Abstract
Although maternal antibodies protect newborn babies from infection1,2, little is known about how protective antibodies are induced without prior pathogen exposure. Here we show that neonatal mice that lack the capacity to produce IgG are protected from infection with the enteric pathogen enterotoxigenic Escherichia coli by maternal natural IgG antibodies against the maternal microbiota when antibodies are delivered either across the placenta or through breast milk. By challenging pups that were fostered by either maternal antibody-sufficient or antibody-deficient dams, we found that IgG derived from breast milk was crucial for protection against mucosal disease induced by enterotoxigenic E. coli. IgG also provides protection against systemic infection by E. coli. Pups used the neonatal Fc receptor to transfer IgG from milk into serum. The maternal commensal microbiota can induce antibodies that recognize antigens expressed by enterotoxigenic E. coli and other Enterobacteriaceae species. Induction of maternal antibodies against a commensal Pantoea species confers protection against enterotoxigenic E. coli in pups. This role of the microbiota in eliciting protective antibodies to a specific neonatal pathogen represents an important host defence mechanism against infection in neonates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
16S rRNA gene profiling data and the ETEC 6 genome are available in the NCBI database under BioProject PRJNA577743 and BioSample SAMN12263012, respectively.
References
Basha, S., Surendran, N. & Pichichero, M. Immune responses in neonates. Expert Rev. Clin. Immunol. 10, 1171–1184 (2014).
Simon, A. K., Hollander, G. A. & McMichael, A. Evolution of the immune system in humans from infancy to old age. Proc. R. Soc. B 282, 20143085 (2015).
Kamada, N., Chen, G. Y., Inohara, N. & Núñez, G. Control of pathogens and pathobionts by the gut microbiota. Nat. Immunol. 14, 685–690 (2013).
Carbonare, C. B., Carbonare, S. B. & Carneiro-Sampaio, M. M. S. Secretory immunoglobulin A obtained from pooled human colostrum and milk for oral passive immunization. Pediatr. Allergy Immunol. 16, 574–581 (2005).
Hanson, L. A. R. & Korotkova, M. The role of breastfeeding in prevention of neonatal infection. Semin. Neonatol. 7, 275–281 (2002).
Madoff, L. C., Michel, J. L., Gong, E. W., Rodewald, A. K. & Kasper, D. L. Protection of neonatal mice from group B streptococcal infection by maternal immunization with beta C protein. Infect. Immun. 60, 4989–4994 (1992).
Zaman, K. et al. Effectiveness of maternal influenza immunization in mothers and infants. N. Engl. J. Med. 359, 1555–1564 (2008).
Englund, J. A. et al. Transplacental antibody transfer following maternal immunization with polysaccharide and conjugate Haemophilus influenzae type b vaccines. J. Infect. Dis. 171, 99–105 (1995).
Kearney, J. F., Patel, P., Stefanov, E. K. & King, R. G. Natural antibody repertoires: development and functional role in inhibiting allergic airway disease. Annu. Rev. Immunol. 33, 475–504 (2015).
Macpherson, A. J., de Agüero, M. G. & Ganal-Vonarburg, S. C. How nutrition and the maternal microbiota shape the neonatal immune system. Nat. Rev. Immunol. 17, 508–517 (2017).
Chen, Y. et al. Microbial symbionts regulate the primary Ig repertoire. J. Exp. Med. 215, 1397–1415 (2018).
Englund, J. A. et al. Maternal immunization with influenza or tetanus toxoid vaccine for passive antibody protection in young infants. J. Infect. Dis. 168, 647–656 (1993).
Boes, M., Prodeus, A. P., Schmidt, T., Carroll, M. C. & Chen, J. A critical role of natural immunoglobulin M in immediate defense against systemic bacterial infection. J. Exp. Med. 188, 2381–2386 (1998).
Ochsenbein, A. F. et al. Control of early viral and bacterial distribution and disease by natural antibodies. Science 286, 2156–2159 (1999).
