Abstract
The most frequently mutated oncogene in cancer is KRAS, which uses alternative fourth exons to generate two gene products (KRAS4A and KRAS4B) that differ only in their C-terminal membrane-targeting region1. Because oncogenic mutations occur in exons 2 or 3, two constitutively active KRAS proteins—each capable of transforming cells—are encoded when KRAS is activated by mutation2. No functional distinctions among the splice variants have so far been established. Oncogenic KRAS alters the metabolism of tumour cells3 in several ways, including increased glucose uptake and glycolysis even in the presence of abundant oxygen4 (the Warburg effect). Whereas these metabolic effects of oncogenic KRAS have been explained by transcriptional upregulation of glucose transporters and glycolytic enzymes3,4,5, it is not known whether there is direct regulation of metabolic enzymes. Here we report a direct, GTP-dependent interaction between KRAS4A and hexokinase 1 (HK1) that alters the activity of the kinase, and thereby establish that HK1 is an effector of KRAS4A. This interaction is unique to KRAS4A because the palmitoylation–depalmitoylation cycle of this RAS isoform enables colocalization with HK1 on the outer mitochondrial membrane. The expression of KRAS4A in cancer may drive unique metabolic vulnerabilities that can be exploited therapeutically.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information.
Code availability
The code used for super-resolution imaging is available at GitHub (https://github.com/yiny02/direct-Triple-Correlation-Algorithm) and in the supplementary information of a previous study29.
References
Tsai, F. D. et al. K-Ras4A splice variant is widely expressed in cancer and uses a hybrid membrane-targeting motif. Proc. Natl Acad. Sci. USA 112, 779–784 (2015).
Voice, J. K., Klemke, R. L., Le, A. & Jackson, J. H. Four human Ras homologs differ in their abilities to activate Raf-1, induce transformation, and stimulate cell motility. J. Biol. Chem. 274, 17164–17170 (1999).
Kimmelman, A. C. Metabolic dependencies in RAS-driven cancers. Clin. Cancer Res. 21, 1828–1834 (2015).
Ying, H. et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149, 656–670 (2012).
Kerr, E. M., Gaude, E., Turrell, F. K., Frezza, C. & Martins, C. P. Mutant Kras copy number defines metabolic reprogramming and therapeutic susceptibilities. Nature 531, 110–113 (2016).
Gottlob, K. et al. Inhibition of early apoptotic events by Akt/PKB is dependent on the first committed step of glycolysis and mitochondrial hexokinase. Genes Dev. 15, 1406–1418 (2001).
Vyssokikh, M. Y. & Brdiczka, D. The function of complexes between the outer mitochondrial membrane pore (VDAC) and the adenine nucleotide translocase in regulation of energy metabolism and apoptosis. Acta Biochim. Pol. 50, 389–404 (2003).
Patra, K. C. et al. Hexokinase 2 is required for tumor initiation and maintenance and its systemic deletion is therapeutic in mouse models of cancer. Cancer Cell 24, 213–228 (2013).
Ahearn, I., Zhou, M. & Philips, M. R. Posttranslational modifications of RAS proteins. Cold Spring Harb. Perspect. Med. 8, a031484 (2018).
Rocks, O. et al. The palmitoylation machinery is a spatially organizing system for peripheral membrane proteins. Cell 141, 458–471 (2010).
Jennings, B. C. et al. 2-Bromopalmitate and 2-(2-hydroxy-5-nitro-benzylidene)-benzo[b]thiophen-3-one inhibit DHHC-mediated palmitoylation in vitro. J. Lipid Res. 50, 233–242 (2009).
John, S., Weiss, J. N. & Ribalet, B. Subcellular localization of hexokinases I and II directs the metabolic fate of glucose. PLoS One 6, e17674 (2011).
Wohlgemuth, S. et al. Recognizing and defining true Ras binding domains I: biochemical analysis. J. Mol. Biol. 348, 741–758 (2005).
