Abstract
Class B G-protein-coupled receptors are major targets for the treatment of chronic diseases, including diabetes and obesity1. Structures of active receptors reveal peptide agonists engage deep within the receptor core, leading to an outward movement of extracellular loop 3 and the tops of transmembrane helices 6 and 7, an inward movement of transmembrane helix 1, reorganization of extracellular loop 2 and outward movement of the intracellular side of transmembrane helix 6, resulting in G-protein interaction and activation2,3,4,5,6. Here we solved the structure of a non-peptide agonist, TT-OAD2, bound to the glucagon-like peptide-1 (GLP-1) receptor. Our structure identified an unpredicted non-peptide agonist-binding pocket in which reorganization of extracellular loop 3 and transmembrane helices 6 and 7 manifests independently of direct ligand interaction within the deep transmembrane domain pocket. TT-OAD2 exhibits biased agonism, and kinetics of G-protein activation and signalling that are distinct from peptide agonists. Within the structure, TT-OAD2 protrudes beyond the receptor core to interact with the lipid or detergent, providing an explanation for the distinct activation kinetics that may contribute to the clinical efficacy of this compound series. This work alters our understanding of the events that drive the activation of class B receptors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All relevant data are available from the authors and/or included in the manuscript or Supplementary Information. Atomic coordinates and the cryo-EM density map have been deposited in the Protein Data Bank (PDB) under accession number 6ORV and Electron Microscopy Data Bank (EMDB) accession EMD-20179.
References
Bortolato, A. et al. Structure of class B GPCRs: new horizons for drug discovery. Br. J. Pharmacol. 171, 3132–3145 (2014).
Liang, Y. L. et al. Cryo-EM structure of the active, Gs-protein complexed, human CGRP receptor. Nature 561, 492–497 (2018).
Liang, Y. L. et al. Phase-plate cryo-EM structure of a biased agonist-bound human GLP-1 receptor-Gs complex. Nature 555, 121–125 (2018).
Liang, Y. L. et al. Phase-plate cryo-EM structure of a class B GPCR-G-protein complex. Nature 546, 118–123 (2017).
Zhao, L. H. et al. Structure and dynamics of the active human parathyroid hormone receptor-1. Science 364, 148–153 (2019).
Zhang, Y. et al. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein. Nature 546, 248–253 (2017).
Siu, F. Y. et al. Structure of the human glucagon class B G-protein-coupled receptor. Nature 499, 444–449 (2013).
Zhang, H. et al. Structure of the full-length glucagon class B G-protein-coupled receptor. Nature 546, 259–264 (2017).
Zhang, H. et al. Structure of the glucagon receptor in complex with a glucagon analogue. Nature 553, 106–110 (2018).
Song, G. et al. Human GLP-1 receptor transmembrane domain structure in complex with allosteric modulators. Nature 546, 312–315 (2017).
Htike, Z. Z. et al. Efficacy and safety of glucagon-like peptide-1 receptor agonists in type 2 diabetes: A systematic review and mixed-treatment comparison analysis. Diabetes Obes. Metab. 19, 524–536 (2017).
Aroda, V. R. et al. PIONEER 1: Randomized clinical trial comparing the efficacy and safety of oral semaglutide monotherapy with placebo in patients with type 2 diabetes. Diabetes Care 42, dc190749 (2019).
Pratley, R. et al. Oral semaglutide versus subcutaneous liraglutide and placebo in type 2 diabetes (PIONEER 4): a randomised, double-blind, phase 3a trial. Lancet 394, 39–50 (2019).
Wootten, D. et al. Differential activation and modulation of the glucagon-like peptide-1 receptor by small molecule ligands. Mol. Pharmacol. 83, 822–834 (2013).
Nolte, W. M. et al. A potentiator of orthosteric ligand activity at GLP-1R acts via covalent modification. Nat. Chem. Biol. 10, 629–631 (2014).
Freeman, J. W. S. et al. TTP273: Oral, G protein pathway selective clinical-stage GLP-1 receptor (GLP-1R) agonist. Keystone Symposium, Keystone, CO February 22 http://vtvtherapeutics.com/wp-content/uploads/pdf/GLP1_poster_animation_Keystone_CO_FINAL.pdf (2016).