Baumgarth, N. et al. B-1 and B-2 cell-derived immunoglobulin M antibodies are nonredundant components of the protective response to influenza virus infection. J. Exp. Med. 192, 271–280 (2000).
Jayasekera, J. P., Moseman, E. A. & Carroll, M. C. Natural antibody and complement mediate neutralization of influenza virus in the absence of prior immunity. J. Virol. 81, 3487–3494 (2007).
Zhou, Z. H. et al. The broad antibacterial activity of the natural antibody repertoire is due to polyreactive antibodies. Cell Host Microbe 1, 51–61 (2007).
Caballero-Flores, G. et al. Maternal immunization confers protection to the offspring against an attaching and effacing pathogen through delivery of IgG in breast milk. Cell Host Microbe 25, 313–323 (2019).
Palmeira, P., Quinello, C., Silveira-Lessa, A. L., Zago, C. A. & Carneiro-Sampaio, M. IgG placental transfer in healthy and pathological pregnancies. Clin. Dev. Immunol. 2012, 985646 (2012).
Masuda, A. et al. Fcγ receptor regulation of Citrobacter rodentium infection. Infect. Immun. 76, 1728–1737 (2008).
Pyzik, M., Rath, T., Lencer, W. I., Baker, K. & Blumberg, R. S. FcRn: the architect behind the immune and nonimmune functions of IgG and albumin. J. Immunol. 194, 4595–4603 (2015).
Israel, E. J. et al. Expression of the neonatal Fc receptor, FcRn, on human intestinal epithelial cells. Immunology 92, 69–74 (1997).
Kotloff, K. L. et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): a prospective, case–control study. Lancet 382, 209–222 (2013).
Kotloff, K. L. et al. Global burden of diarrheal diseases among children in developing countries: incidence, etiology, and insights from new molecular diagnostic techniques. Vaccine 35, 6783–6789 (2017).
Kotloff, K. L. et al. The incidence, aetiology, and adverse clinical consequences of less severe diarrhoeal episodes among infants and children residing in low-income and middle-income countries: a 12-month case–control study as a follow-on to the Global Enteric Multicenter Study (GEMS). Lancet Glob. Health 7, e568–e584 (2019).
Qadri, F., Svennerholm, A.-M., Faruque, A. S. G. & Sack, R. B. Enterotoxigenic Escherichia coli in developing countries: epidemiology, microbiology, clinical features, treatment, and prevention. Clin. Microbiol. Rev. 18, 465–483 (2005).
Thapar, N. & Sanderson, I. R. Diarrhoea in children: an interface between developing and developed countries. Lancet 363, 641–653 (2004).
Skurnik, D., Cywes-Bentley, C. & Pier, G. B. The exceptionally broad-based potential of active and passive vaccination targeting the conserved microbial surface polysaccharide PNAG. Expert Rev. Vaccines 15, 1041–1053 (2016).
Le Gallou, S. et al. A splenic IgM memory subset with antibacterial specificities is sustained from persistent mucosal responses. J. Exp. Med. 215, 2035–2053 (2018).
Wilmore, J. R. et al. Commensal microbes induce serum IgA responses that protect against polymicrobial sepsis. Cell Host Microbe 23, 302–311 (2018).
Apter, F. M. et al. Analysis of the roles of antilipopolysaccharide and anti-cholera toxin immunoglobulin A (IgA) antibodies in protection against Vibrio cholerae and cholera toxin by use of monoclonal IgA antibodies in vivo. Infect. Immun. 61, 5279–5285 (1993).
Michetrti, P., Mahan, M. J., Slauch, J. M., Mekalanos, J. J. & Neutra, M. R. Monoclonal secretory immunoglobulin A protects mice against oral challenge with the invasive pathogen Salmonella typhimurium. Infect. Immun. 60, 1786–1792 (1992).
Moor, K. et al. High-avidity IgA protects the intestine by enchaining growing bacteria. Nature 544, 498–502 (2017).
Stuebe, A. The risks of not breastfeeding for mothers and infants. Rev. Obstet. Gynecol. 2, 222–231 (2009).