Zhu, A., Romero, R. & Petty, H. R. An enzymatic colorimetric assay for glucose-6-phosphate. Anal. Biochem. 419, 266–270 (2011).
Tsai, H. J. & Wilson, J. E. Functional organization of mammalian hexokinases: both N- and C-terminal halves of the rat type II isozyme possess catalytic sites. Arch. Biochem. Biophys. 329, 17–23 (1996).
Sebastian, S., Wilson, J. E., Mulichak, A. & Garavito, R. M. Allosteric regulation of type I hexokinase: a site-directed mutational study indicating location of the functional glucose 6-phosphate binding site in the N-terminal half of the enzyme. Arch. Biochem. Biophys. 362, 203–210 (1999).
Rajalingam, K., Schreck, R., Rapp, U. R. & Albert, S. Ras oncogenes and their downstream targets. Biochim. Biophys. Acta 1773, 1177–1195 (2007).
Chiu, V. K. et al. Ras signalling on the endoplasmic reticulum and the Golgi. Nat. Cell Biol. 4, 343–350 (2002).
Mor, A. & Philips, M. R. Compartmentalized Ras/MAPK signaling. Annu. Rev. Immunol. 24, 771–800 (2006).
Aran, V. & Prior, I. A. Compartmentalized Ras signaling differentially contributes to phenotypic outputs. Cell. Signal. 25, 1748–1753 (2013).
Yun, J. et al. Glucose deprivation contributes to the development of KRAS pathway mutations in tumor cells. Science 325, 1555–1559 (2009).
Newlaczyl, A. U., Coulson, J. M. & Prior, I. A. Quantification of spatiotemporal patterns of Ras isoform expression during development. Sci. Rep. 7, 41297 (2017).
Tanner, L. B. et al. Four key steps control glycolytic flux in mammalian cells. Cell Syst. 7, 49–62 (2018).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Hay, N. Reprogramming glucose metabolism in cancer: can it be exploited for cancer therapy? Nat. Rev. Cancer 16, 635–649 (2016).
Peled, M. et al. Affinity purification mass spectrometry analysis of PD-1 uncovers SAP as a new checkpoint inhibitor. Proc. Natl Acad. Sci. USA 115, E468–E477 (2018).
Zhou, M. & Philips, M. R. Nitrogen cavitation and differential centrifugation allows for monitoring the distribution of peripheral membrane proteins in cultured cells. J. Vis. Exp. 126, e56037 (2017).
Yin, Y., Lee, W. T. C. & Rothenberg, E. Ultrafast data mining of molecular assemblies in multiplexed high-density super-resolution images. Nat. Commun. 10, 119 (2019).
Huang, F. et al. Video-rate nanoscopy using sCMOS camera-specific single-molecule localization algorithms. Nat. Methods 10, 653–658 (2013).
Metallo, C. M. et al. Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. Nature 481, 380–384 (2012).
Grassian, A. R. et al. IDH1 mutations alter citric acid cycle metabolism and increase dependence on oxidative mitochondrial metabolism. Cancer Res. 74, 3317–3331 (2014).
Lewis, C. A. et al. Tracing compartmentalized NADPH metabolism in the cytosol and mitochondria of mammalian cells. Mol. Cell 55, 253–263 (2014).
Fernandez, C. A., Des Rosiers, C., Previs, S. F., David, F. & Brunengraber, H. Correction of 13C mass isotopomer distributions for natural stable isotope abundance. J. Mass Spectrom. 31, 255–262 (1996).