Chen, D. et al. A nonpeptidic agonist of glucagon-like peptide 1 receptors with efficacy in diabetic db/db mice. Proc. Natl Acad. Sci. USA 104, 943–948 (2007).
Jazayeri, A. et al. Crystal structure of the GLP-1 receptor bound to a peptide agonist. Nature 546, 254–258 (2017).
Freeman, J. D. C., Dunn, I. & Valcarce, C. TTP273, Oral (nonpeptide) GLP-1R agonist: improved glycemic control without nausea and vomiting in phase 2. American Diabetes Association 77th Scientific Sessions, San Diego, CA, June 9–13, 2017 http://vtvtherapeutics.com/wp-content/uploads/pdf/ADA_Logra_study_results_poster_1220-P%20Final.pdf (2017).
Runge, S., Thøgersen, H., Madsen, K., Lau, J. & Rudolph, R. Crystal structure of the ligand-bound glucagon-like peptide-1 receptor extracellular domain. J. Biol. Chem. 283, 11340–11347 (2008).
DeVree, B. T. et al. Allosteric coupling from G protein to the agonist-binding pocket in GPCRs. Nature 535, 182–186 (2016).
Yin, Y. et al. An intrinsic agonist mechanism for activation of glucagon-like peptide-1 receptor by its extracellular domain. Cell Discov. 2, 16042 (2016).
Zhao, L. H. et al. Differential requirement of the extracellular domain in activation of class B G protein-coupled receptors. J. Biol. Chem. 291, 15119–15130 (2016).
Ehrenmann, J. et al. High-resolution crystal structure of parathyroid hormone 1 receptor in complex with a peptide agonist. Nat. Struct. Mol. Biol. 25, 1086–1092 (2018).
Yang, L. et al. Conformational states of the full-length glucagon receptor. Nat. Commun. 6, 7859 (2015).
Wootten, D., Simms, J., Miller, L. J., Christopoulos, A. & Sexton, P. M. Polar transmembrane interactions drive formation of ligand-specific and signal pathway-biased family B G protein-coupled receptor conformations. Proc. Natl Acad. Sci. USA 110, 5211–5216 (2013).
dal Maso, E. et al. The molecular control of calcitonin receptor signaling. ACS Pharmacol. Transl. Sci. 2, 31–51 (2019).
Wootten, D. et al. The extracellular surface of the GLP-1 receptor is a molecular trigger for biased agonism. Cell 165, 1632–1643 (2016).
Lei, S. et al. Two distinct domains of the glucagon-like peptide-1 receptor control peptide-mediated biased agonism. J. Biol. Chem. 293, 9370–9387 (2018).
Liang, Y.-L. et al. Dominant negative G proteins enhance formation and purification of agonist-GPCR-G protein complexes for structure determination. ACS Pharmacol. Transl. Sci. 1, 9 (2018).
Furness, S. G. B. et al. Ligand-dependent modulation of G protein conformation alters drug efficacy. Cell 167, 739–749.e711 (2016).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Nakane, T., Kimanius, D., Lindahl, E. & Scheres, S. H. Characterisation of molecular motions in cryo-EM single-particle data by multi-body refinement in RELION. eLife 7, e36861 (2018).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Chan, K. Y., Trabuco, L. G., Schreiner, E. & Schulten, K. Cryo-electron microscopy modeling by the molecular dynamics flexible fitting method. Biopolymers 97, 678–686 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Moriarty, N. W., Grosse-Kunstleve, R. W. & Adams, P. D. electronic Ligand Builder and Optimization Workbench (eLBOW): a tool for ligand coordinate and restraint generation. Acta Crystallogr. D 65, 1074–1080 (2009).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Jacobson, M. P. et al. A hierarchical approach to all-atom protein loop prediction. Proteins 55, 351–367 (2004).
Eswar, N. et al. Comparative protein structure modeling using Modeller. Curr Protoc Bioinformatics Chapter 5, Unit 5.6 (2006).