Goldsmith, S. J., Dickson, J. S., Barnhart, H. M., Toledo, R. T. & Eiten-Miller, R. R. IgA, IgG, IgM and lactoferrin contents of human milk during early lactation and the effect of processing and storage. J. Food Prot. 46, 4–7 (1983).
Fouda, G. G. et al. HIV-specific functional antibody responses in breast milk mirror those in plasma and are primarily mediated by IgG antibodies. J. Virol. 85, 9555–9567 (2011).
Dickinson, B. L. et al. Bidirectional FcRn-dependent IgG transport in a polarized human intestinal epithelial cell line. J. Clin. Invest. 104, 903–911 (1999).
Bournazos, S. & Ravetch, J. V. Diversification of IgG effector functions. Int. Immunol. 29, 303–310 (2017).
Mostov, K. E. Transepithelial transport of immunoglobulins. Annu. Rev. Immunol. 12, 63–84 (1994).
Yoshida, M. et al. Human neonatal Fc receptor mediates transport of IgG into luminal secretions for delivery of antigens to mucosal dendritic cells. Immunity 20, 769–783 (2004).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl Acad. Sci. USA 108, 4516–4522 (2011).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Suzuki, M. T. & Giovannoni, S. J. Bias caused by template annealing in the amplification of mixtures of 16S rRNA genes by PCR. Appl. Environ. Microbiol. 62, 625–630 (1996).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank all of the staff in our animal facility for their support in animal husbandry; all Kasper and Mekalanos laboratory members for comments; Y. Chen for technical help; and F. Qadri for providing the ETEC 6 strain. The study was supported by NIH NIAID Center of Excellence for Translational Research Grant 1U19 AI109764 awarded to D.L.K. and AI01845 awarded to J.J.M. G.S. was supported in part by funding from the European Union’s Horizon 2020 Research and Innovation Programme under Marie Skłodowska Curie Grant Agreement 661138.
Author information
Authors and Affiliations
Contributions
W. Zheng, W. Zhao, J.J.M. and D.L.K. conceived and designed the study and wrote and revised the manuscript. W. Zheng performed all mouse breeding, genotyping and fostering experiments, ELISA and biochemistry experiments. W. Zhao and W. Zheng conducted mouse challenge experiments and immuno-analysis of antibody specificities. M.W. performed the 16S rRNA gene-sequencing study of the mouse gut microbiota. F.C. conducted the mouse transcriptome analysis. X. Song helped to maintain the mouse line with genotyping and provided insights on mouse breeding, western blot and other techniques. X. Sun, F.G., G.S. and S.O. provided discussions during the generation of the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Duane Wesemann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Persistence and development of maternal antibodies.
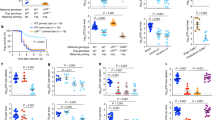
The genotypes of mating pairs are indicated under the large mouse cartoons; the small mouse cartoons represent neonates born to the indicated dams and their genotypes. Red symbols denote the presence in neonates of antibodies that either were acquired transplacentally from μMT+/− mothers or, in the case of μMT+/− pups, were generated endogenously after 4 weeks of age. a, Reciprocal breeding scheme to study maternal antibody persistence and development. b, Serum IgG concentration in 1–8-week-old pups. Data are shown as μg ml−1. n = 5–15 mice in each breeding group for every week of 1–8 weeks. Further details are provided in the Source Data. c, IgA concentrations in small-intestine (SI) and colon (CO) homogenates from 1-week-old pups. Data are shown as μg per small intestine or colon. d, Faecal IgA concentration in 2–8-week-old pups. Data are shown as μg per g of faeces. n = 6–13 mice in each breeding group for every week of 2–8 weeks. Further details are provided in the Source Data. e, Serum IgM concentration in 1-week-old pups. Data are shown as μg ml−1. f, IgG concentration in small-intestine and colon homogenates from 1-week-old pups. Data are shown as μg per small intestine or colon. g, Faecal IgG concentration in 2–8-week-old pups. Data are shown as μg per g of faeces. n = 5–9 mice in each breeding group for every week of 2–8 weeks. Further details are provided in the Source Data. b–g, Data are mean ± s.e.m. Specific n numbers are shown in the figure.