Acknowledgements
We thank D. Esposito and W. Gillette for supplying highly purified recombinant KRAS proteins. This work was funded by the National Institutes of Health (R01CA163489 and R01CA116034 to M.R.P.; R01CA157490, R01CA188048, P01CA117969, R35CA232124 and R01GM095567 to A.C.K.; R35CA210018 and U01CA217864 to A.B.; R01CA214948 to R.P.; T32CA009161 to J.P.M.; and T32GM088118 to C.R.A.); the AACR Basic Cancer Research Fellowship (grant number 15-40-01-MAHA to J.P.M.); the Charles H. Revson Senior Fellowship in Biomedical Science (to J.P.M.), American Cancer Society-New York Cancer Research Fund Postdoctoral Fellowship (grant number PF-18-215-01-TBG to S.J.P.), UCSF Pancreas Center and the Schwartz Family Foundation (to W.-C.C.); and the Lustgarten Foundation and SU2C (to A.C.K.). Proteomic analysis and in vivo imaging were performed in the Proteomics and DART Preclinical Imaging Cores, respectively, each partially funded by the NYU Laura and Isaac Perlmutter Cancer Center Support Grant, NIH/NCI P30CA016087. The Center for Advanced Imaging Innovation and Research (CAI2R, www.cai2r.net) at New York University School of Medicine is supported by NIH/NIBIB P41 EB017183.
Author information
Authors and Affiliations
Contributions
C.R.A., J.P.M. and M.R.P. designed and interpreted all experiments and wrote the manuscript. Unless otherwise stipulated, J.P.M. and C.R.A. performed all experiments. W.-C.C. and A.B. provided advice and A549 and SUIT2 cells that were engineered by CRISPR–Cas9 to lack KRAS4A. M.Z. performed mitochondrial purifications. I.M.A. and H.C. performed the 2-DG growth inhibition studies. J.S. performed hexokinase activity assays. S.L.M. performed the PET CT studies. S.J.P. performed the Seahorse analysis and 13C-glucose labelling. M.J.M. performed the super-resolution microscopy. E.G., A.C.K., Y.Z.W., R.P., S.R.H., and E.R. assisted with the interpretation of the results and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.C.K. has financial interests in Vescor Therapeutics. A.C.K. is an inventor on patents that pertain to KRAS-regulated metabolic pathways, redox control pathways in pancreatic cancer, targeting GOT1 as therapeutic approach and the autophagic control of iron metabolism. A.C.K. is on the Scientific Advisory Board of Rafael Pharmaceuticals and has been a consultant for Diciphera Pharmaceuticals.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Dimitrios Anastasiou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 KRAS4A binds to HK2 in an isoform-specific and GTP-dependent manner.
The indicated Flag-tagged RAS constructs (with or without G12V mutations) were expressed in HeLa cells and immunoprecipitated with anti-Flag beads. Blots were probed for Flag-tagged proteins and endogenous HK2. Flag–RAC1(Q61L) served as the negative control. The immunoblot shown is representative of four independent experiments.
Extended Data Fig. 2 Association of KRAS4A with HK1 and HK2 requires prenylation but is diminished by palmitoylation.
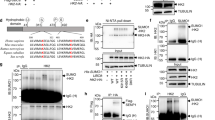
a, HeLa cells expressing the indicated, Flag-tagged GTPase were lysed, KRAS4A or RAC1 were immunoprecipitated and the precipitates were blotted for Flag-tagged GTPases or endogenous HK1 or HK2. b, To confirm that the results in a reflect membrane targeting rather than GTP loading, the relative GTP loading of the KRAS4A proteins was determined by GST–RAF1-RBD affinity capture. Left, Flag–KRAS4A with an activating G12V mutation and either a native (N) membrane-targeting sequence or one with the indicated substitution were expressed in HEK293 cells. Right, in addition, cells expressing Flag–KRAS4A(G12V) were treated with 2-BP to inhibit palmitoylation or FTI to inhibit farnesylation. The total level of Flag–KRAS(G12V) was measured by anti-Flag immunoblot of 1% of the lysate (bottom) and GTP-bound Flag–KRAS4A was measured by affinity purification of the remaining lysate with GST–RAF1-RBD (top). The number under each lane is the amount of GTP-bound KRAS4A relative to lane 1 after normalization for expression (bottom). The immunoblots shown are representative of two independent experiments (a, b).