Vohra, S. et al. Similarity between class A and class B G-protein-coupled receptors exemplified through calcitonin gene-related peptide receptor modelling and mutagenesis studies. J. R. Soc. Interface 10, 20120846 (2012).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph 14, 33–38 (1996).
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011).
Carpenter, B., Nehmé, R., Warne, T., Leslie, A. G. & Tate, C. G. Structure of the adenosine A2A receptor bound to an engineered G protein. Nature 536, 104–107 (2016).
Huang, J. & MacKerell, A. D. Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).
Doerr, S., Harvey, M. J., Noé, F. & De Fabritiis, G. HTMD: high-throughput molecular dynamics for molecular discovery. J. Chem. Theory Comput. 12, 1845–1852 (2016).
Dolinsky, T. J., Nielsen, J. E., McCammon, J. A. & Baker, N. A. PDB2PQR: an automated pipeline for the setup of Poisson–Boltzmann electrostatics calculations. Nucleic Acids Res. 32, W665–W667 (2004).
Olsson, M. H., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pka predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Lomize, M. A., Lomize, A. L., Pogozheva, I. D. & Mosberg, H. I. OPM: orientations of proteins in membranes database. Bioinformatics 22, 623–625 (2006).
Sommer, B. Membrane packing problems: a short review on computational membrane modeling methods and tools. Comput. Struct. Biotechnol. J. 5, e201302014 (2013).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Vanommeslaeghe, K. & MacKerell, A. D. Jr. Automation of the CHARMM General Force Field (CGenFF) I: bond perception and atom typing. J. Chem. Inf. Model. 52, 3144–3154 (2012).
Vanommeslaeghe, K., Raman, E. P. & MacKerell, A. D. Jr. Automation of the CHARMM General Force Field (CGenFF) II: assignment of bonded parameters and partial atomic charges. J. Chem. Inf. Model. 52, 3155–3168 (2012).
Harvey, M. J., Giupponi, G. & Fabritiis, G. D. ACEMD: Accelerating Biomolecular Dynamics in the Microsecond Time Scale. J. Chem. Theory Comput. 5, 1632–1639 (2009).
Loncharich, R. J., Brooks, B. R. & Pastor, R. W. Langevin dynamics of peptides: the frictional dependence of isomerization rates of N-acetylalanyl-N'-methylamide. Biopolymers 32, 523–535 (1992).
Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F., DiNola, A. & Haak, J. R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684 (1984).
Kräutler, V. G., van Gunsteren, W. F. & Hünenberger, P. H. A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. J. Comput. Chem. 22, 501–508 (2001).
Essmann, U. P. & Berkowitz, L. M.L. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577 (1995).
Cuzzolin, A., Deganutti, G., Salmaso, V., Sturlese, M. & Moro, S. AquaMMapS: an alternative tool to monitor the role of water molecules during protein-ligand association. ChemMedChem 13, 522–531 (2018).
Wall, M. E., Calabró, G., Bayly, C. I., Mobley, D. L. & Warren, G. L. Biomolecular solvation structure revealed by molecular dynamics simulations. J. Am. Chem. Soc. 141, 4711–4720 (2019).
Koole, C. et al. Allosteric ligands of the glucagon-like peptide 1 receptor (GLP-1R) differentially modulate endogenous and exogenous peptide responses in a pathway-selective manner: implications for drug screening. Mol. Pharmacol. 78, 456–465 (2010).
Savage, E. E., Wootten, D., Christopoulos, A., Sexton, P. M. & Furness, S. G. A simple method to generate stable cell lines for the analysis of transient protein-protein interactions. Biotechniques 54, 217–221 (2013).
Jun, L. S. et al. A novel humanized GLP-1 receptor model enables both affinity purification and Cre-LoxP deletion of the receptor. PLoS One 9, e93746 (2014).
