Abstract
Oncogenic activation of RAS is associated with the acquisition of a unique set of metabolic dependencies that contribute to tumour cell fitness. Cells that express oncogenic RAS are able to internalize and degrade extracellular protein via a fluid-phase uptake mechanism termed macropinocytosis1. There is increasing recognition of the role of this RAS-dependent process in the generation of free amino acids that can be used to support tumour cell growth under nutrient-limiting conditions2. However, little is known about the molecular steps that mediate the induction of macropinocytosis by oncogenic RAS. Here we identify vacuolar ATPase (V-ATPase) as an essential regulator of RAS-induced macropinocytosis. Oncogenic RAS promotes the translocation of V-ATPase from intracellular membranes to the plasma membrane via a pathway that requires the activation of protein kinase A by a bicarbonate-dependent soluble adenylate cyclase. Accumulation of V-ATPase at the plasma membrane is necessary for the cholesterol-dependent plasma-membrane association of RAC1, a prerequisite for the stimulation of membrane ruffling and macropinocytosis. These observations establish a link between V-ATPase trafficking and nutrient supply by macropinocytosis that could be exploited to curtail the metabolic adaptation capacity of RAS-mutant tumour cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The full-genome siRNA screen datasets have been deposited at NCBI PubChem under the accession code AID 1347130. Uncropped immunoblot images are available in Supplementary Fig. 1. Datasets that support the findings of this study are available in Source Data. Full list of siRNA and shRNA used in this study are in Supplementary Table 1. Full list of primary antibodies used in this study are in Supplementary Table 2.
References
Commisso, C. et al. Macropinocytosis of protein is an amino acid supply route in Ras-transformed cells. Nature 497, 633–637 (2013).
Kamphorst, J. J. et al. Human pancreatic cancer tumors are nutrient poor and tumor cells actively scavenge extracellular protein. Cancer Res. 75, 544–553 (2015).
Fennell, M., Commisso, C., Ramirez, C., Garippa, R. & Bar-Sagi, D. High-content, full genome siRNA screen for regulators of oncogenic HRAS-driven macropinocytosis. Assay Drug Dev. Technol. 13, 347–355 (2015).
Stransky, L., Cotter, K. & Forgac, M. The function of v-ATPases in cancer. Physiol. Rev. 96, 1071–1091 (2016).
Furuchi, T., Aikawa, K., Arai, H. & Inoue, K. Bafilomycin A1, a specific inhibitor of vacuolar-type H+-ATPase, blocks lysosomal cholesterol trafficking in macrophages. J. Biol. Chem. 268, 27345–27348 (1993).
Iliev, A. I., Djannatian, J. R., Nau, R., Mitchell, T. J. & Wouters, F. S. Cholesterol-dependent actin remodeling via RhoA and Rac1 activation by the Streptococcus pneumoniae toxin pneumolysin. Proc. Natl Acad. Sci. USA 104, 2897–2902 (2007).
del Pozo, M. A. et al. Integrins regulate Rac targeting by internalization of membrane domains. Science 303, 839–842 (2004).
Kozik, P. et al. A human genome-wide screen for regulators of clathrin-coated vesicle formation reveals an unexpected role for the V-ATPase. Nat. Cell Biol. 15, 50–60 (2013).
Plowman, S. J., Muncke, C., Parton, R. G. & Hancock, J. F. H-ras, K-ras, and inner plasma membrane raft proteins operate in nanoclusters with differential dependence on the actin cytoskeleton. Proc. Natl Acad. Sci. USA 102, 15500–15505 (2005).
Li, T. & Chiang, J. Y. L. Regulation of bile acid and cholesterol metabolism by PPARs. PPAR Res. 2009, 501739 (2009).
Garbarino, J. et al. STARD4 knockdown in HepG2 cells disrupts cholesterol trafficking associated with the plasma membrane, ER, and ERC. J. Lipid Res. 53, 2716–2725 (2012).
Capecci, J. & Forgac, M. The function of vacuolar ATPase (V-ATPase) a subunit isoforms in invasiveness of MCF10a and MCF10CA1a human breast cancer cells. J. Biol. Chem. 288, 32731–32741 (2013).
