. Each cell colour represents the cell type. Expression of known markers is shown using the same layout on the right; grey, no expression; red, relative expression. Microglia: 638, 489, 246, 259, 465, 229, 84, 84, 68, 54 and 44 cells. c, d, Distribution of G1, S, and G2/M stages of the cell cycle for microglia of different subtypes (c) and at different gestational weeks (d). e, Immunostaining images of PTPRC and MKI67 at GW25. Scale bars, 500 μm (left), 100 μm (right). The experiment was repeated three times independently with similar results.
Extended Data Fig. 9 NBPF family genes in the human hippocampus.
a, Expression of NBPF family genes shown using the same layout as in Fig. 1b. Grey, no expression; blue, relative expression. b, Domains of NBPF1. c, Evolutionary history inferred using the neighbour-joining method. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Poisson correction method and are in the units of the number of amino acid substitutions per site. The analysis involved six amino acid sequences. Evolutionary analyses were conducted in MEGA X. d, Overexpression of NBPF1 promotes DG formation at E13.5, and is observed at E15.5 in mouse. Scale bars, 500 μm. The experiment was repeated six times independently with similar results. e, Scheme depicting the position in the mouse brain at E18.5 of the slice in f. f, Overexpression of NBPF1 promotes DG formation at E13.5, and this is observed at E18.5 in mouse. Scale bars, 1,000 μm. The experiment was repeated six times independently with similar results. g, Flow chart of patch-qRT–PCR. h, Relative expression of specific genes of GFP+ cells. **P = 0.0020, *P = 0.0408, two-sided t-test; n = 10 GFP cells; 8 NBPF1–GFP cells. Mean ± s.e.m. i, Normalized ATAC-seq profiles of PROX1 in GW25 hippocampus with three independent biological replicates (Rep1, Rep2 and Rep3) showing the activation of PROX1. The amplifying panel shows the predicted LHX2 binding sites.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Zhong, S., Ding, W., Sun, L. et al. Decoding the development of the human hippocampus. Nature 577, 531–536 (2020). https://doi.org/10.1038/s41586-019-1917-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1917-5
This article is cited by
-
SRT-Server: powering the analysis of spatial transcriptomic data
Genome Medicine (2024)
-
Drug targeting in psychiatric disorders — how to overcome the loss in translation?
Nature Reviews Drug Discovery (2024)
-
Genetics of human brain development
Nature Reviews Genetics (2024)
-
Multimodal Nature of the Single-cell Primate Brain Atlas: Morphology, Transcriptome, Electrophysiology, and Connectivity
Neuroscience Bulletin (2024)
-
A Review of the Application of Spatial Transcriptomics in Neuroscience
Interdisciplinary Sciences: Computational Life Sciences (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.