Acknowledgements
The work was supported by the Monash University Ramaciotti Centre for Cryo-Electron Microscopy, the Monash MASSIVE high-performance computing facility, the National Health and Medical Research Council of Australia (NHMRC) project grants (1061044, 1065410, 1120919 and 1126857) and NHMRC program grants (1055134 and 1150083), the Japan Society for the Promotion of Science (JSPS) KAKENHI no. 18H06043 and Japan Science and Technology Agency (JST) PRESTO no. 18069571 (to R.D.). P.M.S. and A.C. are NHMRC Senior Principal Research Fellows and D.W. is an NHMRC Senior Research Fellow. S.G.B.F. is an ARC Future Fellow. A.I. was funded by the PRIME JP17gm5910013 and the LEAP JP17gm0010004 from the Japan Agency for Medical Research and Development, and JSPS KAKENHI 17K08264. We are grateful to G. Christopoulos, V. Julita, T. Fields, C. Lafuente, J. M. Minguez, G. C. Sanz and F. Qu for assay and technical support.
Author information
Authors and Affiliations
Contributions
P.Z. designed and performed most of the pharmacological studies with assistance from T.T.T.; Y.-L.L. expressed and purified the complex; R.D. performed cryo-sample preparation and imaging to acquire electron microscopy data; M.J.B. and R.D. processed the electron microscopy data and performed electron microscopy map calculations; M.J.B. built the model and performed refinement; M.M.F. performed the mutagenesis studies, L.C. performed studies in the HEK293 CRISPR-knockout cells; G.D. and C.A.R. designed, performed and analysed the molecular dynamics simulations; F.S.W. and M.G.B. provided TT-OAD2. M.E.C., M.G.B. and K.W.S. designed and oversaw the in vivo studies; P.Z., Y.-L.L., M.J.B., G.D., C.A.R., F.S.W., K.W.S., R.D., P.M.S. and D.W. performed data analysis; P.Z., Y.-L.L., M.J.B., G.D., C.A.R., F.S.W., K.W.S., A.C., L.J.M., M.-W.W. and R.D. assisted with data interpretation, figure and manuscript preparation; P.M.S. and D.W. designed and supervised the project, interpreted the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
F.W.S., M.E.C. and K.W.S. are employees of Eli Lilly and Company.
Additional information
Peer review information Nature thanks Doryen Bubeck, Dave D’Alessio, Nita R. Shah and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Binding, transducer coupling and signalling mediated by TT-OAD2.
a, Kinetic ligand-binding assay using ROX-exendin-4 as the fluorescent probe. TT-OAD2 is only able to partially displace the probe and with slower kinetics relative to exendin-4 that shows complete displacement of the probe with rapid kinetics. b, cAMP accumulation studies using GLP-1 and TT-OAD2 as the agonist in wild-type HEK293 cells and HEK293 cells in which Gs/olf (ΔGs) or all Gi/o/z (ΔGi/o/z) have been depleted using CRISPR–Cas9. c, HEK293A cells transiently transfected with the GLP-1R and the NanoBit constructs for Gαs and Gαi2 (Gα-LgBIT, Gγ2-SmBIT). Luminescence signal was assessed over time (0–20 min) in the presence of increasing concentrations of GLP-1 and TT-OAD2. Concentration response curves are expressed as AUC (0–20 min) for each concentration and normalized to the negative response observed by GLP-1 at 1 μM. d, Agonist-induced changes in trimeric Gs protein conformation. Ligand-induced changes in BRET were measured in plasma membrane preparations performed in kinetic mode until kinetic equilibrium was reached for vehicle or increasing concentrations of GLP-1 (left) and TT-OAD2 (right). The addition of GTP dissociated the trimeric G protein complex stabilized by GLP-1-occupied and TT-OAD2-occupied GLP-1R. e, Agonist-induced changes in trimeric Gi2 protein conformation. Left, ligand-induced changes in BRET were measured in plasma membrane preparations performed in kinetic mode until kinetic equilibrium with a saturating concentration of GLP-1 and TT-OAD2. The BRET signal decreased in the presence of GTP, which suggests that GTP dissociated the Gi2 protein complex stabilized by GLP-1-occupied and TT-OAD2-occupied GLP-1R. Quantification of the plateau (middle) and the rate of ligand-induced conformational change (right) for each agonist (1 μM GLP-1 and 10 μM TT-OAD2) was calculated by applying a one-phase association curve to the kinetic data with values from each individual experiment show in black circles. f, Concentration–response curves of production in live HEK293 cells expressing the GLP-1R and an EPAC BRET biosensor in the presence of different concentrations of GLP-1 and TT-OAD2. Left, cAMP response taken 25 min after ligand addition. Right, area under the curve (AUC) analysis of the response calculated as AUC across the full kinetic trace for each ligand concentration (from data in Fig. 2d). Data are mean + s.e.m. of 4–6 independent experiments performed in duplicate or triplicate.