Nishisho, T. et al. The a3 isoform vacuolar type H+-ATPase promotes distant metastasis in the mouse B16 melanoma cells. Mol. Cancer Res. 9, 845–855 (2011).
Chung, C. et al. The vacuolar-ATPase modulates matrix metalloproteinase isoforms in human pancreatic cancer. Lab. Invest. 91, 732–743 (2011).
Deffieu, M. S. & Pfeffer, S. R. Niemann–Pick type C 1 function requires lumenal domain residues that mediate cholesterol-dependent NPC2 binding. Proc. Natl Acad. Sci. USA 108, 18932–18936 (2011).
Breton, S., Lisanti, M. P., Tyszkowski, R., McLaughlin, M. & Brown, D. Basolateral distribution of caveolin-1 in the kidney. Absence from H+-ATPase-coated endocytic vesicles in intercalated cells. J. Histochem. Cytochem. 46, 205–214 (1998).
Breton, S. & Brown, D. Regulation of luminal acidification by the V-ATPase. Physiology (Bethesda) 28, 318–329 (2013).
Chen, Y. et al. Soluble adenylyl cyclase as an evolutionarily conserved bicarbonate sensor. Science 289, 625–628 (2000).
Gorbatenko, A., Olesen, C. W., Boedtkjer, E. & Pedersen, S. F. Regulation and roles of bicarbonate transporters in cancer. Front. Physiol. 5, 130 (2014).
Romero, M. F., Chen, A.-P., Parker, M. D. & Boron, W. F. The SLC4 family of bicarbonate (HCO3 −) transporters. Mol. Aspects Med. 34, 159–182 (2013).
Gorbatenko, A. et al. ErbB2 upregulates the Na+,HCO3 −-cotransporter NBCn1/SLC4A7 in human breast cancer cells via Akt, ERK, Src, and Kruppel-like factor 4. FASEB J. 28, 350–363 (2014).
Haigler, H. T., McKanna, J. A. & Cohen, S. Rapid stimulation of pinocytosis in human carcinoma cells A-431 by epidermal growth factor. J. Cell Biol. 83, 82–90 (1979).
Amyere, M. et al. Constitutive macropinocytosis in oncogene-transformed fibroblasts depends on sequential permanent activation of phosphoinositide 3-kinase and phospholipase C. Mol. Biol. Cell 11, 3453–3467 (2000).
Koivusalo, M. et al. Amiloride inhibits macropinocytosis by lowering submembranous pH and preventing Rac1 and Cdc42 signaling. J. Cell Biol. 188, 547–563 (2010).
Nimnual, A. S., Taylor, L. J., Nyako, M., Jeng, H. H. & Bar-Sagi, D. Perturbation of cytoskeleton dynamics by the opposing effects of Rac1 and Rac1b. Small GTPases 1, 89–97 (2010).
Yeung, T. et al. Receptor activation alters inner surface potential during phagocytosis. Science 313, 347–351 (2006).
Orellana, S. A. & McKnight, G. S. Mutations in the catalytic subunit of cAMP-dependent protein kinase result in unregulated biological activity. Proc. Natl Acad. Sci. USA 89, 4726–4730 (1992).
Franceschini, A. et al. STRING v9.1: protein–protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 41, D808–D815 (2013).
Commisso, C., Flinn, R. J. & Bar-Sagi, D. Determining the macropinocytic index of cells through a quantitative image-based assay. Nat. Protoc. 9, 182–192 (2014).
Bolte, S. & Cordelières, F. P. A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 224, 213–232 (2006).
Pike, J. A., Styles, I. B., Rappoport, J. Z. & Heath, J. K. Quantifying receptor trafficking and colocalization with confocal microscopy. Methods 115, 42–54 (2017).
Kamiyama, D. et al. Versatile protein tagging in cells with split fluorescent protein. Nat. Commun. 7, 11046 (2016).
Sato, I. et al. Differential trafficking of Src, Lyn, Yes and Fyn is specified by the state of palmitoylation in the SH4 domain. J. Cell Sci. 122, 965–975 (2009).
Walsh, A. B. & Bar-Sagi, D. Differential activation of the Rac pathway by Ha-Ras and K-Ras. J. Biol. Chem. 276, 15609–15615 (2001).