Extended Data Fig. 2 Comparison of mNAb+ and mNab− pups.
a, Survival of 2-week-old mNAb+ and mNab− pups on day 1 after intraperitoneal challenge with 107 CFU of ETEC 6. mNab+ group, n = 6 pups; mNab− group, n = 15 pups. b, 16S rRNA gene analysis of the composition of the microbiota in 1-week-old reciprocally bred pups. Data are the average of 8 or 9 individual pups from each group; mNab+ group, n = 9 pups; mNab− group, n = 8 pups. c, Transcriptome analysis of small intestines of ETEC-6-infected mNab+ and mNab− pups using RNA sequencing. n = 8 mNab+ pups; n = 4 mNab− pups. Specific n numbers are shown in the figure.
Extended Data Fig. 3 Comparison of pups from the cross-fostering experiment.
For all panels, cross-fostering of neonates is denoted by horizontal arrows that provide the genotype of the pup followed by the genotype of the fostering dam. a, Cross-fostering experimental scheme. The genotypes of dams are indicated under the large mouse cartoons; the small mouse cartoons represent neonates that are born to those dams and have the same genotype as their mothers. Thicker arrows define the mother that fostered the indicated neonates. Red symbols denote the presence in neonates of antibodies that were acquired transplacentally from their μMT+/− mothers or, in the case of μMT−/− pups, from a μMT+/− fostering dam. b, Colon IgG concentration of 1-week-old cross-fostered pups. Data are shown as μg per colon. c, Colon IgA concentration of 1-week-old cross-fostered pups. Data are shown as μg per colon. d, Fostering scheme of μMT−/− pups cross-fostered by a μMT+/− dam. e, Survival of fostered μMT−/− pups at 20-h after ETEC infection. In the first experiment, n = 5 μMT−/− pups were fostered by μMT+/− dams; n = 5 μMT−/− pups were fostered by μMT−/− dams. In the second experiment, n = 6 μMT−/− pups were cross-fostered by μMT+/− dams; n = 8 μMT−/− pups were fostered by μMT−/− dams. b, c, Data are mean ± s.e.m. Specific n numbers are shown in the figure.
Extended Data Fig. 4 Relative concentrations of all subclasses of IgG between dams and pups.
a–e, Serum IgG subclass concentrations in μMT−/−FcRn+/+ (or FcRn+/−) and μMT−/−FcRn−/− pups fostered by μMT+/− dams for 1 week. f–j, Relative concentrations of all subclasses of IgG between dam and pups. k, Adult (8-week-old) mice were orally gavaged with 5 mg of IgG, and serum IgG concentrations were quantified as ng ml−1. NS, no significant difference; calculated using a Mann–Whitney U-test. Specific n numbers are shown in the figure.
Extended Data Fig. 5 Serum from conventionally colonized (SPF) mice broadly recognizes human commensal bacteria and other enteric pathogens.
a, Total immunoglobulin titres against E. coli strain Nissle 1917 in germ-free and SPF mouse serum. b, Total immunoglobulin titres against Salmonella typhimurium in germ-free and SPF mouse serum. Specific n numbers are shown in the figure.
Extended Data Fig. 6 Characteristics of commensal-immunized and unimmunized serum.