Extended Data Fig. 2 Purification, cryo-EM data imaging and processing of the TT-OAD2–GLP-1R–Gs complex.
a, Representative elution profile of Flag-purified complex on Superdex 200 Increase 10/30 SEC. b, Representative micrograph of the TT-OAD2–GLP-1R–Gs complex. Red circles highlight examples of individual particles. c, Two-dimensional class averages of the complex in maltose-neopentyl glycol (MNG) micelle. d, Cryo-EM data processing workflow. e, Gold-standard Fourier shell correlation (FSC) curves, showing the overall nominal resolution at 3.0 Å. f, 3D histogram representation of the Euler angle distribution of all the particles used for the in the reconstruction overlaid on the density map drawn on the same coordinate axis (map is coloured according to local resolution as in g). g, Cryo-EM density map coloured according to resolution. Left, map with the GLP-1R ECD masked; right, map including the ECD of GLP-1R.
Extended Data Fig. 3 The atomic resolution model of the TT-OAD2–GLP-1R–Gαs heterotrimer in the cryo-EM density map.
Electron microscopy density map and the model are shown for all seven transmembrane helices and helix 8 (H8) of the receptor, the α5 helix of the GαS Ras-like domain and TT-OAD2. All transmembrane helices exhibit good density, with TM6 that displays flexibility being the least well resolved region.
Extended Data Fig. 4 Cryo-EM density supports ligand interactions in the TT-OAD2-binding site.
a, Interacting residues predicted by LigPlot using the full-length model with ECD. b, The pose of TT-OAD2 and interactions with residues within TM1, TM2, TM3, ECL1 and ECL2 are supported by well-resolved density in the cryo-EM map. c, Density for the ECD was visible in the cryo-EM and supports extended interactions of the ECD with ECL1 and ECL2, as well as with the ligand TT-OAD2.
Extended Data Fig. 5 Comparison of the TT-OAD2–GLP-1R–Gs complex with peptide agonist-bound GLP-1R structures and the inactive class B GPCR glucagon receptor transmembrane helices.
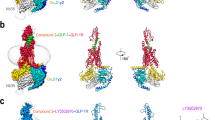
a, Structures of agonist-bound GLP-1R; from left to right: GLP-1R (orange) bound to GLP-1 peptide (green) in an active conformation, GLP-1R (pink) bound to ExP5 peptide (cyan) in an active conformation, GLP-1R (blue) bound to non-peptide TT-OAD2 (red) in an active conformation, GLP-1R (pale green) bound to 11-mer peptide HepP5 (purple) in a partially active conformation. Far right, overlay of GLP-1R agonist-bound structures highlighting variations within the ECD position in the different structures. Inset, differences in the location of the ECD are supported by density in the cryo-EM maps; shown are the GLP-1-bound (orange) and TT-OAD2 bound (blue) GLP-1R. b, c, Various overlays of these structures (using the same colours) to compare conformational differences between the different structures. b, Overlay of TT-OAD2-bound GLP-1R Gs structure with the full-length peptide bound Gs structures and the inactive glucagon receptor (GCGR; grey) bundle reveals common conformational transitions occur in all agonist-bound structures relative to the inactive GCGR, but the extent of these movements differ. A more open helical bundle is observed for the TT-OAD2-bound GLP-1R than either GLP-1- or ExP5-bound owing to a distinction in the conformations of TM1, TM6, TM7 and ECL3 at the extracellular side of the receptor induced by the binding of the different ligands (left and middle). Middle, differences in the conformation of TM2 between the inactive and peptide-agonist-bound structures is also evident. Right, at the intracellular face all active structures display a similar large outward movement of TM6 and a smaller movement within TM5. c, Comparison of TT-OAD2-bound GLP-1R with the small peptide HepP5-bound GLP-1R structure. Left, TT-OAD2 and Hep-P5 occupy a partially overlapping binding site but promote distinct conformations of the ECD and transmembrane bundle of the GLP-1R. Middle, HepP5 engages deeper in the helical bundle than TT-OAD2 and promotes a more closed helical bundle owing to differences induced in the conformation of TM1, TM6, TM7 and ECL3. Right, overlay of the TT-OAD2-, Hep-P5- and GLP-1-bound GLP-1R transmembrane bundles reveals HepP5 induces a similar conformation of the helical bundle to GLP-1 whereas TT-OAD2 induces a distinct conformation.