Pylayeva-Gupta, Y., Lee, K. E., Hajdu, C. H., Miller, G. & Bar-Sagi, D. Oncogenic Kras-induced GM-CSF production promotes the development of pancreatic neoplasia. Cancer Cell 21, 836–847 (2012).
Badea, L., Herlea, V., Dima, S. O., Dumitrascu, T. & Popescu, I. Combined gene expression analysis of whole-tissue and microdissected pancreatic ductal adenocarcinoma identifies genes specifically overexpressed in tumor epithelia. Hepatogastroenterology 55, 2016–2027 (2008).
Pei, H. et al. FKBP51 affects cancer cell response to chemotherapy by negatively regulating Akt. Cancer Cell 16, 259–266 (2009).
Acknowledgements
We thank R. Garippa and M. Fennell (Memorial Sloan Kettering Cancer Center) for their help with RNAi screen data analysis; members of the Bar-Sagi laboratory for their comments and discussions and M. Philips for sharing cDNA constructs. This work was supported by a grant from The Lustgarten Foundation and National Institutes of Health (NIH)/National Cancer Institute (NCI) (CA210263/CA055360) to D.B.-S. C.R. was supported by a grant from NIH (5 T32 GM007238), A.D.H. was supported by a grant from NIH/NCI (T32CA009161) and E.A.V. was supported by a Canadian Institutes of Health Research Fellowship (146792).
Author information
Authors and Affiliations
Contributions
C.R., A.D.H. and D.B.-S. conceived the cell biological experiments. C.R. carried out the macropinocytic assays, immunofluorescence and microscopy. A.D.H. carried out the biochemical assays. C.R. and A.D.H. carried out the xenograft experiments. E.A.V. conceived and carried out the human gene expression analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 V-ATPase is required for RAS-induced macropinocytosis.
a, Functional clusters within the macropinocytosis screen hits defined by STRING analysis (pink, primary screen; red, primary and confirmation screen). b, Quantification of TMR–dextran uptake following knockdown of the indicated V-ATPase subunits in HeLa HRAS(G12V) cells (HV12). c, Immunoblot of V1A expression from whole-cell lysate (vinculin loading control). d, Effect of V-ATPase depletion (siV1A) on cholesterol localization in Hela HRAS(G12V) cells. Fluorescence micrographs of filipin staining (left), membrane labelling with R-pre (a transfected construct containing a modified sequence of the membrane targeting domain of KRAS linked to RFP, middle), merge of filipin and R-pre with boxed areas enlarged to show plasma membrane localization (right), and the quantification of the ratio of filipin to R-pre membrane localization (bar graph) in control (siCtl) or V1A-knockdown conditions. e, Effect of V1A knockdown on total cholesterol in HeLa HRAS(G12V) (left) and KRAS(G12V) (KV12) (right) cells. f, Effect of bafilomycin A1 (BafA1) and rescue by exogenous cholesterol on the localization of cholesterol and RAC1 in HeLa HRAS(G12V) cells. Fluorescence micrographs of filipin (top), GFP–RAC1 (bottom) and quantification of relative surface GFP–RAC1 (bar graph). g, Effect of oncogenic RAS and V1A expression on RAC1 localization. Immunoblots of RAC1 and V1A in the plasma membrane fraction and whole-cell lysate from HeLa T7-vector control (Ctl) and HRAS(G12V) or oncogenic KRAS cell lines with or without V1A knockdown. Images (d, f) and immunoblots (c, g) are representative of three biological replicates. In f, the dashed lines delineate the cell and nucleus and data (mean ± s.e.m.) represent the fraction of cells that display plasma membrane localization of cholesterol. Scale bars, 10 μm. At least 500 (b, f) and 20 (d) cells were quantified in biological replicates (n = 3). In e, cholesterol quantification is representative of four biological replicates. All data are mean ± s.e.m. for the indicated sample size; unpaired two-tailed Student’s t-test. Gel source data for c, g are shown in Supplementary Fig. 1.