a, Western blot analysis of serum IgG from pups born to Pantoea-1-immunized SPF mice shows epitopes of ETEC 6, Pantoea 1 and Enterobacter, similar in size to those in serum from pups born to Pantoea-1-immunized germ-free mice. b, Serum IgG of pups born to Pantoea-immunized germ-free dams cross-reacts with different Enterobacteriaceae isolates from different facilities. 1, Harvard SGM Pantoea; 2, ETEC 6; 3, Harvard SGM Enterobacter; 4, Bacteroides fragilis NCTC9343; 5, Charles River B6 Proteus mirabilis; 6, Charles River B6 E. coli isolate 1; 7, Charles River B6 E. coli isolate 2; 8, Charles River CD1 E. coli isolate 1; 9, Charles River CD1 E. coli isolate 2; 10, Taconic B6 E. coli isolate 1; 11, Taconic B6 E. coli isolate 2; 12, Taconic B6 Proteus isolate; 13, Taconic B6 Enterobacter isolate; 14, C. rodentium. c, Pronase-treated bacterial lysates blotted with serum IgG of pups born to Pantoea-immunized germ-free dams. The concentrations of pronase are specified in the figure. a–c, Proteins were detected using a goat anti-mouse IgG antibody. For gel source data, see Supplementary Fig. 1. d, Mouse milk IgG and IgA concentrations. Data are shown in μg ml−1. e, Mouse milk IgG titre against microbiota. Each line represents an independent mouse. d, Data are mean ± s.e.m.
Extended Data Fig. 7 Schematic summary of the findings in this study.
Top, mNabs induced by commensal microbiota in dams were transferred to neonates through the breast milk. Cross-reacting mNabs (especially IgG antibodies) were detected that bound to the pathogenic, non-indigenous bacterial species ETEC and correlated with protection against disease in pups challenged with ETEC. IgG antibodies were also shown to be transported from the milk to the bloodstream of pups by a process that we call IgG retro-transport. Bottom, mNabs react with many commensal species and among them an Enterobacteriaceae isolate (Pantoea) was found to induce antibodies that cross-react with ETEC. The immunogenicity of this commensal species is hypothesized to be a result of local antigen-sampling processes that involve dendritic cells and uptake by Peyer’s patch germinal centres. This ultimately leads to the induction of high-affinity IgGs directed against a Pantoea antigen that cross-reacts with ETEC. IgG was also shown to be transported from the blood stream to the intestinal lumen by FcRn in adult mice. Illustrations were created with BioRender (https://biorender.com/).
Supplementary information
Supplementary Figure 1 | Uncropped blots
Figure 4e: Bacterial lysates blot with sera of pups born to Pantoea immunized GF dam or non immunized GF dam. Bacterial lysates ran twice on one NC membrane, cut in half, left blot with sera of pups born to Pantoea immunized GF dam, right blot with sera of pups born to non immunized GF serum, same serum dilution, two blots are scanned at the same time under the same exposure. Extended Data figure 6a left and center gels: Bacterial lysates blot with sera of pups born to Pantoea immunized SPF dam and non-immunized SPF dam. Extended Data figure 6a right gel: Bacterial lysates blot with sera of GF pups born to GF non immunized dam (same dilution as SPF immunized and SPF non-immunized serum). Extended Data figure 6b: Gram negative Enterobacteriaceae isolates from different sources blot with sera of pups born to Pantoea immunized GF dam. Extended Data figure 6c: Bacterial lysates treated with different concentrations of pronase with sera of pups born to Pantoea immunized GF dam.
Rights and permissions
About this article
Cite this article
Zheng, W., Zhao, W., Wu, M. et al. Microbiota-targeted maternal antibodies protect neonates from enteric infection. Nature 577, 543–548 (2020). https://doi.org/10.1038/s41586-019-1898-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1898-4
This article is cited by
-
The biological functions and metabolic pathways of valine in swine
Journal of Animal Science and Biotechnology (2023)
-
Microbiota-mediated colonization resistance: mechanisms and regulation
Nature Reviews Microbiology (2023)
-
Early-life interactions between the microbiota and immune system: impact on immune system development and atopic disease
Nature Reviews Immunology (2023)
-
Gut–liver axis: barriers and functional circuits
Nature Reviews Gastroenterology & Hepatology (2023)
-
Mouse milk immunoglobulin G Fc-linked N-glycosylation nano-LC–MS analysis in a model of vancomycin exposure during pregnancy
Analytical and Bioanalytical Chemistry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.