Extended Data Fig. 6 Pharmacological responses exhibited by endogenous ligands GLP-1 and oxyntomodulin in the presence of TT-OAD2.
Signalling profiles of GLP-1 and oxyntomodulin, after 30 min preincubation of vehicle (0) or increasing concentrations of TT-OAD2. Data were performed in HEK293A cells stably expressing the GLP-1R, and are mean + s.e.m. of 3–4 independent experiments performed in duplicate.
Extended Data Fig. 7 GLP-1R domains are stabilized by either ligand contacts or lipid interactions.
a, Top, RMSF values of alpha carbons computed during MD simulations of the GLP-1R–GLP-1–Gs complex (black line) and the GLP-1R–TT-OAD2–Gs complex (red line); transmembrane helices, intracellular loops (ICLs), and ECLs positions are indicated. Bottom left, RMSF values plotted on the GLP-1R structure bound to GLP-1 (transparent ribbon). Bottom right, RMSF values plotted on the GLP-1R structure bound to TT-OAD2 (transparent stick representation). ECL1 and ECL3 were more dynamic in the GLP-1-bound receptor than the TT-OAD2-bound structure. By contrast, ECL2 and the top end of TM5 were more mobile in the GLP-1R–TT-OAD2–Gs complex. b, GLP-1R contacts formed with membrane lipids during molecular dynamic simulations of the GLP-1R–TT-OAD2–Gs and the GLP-1R–GLP-1–Gs systems. Two sides views of the receptor are shown (ribbon and transparent surface). When bound to TT-OAD2, ECL1, TM3, the distal end of TM6, and ECL3 are more in contact with the membrane lipids (magenta). By contrast, TM1 and TM7 are more prone to interact with the membrane when GLP-1 is bound (green). The outward movement of ECL3 in the GLP-1R–TT-OAD2–Gs complex (stabilized by a hydrogen bond network different than GLP-1R–GLP-1–Gs; Extended Data Table 2) produces more interactions with the lipids, possibly further stabilizing the open conformation of TM6, ECL3 and TM7.
Extended Data Fig. 8 Dynamics of the ECD of GLP-1R.
a, The vector (shown here as a green arrow) connecting S49ECD and E34ECD alpha carbons (ECD N-terminal helix) are shown in the box. b, Left, ECD N-terminal helix orientations observed during the molecular dynamics simulation of the GLP-1R–GLP-1–Gs (black arrows), the GLP-1R–GLP-1 complex (obtained by removing G protein; blue arrows), and the apo-GLP-1R (obtained by removing both the Gs protein and GLP-1; cyan arrows) are shown on the left viewed from the top and side of the bundle. The receptor is shown as a dark grey ribbon. During molecular dynamic simulations with GLP-1 bound, the N-terminal helix was oriented vertically (black and blue arrows), whereas in the apo-form the ECD N-terminal helix was more dynamic and experienced both open and closed conformations (this is analogous to the suggested ECD dynamics for the glucagon receptor). Right, ECD N-terminal helix orientations of the GLP-1R–TT-OAD2–Gs (red arrows), the GLP-1R–TT-OAD2 complex (obtained by removing G protein; orange arrows), and the apo-GLP-1R (obtained by removing both the Gs protein and TT-OAD2; yellow arrows) are shown. The receptor is shown as a red ribbon. The distal end (S49ECD) of the helix was more mobile than the proximal one (E34ECD), which had an overall tendency to remain in the proximity of the TT-OAD2-binding site, driven by transient interactions with the ligand (Extended Data Table 1) and hydrogen bonds with the R299ECL2 side chain (Extended Data Table 2). Molecular dynamics simulations therefore suggest a different behaviour for residue R299ECL2, which is stably involved in interactions with the peptide in the GLP-1-R–GLP-1–Gs complex (Extended Data Table 1), and instead interacts with E34ECD and other residues located at the ECL2 (E294ECD, D293ECD and N300ECD) in the GLP-1R–TT-OAD2–Gs complex (Extended Data Table 2).