Extended Data Fig. 2 Plasma membrane-localized RAC1 is required for RAS-induced macropinocytosis.
a, b, Effect of V1A expression on RAC1 activity. a, Immunoblots of RAC1 wild type and RAC1(L61) activity (GST–PBD, pull-down of GFP–RAC1–GTP; vinculin loading control) with or without V1A knockdown in HeLa HRAS(G12V) cells (Ctl, GFP). b, Immunoblot of endogenous RAC1 activity (GST–PBD, pull-down of RAC1–GTP; tubulin loading control) in HeLa Ctl and HRAS(G12V) cells with or without V1A knockdown. c, Effect of V-ATPase depletion (siV1A) and rescue by plasma membrane-targeted RAC1 (GFP–RAC1(L61) K-tail) on RAC1(L61) localization and macropinocytosis in HeLa HRAS(G12V) cells. Fluorescence micrographs of GFP–RAC1 (top), TMR–dextran uptake (bottom) and quantification of TMR–dextran (bar graph). d, e, Effect of V-ATPase or RAC1 depletion on macropinocytosis. Quantification of TMR–dextran uptake following bafilomycin A1 treatment (d) or RAC1 knockdown (e) in the absence or presence of exogenous cholesterol in HeLa HRAS(G12V) cells. f, Fluorescence micrographs of cholesterol localization (filipin, top), GFP–RAC1 localization (middle) and TMR–dextran uptake (bottom) in V-ATPase-depleted HeLa KRAS(G12V) cells in the absence or presence of exogenous cholesterol. g, Quantification of TMR–dextran uptake in mutant RAS cells with V1A knockdown and rescue by plasma membrane-targeted RAC1 (GFP–RAC1(L61) K-tail). Images (c, f) and immunoblots (a, b) are representative of three biological replicates. Data in c are mean ± s.e.m. representing the fraction of cells that display plasma membrane localization of GFP–RAC1(L61) or GFP–RAC1(L61) K-tail. In f, the dashed lines delineate the cell and nucleus and data are mean ± s.e.m. representing the fraction of cells that display plasma membrane localization of cholesterol (top) or GFP–RAC1 (middle). Scale bars, 10 μm. At least 500 (c–g) cells were quantified in each biological replicate (n = 3); data are mean ± s.e.m.; unpaired two-tailed Student’s t-test. Gel source data for a, b are shown in Supplementary Fig. 1.
Extended Data Fig. 3 Plasma membrane V-ATPase regulates cholesterol distribution and macropinocytosis.
a, Fluorescence micrographs of HeLa control and HeLa HRAS(G12V) or T7-KRAS(G12V) cells immunostained with V1A antibody. b, Validation of V1A antibody for immunofluorescence. Fluorescence micrographs of V1A immunostaining of HeLa HRAS(G12V) cells transfected with the indicated siRNAs. c, Fluorescence micrographs of V1A immunostaining and filipin labelling of HeLa KRAS(G12V) cells transfected with the indicated siRNAs. d, Quantification of TMR–dextran uptake in mutant RAS cell lines transfected with siV0a3 in the presence or absence of exogenous cholesterol. Images (a–c) are representative of three biological replicates. Scale bars, 10 μm. In b, c, dashed lines delineate the cell and/or nucleus. Data in a, c, are mean ± s.e.m. representing the fraction of cells that display V1A plasma membrane localization. At least 500 (a, c, d) cells were quantified in each biological replicate (n = 3); data are mean ± s.e.m.; unpaired two-tailed Student’s t-test.
Extended Data Fig. 4 Plasma membrane V-ATPase feeds into Rab7-positive endosomes.
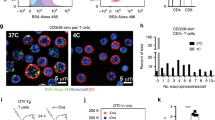
a, b, Fluorescence micrographs of HeLa control and HRAS(G12V) cells with the indicated treatment showing V1A (green) and Rab7 (red) immunostaining. Representative original image used to calculate Mander’s overlap coefficient (a) and processed image (b). c, d, Quantification of Rab7 with V1A colocalization in HeLa cells treated as indicated using Mander’s overlap coefficient (M coefficient). e, f, Epifluorescence imaging of self-complementing GFP from HeLa HRAS(G12V) cells transfected with V1A–GFP11 and plasma membrane-targeting Lyn–GFP(1–10). Positive fluorescence indicates V1A localization to the plasma membrane. e, Fluorescence micrographs of time-lapse imaging. The boxed area of the cell (left) was enlarged (four right images) to show plasma membrane V-ATPase being internalized over time and forming a vesicle (arrow). Time (in s) is shown on each micrograph. f, Fluorescence micrographs of self-complementing GFP (left), immunofluorescence of Rab7 (middle) and merge of V1A and Rab7 (right) with boxed area enlarged beneath the image to show Rab7 colocalization with plasma membrane-derived V1A. In a, b, images are representative of three biological replicates. For c, d, 25 cells were quantified in each biological replicate (n = 3); data are mean ± s.e.m.; unpaired two-tailed Student’s t-test. Scale bars, 10 μm.