Extended Data Fig. 3 Colocalization of palmitoylation-deficient KRAS4A but not NRAS with HK1 on mitochondria.
a, Representative live-cell images of COS1 cells co-transfected with the indicated mCherry–KRAS4A constructs and GFP extended with the mitochondrial targeting sequence of HK1 (Mito-GFP). The arrowhead and the arrow indicate the plasma membrane and nuclear envelope, respectively. The cell shown is representative of hundreds on each plate of five independent transfections. Scale bar, 10 µm. b, Colocalization of KRAS4A (with or without palmitoylation) with HK1 and HK2. COS1 or U2OS cells were co-transfected with GFP-tagged HK1 or HK2 and mCherry-tagged, constitutively active KRAS4A(G12V), with or without mutation of cysteine 180 to serine (C180S) to block palmitoylation. The cells were imaged alive using a Zeiss 800 laser scanning confocal microscope and the Pearson’s correlation coefficient between the red and green channels was measured. Data are mean ± s.d. of the values measured in n = 15 cells examined. Significance was determined by unpaired, two-tailed Student’s t-test. c, Neither wild-type nor palmitoylation-deficient NRAS colocalizes with Mito-GFP on mitochondria. mCherry tagged wild-type, palmitoylation-deficient (C181S) or prenylation-deficient (C186S) NRAS were co-expressed in COS1 cells with Mito-GFP and imaged alive using an inverted Zeiss 800 laser scanning confocal microscope. Scale bar, 10 µm. The images shown are representative of hundreds of transfected cells on each plate in two independent experiments. d, Super-resolution (STORM) image of U2OS cells transfected with Flag–KRAS4A(G12V/C180S) or Flag–KRAS4B(G12V), showing colocalization with HK1 on the OMM (arrow) of KRAS4A(G12V/C180S) but not KRAS4B(G12V) (n = 3). Mito, mitochondria. e, Mitochondria were purified from HCT-15 cells that were pre-treated with vehicle or 2-BP and analysed by immunoblot with the indicated antibodies: succinate dehydrogenase (SDHA; mitochondrial matrix), F1-ATPase (inner mitochondrial membrane), RHOGDI (cytosol), fibrillarin (nucleolus) or EEA1 (endosomes). RAS indicates the total level of RAS detected by a pan-RAS antibody. The KRAS4A immunoblot was quantified by a Li-Cor Odyssey infrared scanner. The immunoblot shown is representative of two independent experiments. f, The interaction of KRAS4A with HK1 requires the HK1 OMM-targeting sequence. The indicated Flag-tagged KRAS4A constructs were co-expressed in HEK293 cells with HA-tagged full-length HK1 (FL) or HK1 missing its OMM-targeting region (Δ1–21; ΔMito). Flag–KRAS4A was immunoprecipitated, and binding to HK1 was assessed with an anti-HA immunoblot. The immunoblot shown is representative of four independent experiments.
Extended Data Fig. 4 A putative RAS-binding region in HK1.
a, Crystal structure (PDB 4G0N) of the CRAF RBD (cyan and magenta) in complex with the G domain of HRAS (orange; nucleotide (grey) and magnesium (black)). The regions of the CRAF RBD that mediate the interaction with HRAS are magenta. b, Superposition of the N-terminal lobe of HK1 (green and magenta; PDB 4F9O) alongside HRAS (orange). The putative region of HK1 that interacts with RAS, corresponding to that of the CRAF RBD, is coloured magenta. c, A section of the helix–loop–sheet structure that is common to RBDs is superimposed on the region of HK1 highlighted in b. The RBD of CRAF is shown in cyan, BRAF (PDB 3NY5) in yellow and the putative RBD of HK1 in green. d, Sequence alignments of validated RBDs and the HK1 putative RBD shown with structural motifs of BRAF (yellow) and HK1 (green). Despite the highly conserved structural features shown in c, RBDs have little sequence homology.