Extended Data Fig. 9 Proposed activation mechanism of class B GPCRs.
Left, in the inactive conformation, the top of the transmembrane domain is stabilized by interactions of the ECD with the TM6–ECL3–TM7 region. Top, activation of class B GPCRs by peptides occurs via a two domain mechanism. Top left, engagement of the peptide with the receptor ECD releases ECD constraints on the transmembrane domain promoting outward movements of TM1, TM6 and TM7 by peptide. Middle, interaction of the peptide N terminus in the bundle within TM1, TM2, TM3, TM5, TM6 and TM7 promotes TM1, TM6 and TM7 to close in around the peptide. Direct engagement of peptides with the central polar network facilitates conformational transitions required for G protein coupling and activation. Top right, the active conformation of the central polar network is stabilized by a series of structural waters. Bottom, interaction of the non-peptide TT-OAD2 at the top of the GLP-1R transmembrane bundle releases ECD constraints on the transmembrane bundle resulting in movements of TM1, TM6 and TM7 outwards. TT-OAD2 does not engage TM5–TM7 and the bundle remains open. TT-OAD allosterically promotes conformational rearrangement of the central polar network to stabilize the fully active receptor conformation that allows coupling to G protein. Bottom right, the central polar network is stabilized by a distinct network of structural waters relative to peptide-mediated activation.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-3.
Video 1
Different hydration of polar networks within the GLP-1R transmembrane (TM) domain. In the GLP-1R:TT-OAD2 Gs complex (left), structural water molecules form stable hydrogen bonds with the N3205.50 side chain and the Y2413.44 backbone atoms as well with E3646.53. The bound GLP-1 (right) stabilizes a water molecule network in the proximity of the peptide N-terminal residues H7 and A8 as well of Y1521.47, T3917.46, R1902.60, E3645.53.
Video 2
TT-OAD2 interactions lead to reorganisation and stabilisation of the central polar network via a distinct mechanism to GLP-1. The GLP-1R:TT-OAD2 interactions modify the hydrogen bond network between TM1 and TM2. Left; GLP-1 (brown ribbon representation) residue 3 (D9) (white stick) forms an ionic interaction (red dotted lines) with R1902.60, which is involved in key hydrogen bonds with N2403.43 (in turn interacting with S1862.56). At the top of TM2, K1972.67, D1982.68, and Y1451.40 are stabilized in polar interactions (red dotted lines). Right; The TT-OAD2 (brown stick and transparent van der Waals surface) binding triggers the reorganization of the interaction network at the top of TM1 and leads the Y1481.43 and Y1521.47 to make contact and form hydrogen bond interactions with the R1902.60. The ECL3 of the GLP-1R:TT-OAD complex was removed for clarity.
Rights and permissions
About this article
Cite this article
Zhao, P., Liang, YL., Belousoff, M.J. et al. Activation of the GLP-1 receptor by a non-peptidic agonist. Nature 577, 432–436 (2020). https://doi.org/10.1038/s41586-019-1902-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1902-z
This article is cited by
-
G protein-coupled receptors (GPCRs): advances in structures, mechanisms, and drug discovery
Signal Transduction and Targeted Therapy (2024)
-
Cryo-electron microscopy for GPCR research and drug discovery in endocrinology and metabolism
Nature Reviews Endocrinology (2024)
-
GLP-1R signaling neighborhoods associate with the susceptibility to adverse drug reactions of incretin mimetics
Nature Communications (2023)
-
Conserved class B GPCR activation by a biased intracellular agonist
Nature (2023)
-
GWAS of random glucose in 476,326 individuals provide insights into diabetes pathophysiology, complications and treatment stratification
Nature Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.