Extended Data Fig. 5 sAC–PKA pathway is necessary for oncogenic RAS-induced macropinocytosis.
a–g, Effect of sAC (KH7) or PKA (H89) inhibition on membrane cholesterol, RAC1 activation and RAC1 localization. a, Fluorescence micrographs of HeLa HRAS(G12V) cells in indicated treatments with filipin labelling (top), membrane labelling with R-pre (middle), and merge of filipin and R-pre fluorescence micrographs with boxed areas enlarged beneath the image to show plasma membrane localization (bottom). b, Quantification of the ratio of filipin to R-pre membrane localization with the indicated treatments in HeLa HRAS(G12V) cells. c, Immunoblot of endogenous RAC1 activity (GST–PBD, pull-down of RAC1–GTP; tubulin loading control) in HeLa control and HRAS(G12V) cells treated as indicated. d, Immunoblot of endogenous RAC1 in the plasma membrane fraction and whole-cell lysate in HeLa control and HRAS(G12V) cells treated as indicated. e, Fluorescence micrographs of cholesterol (filipin) distribution in HeLa control cells with indicated treatments. f, Quantification of plasma membrane cholesterol in HeLa control cells with the indicated treatments. g, Fluorescence micrographs of cholesterol (filipin) distribution in BxPC-3 cells in the absence or presence of ectopically expressed KRAS(G12V) treated as indicated. g, Data are mean ± s.e.m. representing the fraction of cells that display cholesterol plasma membrane localization. Images (a, e, g) and immunoblots (c, d) are representative of three biological replicates. Scale bars, 10 μm. At least 20 (b), 50 (f) and 500 (g) cells were quantified in each biological replicate (n = 3); data are mean ± s.e.m.; unpaired two-tailed Student’s t-test. Gel source data for c, d are shown in Supplementary Fig. 1.
Extended Data Fig. 6 PKA activation rescues RAS-induced macropinocytosis from sAC inhibition.
a, b, Effect of sAC inhibition (KH7) and rescue (KH7 + CA-PKA) on macropinocytosis, V-ATPase localization and cholesterol distribution in HeLa HRAS(G12V) cells. a, Quantification of TMR–dextran uptake following indicated treatments. b, Fluorescence micrographs of V1A immunostaining and filipin labelling following indicated treatments. c, d, Effect of sAC inhibition (KH7), plasma membrane V-ATPase inhibition (siV0a3), and rescue (KH7 + CA-PKA + siV0a3) on macropinocytosis, V-ATPase localization and cholesterol distribution in HeLa HRAS(G12V) cells. c, Quantification of TMR–dextran uptake following indicated treatments. d, Fluorescence micrographs of V1A immunostaining and filipin labelling following indicated treatments. e, Effect of KRAS inhibition (siKRAS) and rescue (siKRAS + CA-PKA) on macropinocytosis by quantification of TMR–dextran uptake in mutant RAS cell lines. Images in b, d are representative of three biological replicates. In b, d, dashed lines delineate the cell and nucleus; data are mean ± s.e.m. representing the fraction of cells that display V1A plasma membrane localization. Scale bars, 10 μm. At least 500 cells (a, c, e) were quantified in each biological replicate (n= 3); data are mean ± s.e.m.; unpaired two-tailed Student’s t-test.