Extended Data Fig. 5 Enzyme kinetics of hexokinases with and without recombinant KRAS4A and with and without 2-DG.
a, The activity of recombinant full-length HK1 is unaffected by recombinant KRAS4A. Reaction velocity is plotted (mean ± s.e.m.) as a function of glucose concentration. Velocities are plotted with or without the addition of recombinant, GTP-loaded KRAS4A or RAC2. Plots combine independent assays (n = 4). b, Enzyme kinetics of hexokinases. Full-length HK1 and HK2 and the catalytic C-terminal domain of HK1 were expressed in E. coli as GST fusion proteins and affinity purified with glutathione–agarose beads. Hexokinase activity on the beads was measured with a linked assay kit (BioVision) in which the glucose-6-phosphate produced is oxidized by glucose-6-phosphate dehydrogenase to form NADH, which reduces a colourless probe to a coloured product with strong absorbance at 450 nm. Vmax and Km were calculated by nonlinear regression using GraphPad Prism v 8.1.1 and the goodness of fit is given as R2.
Extended Data Fig. 6 Dissociation between MAPK signalling and differential stimulation of glucose consumption and basal ECAR by KRAS4A versus palmitoylation-deficient KRAS4A and KRAS4B.
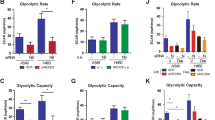
a–c, Flp-In T-REx 293 cells were generated that express the indicated KRAS proteins after induction with doxycycline. a, b, Glucose consumption (mean ± s.e.m.; n = 5) (a) and basal ECAR (mean ± s.e.m.; n = 10) (b) were measured in doxycycline-induced cells. This revealed the order of potency to be KRAS4A(G12V/C180S) > KRAS4A(G12V) > KRAS4B(G12V). Significance was determined by Student’s t-test (paired in a; unpaired in b). c, Immunoblot reveals equivalent expression of the three KRAS proteins in the cells used in a and b. Whereas KRAS4A(G12V) and KRAS4B(G12V) induced equivalent levels of phosphorylated (p)ERK and phosphorylated MEK, KRAS4A(G12V/C180S) (which is palmitoylation deficient) was less potent. These cells have constitutively high levels of AKT phosphorylation that were not altered by expression of any form of KRAS. Note also that despite MAPK stimulation, protein levels of HK1 and HK2 were not altered. The immunoblots shown are representative of two independent experiments. t, total. d, Parental HEK293 cells with lower basal levels of phosphorylated AKT were transfected with the indicated constructs, transferred to 0.1% serum 18 h after transfection and lysed 24 h later. Lysates were analysed for the indicated proteins by immunoblot (n = 2).
Extended Data Fig. 7 Glucose consumption and 2-DG sensitivity in KRAS-mutant human tumour cells with or without exon 4A.
a, Rate of glucose consumption (mean ± s.e.m.; n = 3) in parental (4A+/+) and exon 4A-targeted (4A−/−) A549 cells transfected with Flag–GFP or Flag–KRAS4A(G12V). Representative expression is shown by immunoblot. b, Flux of lactate secretion (mean ± s.e.m.; n = 3) measured over 24 h in A549 cells with the indicated genotype. c, Incorporation of 13C from glucose into lactate (mean ± s.e.m.; n = 3) in A549 cells with the indicated genotype. Significance in a–c was determined by paired Student’s t-test. d, e, Growth inhibition by 2-DG of SUIT2 (d; 48 h) and A549 (e; 24 h) cells with or without the 4A exon of KRAS (mean ± s.e.m.; n = 3; significance by two-way ANOVA). f, g, SUIT2 (f) or A549 (g) cells were used to establish xenograft tumours on the contralateral flanks (4A+/+ versus 4A−/−) of NCG mice. Six weeks later, when the tumours were established and of equivalent size, glucose uptake was measured by 18F-FDG-PET CT scan. f, Left, coronal scan of a representative mouse with SUIT2 xenografts. Glucose uptake is represented by a colour look-up table. Right, SUVs of the entire tumour are plotted (mean ± s.d.; n = 5 mice; significance by two-way ANOVA) as a function of time after 18FDG injection. g, Left, coronal scan of a representative mouse (of n = 5 mice) with A549 xenografts. Right, SUV versus time after injection or that mouse.