Extended Data Fig. 7 The SLC4 family is required for PKA-dependent, RAS-induced macropinocytosis.
a, Quantification of TMR–dextran uptake in the absence (bicarbonate-free) or presence (bicarbonate) of extracellular bicarbonate in mutant RAS cells. b, Fluorescence micrographs of V1A immunostaining following treatment of mutant RAS cells with vehicle or pan-SLC4 inhibitor (S0859). c, Fluorescence micrographs of HeLa HRAS(G12V) cells in indicated treatments with V1A immunostaining (top), membrane labelling with R-pre (middle), and merge of V1A and R-pre fluorescence micrographs with boxed areas enlarged beneath the image to show plasma membrane localization (bottom). d, Quantification of the ratio of V1A to R-pre membrane localization with the indicated treatments in HeLa HRAS(G12V) cells from c. e, f, Effect of SLC4 inhibition (S0859) and rescue (S0859 + CA-PKA) in HeLa HRAS(G12V) cells. e, Fluorescence micrographs of V1A immunostaining and filipin labelling following indicated treatments. Dashed lines delineate the cell and nucleus. f, Quantification of TMR–dextran uptake. g, Fluorescence micrographs of cholesterol (filipin) distribution in BxPC-3 cells in the absence or presence of ectopically expressed KRAS(G12V) treated with SLC4 family inhibitor (S0859). h, mRNA levels of SLC4A7 expression in HeLa control, HRAS(G12V) or KRAS(G12V) cells. i, mRNA levels of SLC4A7 expression following KRAS knockdown in MIA-PaCa-2 cells. j, Effect of PI3K (LY294002, left) or MEK (U0126, right) inhibition on SLC4A7 expression in HeLa control and HRAS(G12V) cells. Immunoblots of SLC4A7 expression from whole-cell lysate (vinculin loading control). p-AKT (left) and p-ERK (right) immunoblots show inhibition of pathways by the indicated treatments. k, Effect of PKA (H89), NHE (EIPA), SLC4A7 (siSLC4A7) and V1A (siV1A) inhibition on submembranous pH (pHsm) in HeLa HRAS(G12V) cells transfected with SEpHluorin–mCherry construct (genetically encoded ratiometric pH probe that is targeted to the inner leaflet of the plasma membrane). Calibration curve of SEpHluorin–mCherry (line graph) was performed with K+ nigericin buffer. Quantification of submembranous pH with H89 and EIPA treatment (bar graph, middle) or with knockdown of SLC4A7 and V1A (bar graph, right). l, Immunoblot of SLC4A7 (vinculin loading control) in MIA-PaCa2 cells with doxycycline-inducible SLC4A7 depletion. m, n, Effect of doxycycline-inducible SLC4A7 depletion in BxPC-3 cells on tumour growth. m, Immunoblot of SLC4A7 expression from whoe-cell lysate (vinculin loading control). n, Waterfall plots of xenografts treated as shown relative to baseline. Each bar represents a tumour. Images (b, c, e, g), immunoblots (j, l, m), and mRNA levels (h, i) are representative of three biological replicates. b, e, g, Data are mean ± s.e.m. representing the fraction of cells that display V1A (b, e) or cholesterol (g) plasma membrane localization. Scale bars, 10 μm. At least 20 (d) and 500 (a, b, e–g) cells were quantified in each biological replicate (n = 3); data are mean ± s.e.m.; unpaired two-tailed Student’s t-test. Gel source data for j, l, m are shown in Supplementary Fig. 1.
Supplementary information
Supplementary Figure
This file contains the uncropped blots used in the main figures and extended data figures.
Supplementary Table 1
Full list of siRNA and shRNA used in this study.
Supplementary Table 2
Full list of primary antibodies used in this study.
Source data
Rights and permissions
About this article
Cite this article
Ramirez, C., Hauser, A.D., Vucic, E.A. et al. Plasma membrane V-ATPase controls oncogenic RAS-induced macropinocytosis. Nature 576, 477–481 (2019). https://doi.org/10.1038/s41586-019-1831-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1831-x
This article is cited by
-
Lysosomes as coordinators of cellular catabolism, metabolic signalling and organ physiology
Nature Reviews Molecular Cell Biology (2024)
-
Combinatorial strategies to target RAS-driven cancers
Nature Reviews Cancer (2024)
-
Acid–base transporters in the context of tumor heterogeneity
Pflügers Archiv - European Journal of Physiology (2024)
-
Nutlin-3a induces KRAS mutant/p53 wild type lung cancer specific methuosis-like cell death that is dependent on GFPT2
Journal of Experimental & Clinical Cancer Research (2023)
-
New Developments in Treating RAS-Mutated Metastatic Colorectal Cancer
Current Treatment Options in Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.