Extended Data Fig. 8 Diminished basal ECAR in KRAS-mutant tumour cells in which KRAS exon 4A is disrupted.
a, b, ECAR measured by Seahorse XFe96 of A549 (a) and SUIT2 (b) cells with or without disruption of KRAS exon 4A by CRISPR–Cas9. Oligomycin inhibits oxidative phosphorylation and allows the glycolytic reserve to be measured. 2-DG inhibits glycolysis. Data are mean ± s.e.m. (n = 10 technical replicates).
Extended Data Fig. 9 Quantification of HK1 and total KRAS in tumour cell lines.
a, Top, a standard curve for the quantification of HK1 by immunoblot was generated by titrating recombinant GST–HK1 (rHK1) and probing with a rabbit monoclonal antibody (Cell Signaling Technology, C3534). Bottom, immunoblots of the indicated amounts of lysate (5,000 cells per µl). Calculations based on these results and a molecular mass of 100 kDa indicate 200,000 and 150,000 molecules per cell for A549 and HCT-15 cells, respectively. b, Top, a standard curve for the quantification of total KRAS by immunoblot was generated by titrating recombinant KRAS4B truncated at amino acid 171 and probing with a mouse monoclonal antibody (Sigma-Aldrich, WH0003845M1). Bottom, immunoblots of the indicated amounts of lysate (5,000 cells per µl). Calculations based on these results and a molecular mass of 21 kDa indicate 700,000 and 200,000 molecules per cell for A549 and HCT-15 cells, respectively. The standard curves in a, b were plotted by linear regression using GraphPad Prism v.8.1.1 and the goodness of fit is given as R2.
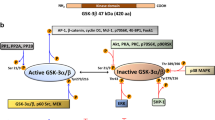
Extended Data Fig. 10 Model of KRAS4A regulation of HK1.
Like all palmitoylated GTPases, KRAS4A cycles between a palmitoylated and a depalmitoylated state. When palmitoylated, the protein has relatively high affinity for the plasma membrane, owing to farnesylation of the C-terminal CAAX sequence and an adjacent polybasic region that operates in conjunction with palmitoylation. After depalmitoylation, KRAS4A loses affinity for the plasma membrane and gains affinity for endomembranes, including the OMM. Tethering of KRAS4A to the OMM allows it to interact with HK1, which is resident on this compartment owing to an N-terminal OMM-targeting sequence and protein–protein interaction with VDAC. Interaction of KRAS4A with HK1 on the OMM decreases allosteric inhibition by glucose-6-phosphate and thereby enhances HK1 activity and glycolytic flux.
Supplementary information
Supplementary Figure 1
This file contains Uncropped SDS-PAGE Blots and Western Blots.
Supplemental Table 1
FLAG-NRAS binding partners in HEK293 cells identified by mass spectrometry.
Rights and permissions
About this article
Cite this article
Amendola, C.R., Mahaffey, J.P., Parker, S.J. et al. KRAS4A directly regulates hexokinase 1. Nature 576, 482–486 (2019). https://doi.org/10.1038/s41586-019-1832-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1832-9
This article is cited by
-
Protein lipidation in cancer: mechanisms, dysregulation and emerging drug targets
Nature Reviews Cancer (2024)
-
From bench to bedside: current development and emerging trend of KRAS-targeted therapy
Acta Pharmacologica Sinica (2024)
-
Endocytic vesicles act as vehicles for glucose uptake in response to growth factor stimulation
Nature Communications (2024)
-
Hypoxia and low-glucose environments co-induced HGDILnc1 promote glycolysis and angiogenesis
Cell Death Discovery (2024)
-
Origin and Evolution of RAS Membrane Targeting
Oncogene (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.