Abstract
Mitochondria are essential regulators of cellular energy and metabolism, and have a crucial role in sustaining the growth and survival of cancer cells. A central function of mitochondria is the synthesis of ATP by oxidative phosphorylation, known as mitochondrial bioenergetics. Mitochondria maintain oxidative phosphorylation by creating a membrane potential gradient that is generated by the electron transport chain to drive the synthesis of ATP1. Mitochondria are essential for tumour initiation and maintaining tumour cell growth in cell culture and xenografts2,3. However, our understanding of oxidative mitochondrial metabolism in cancer is limited because most studies have been performed in vitro in cell culture models. This highlights a need for in vivo studies to better understand how oxidative metabolism supports tumour growth. Here we measure mitochondrial membrane potential in non-small-cell lung cancer in vivo using a voltage-sensitive, positron emission tomography (PET) radiotracer known as 4-[18F]fluorobenzyl-triphenylphosphonium (18F-BnTP)4. By using PET imaging of 18F-BnTP, we profile mitochondrial membrane potential in autochthonous mouse models of lung cancer, and find distinct functional mitochondrial heterogeneity within subtypes of lung tumours. The use of 18F-BnTP PET imaging enabled us to functionally profile mitochondrial membrane potential in live tumours.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
03 January 2020
An Amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Mitchell, P. & Moyle, J. Evidence discriminating between the chemical and the chemiosmotic mechanisms of electron transport phosphorylation. Nature 208, 1205–1206 (1965).
Morais, R. et al. Tumor-forming ability in athymic nude mice of human cell lines devoid of mitochondrial DNA. Cancer Res. 54, 3889–3896 (1994).
Cavalli, L. R., Varella-Garcia, M. & Liang, B. C. Diminished tumorigenic phenotype after depletion of mitochondrial DNA. Cell Growth Differ. 8, 1189–1198 (1997).
Madar, I. et al. Characterization of uptake of the new PET imaging compound 18F-fluorobenzyl triphenyl phosphonium in dog myocardium. J. Nucl. Med. 47, 1359–1366 (2006).
Weinberg, F. et al. Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc. Natl Acad. Sci. USA 107, 8788–8793 (2010).
Ji, H. et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 448, 807–810 (2007).
Shackelford, D. B. et al. LKB1 inactivation dictates therapeutic response of non-small cell lung cancer to the metabolism drug phenformin. Cancer Cell 23, 143–158 (2013).
Madar, I. et al. Characterization of membrane potential-dependent uptake of the novel PET tracer 18F-fluorobenzyl triphenylphosphonium cation. Eur. J. Nucl. Med. Mol. Imaging 34, 2057–2065 (2007).
Smith, R. A., Hartley, R. C. & Murphy, M. P. Mitochondria-targeted small molecule therapeutics and probes. Antioxid. Redox Signal. 15, 3021–3038 (2011).
Kim, D. Y. et al. Evaluation of a mitochondrial voltage sensor, (18F-fluoropentyl)triphenylphosphonium cation, in a rat myocardial infarction model. J. Nucl. Med. 53, 1779–1785 (2012).
Madar, I. et al. Detection and quantification of the evolution dynamics of apoptosis using the PET voltage sensor 18F-fluorobenzyl triphenyl phosphonium. J. Nucl. Med. 50, 774–780 (2009).
Logan, A. et al. Assessing the mitochondrial membrane potential in cells and in vivo using targeted click chemistry and mass spectrometry. Cell Metab. 23, 379–385 (2016).
Waldmann, C. M. et al. An automated multidose synthesis of the potentiometric PET probe 4-[18F]fluorobenzyl-triphenylphosphonium ([18F]FBnTP). Mol. Imaging Biol. 20, 205–212 (2017).
Dykens, J. A. et al. Biguanide-induced mitochondrial dysfunction yields increased lactate production and cytotoxicity of aerobically-poised HepG2 cells and human hepatocytes in vitro. Toxicol. Appl. Pharmacol. 233, 203–210 (2008).
Li, F. et al. LKB1 inactivation elicits a redox imbalance to modulate non-small cell lung cancer plasticity and therapeutic response. Cancer Cell 27, 698–711 (2015).
Giordano, S., Lee, J., Darley-Usmar, V. M. & Zhang, J. Distinct effects of rotenone, 1-methyl-4-phenylpyridinium and 6-hydroxydopamine on cellular bioenergetics and cell death. PLoS One 7, e44610 (2012).
Singer, T. P. & Ramsay, R. R. The reaction sites of rotenone and ubiquinone with mitochondrial NADH dehydrogenase. Biochim. Biophys. Acta 1187, 198–202 (1994).
Caboni, P. et al. Rotenone, deguelin, their metabolites, and the rat model of Parkinson’s disease. Chem. Res. Toxicol. 17, 1540–1548 (2004).
Bridges, H. R., Jones, A. J., Pollak, M. N. & Hirst, J. Effects of metformin and other biguanides on oxidative phosphorylation in mitochondria. Biochem. J. 462, 475–487 (2014).
Owen, M. R., Doran, E. & Halestrap, A. P. Evidence that metformin exerts its anti-diabetic effects through inhibition of complex 1 of the mitochondrial respiratory chain. Biochem. J. 348, 607–614 (2000).
Wheaton, W. W. et al. Metformin inhibits mitochondrial complex I of cancer cells to reduce tumorigenesis. eLife 3, e02242 (2014).
Sanchez-Rangel, E. & Inzucchi, S. E. Metformin: clinical use in type 2 diabetes. Diabetologia 60, 1586–1593 (2017).
Birsoy, K. et al. Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides. Nature 508, 108–112 (2014).
Hensley, C. T. et al. Metabolic heterogeneity in human lung tumors. Cell 164, 681–694 (2016).
de Bruin, E. C. et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346, 251–256 (2014).
Momcilovic, M. et al. The GSK3 signaling axis regulates adaptive glutamine metabolism in lung squamous cell carcinoma. Cancer Cell 33, 905–921.e905 (2018).
Momcilovic, M. et al. Heightening energetic stress selectively targets LKB1-deficient non-small cell lung cancers. Cancer Res. 75, 4910–4922 (2015).
Su, C. Y., Chang, Y. C., Yang, C. J., Huang, M. S. & Hsiao, M. The opposite prognostic effect of NDUFS1 and NDUFS8 in lung cancer reflects the oncojanus role of mitochondrial complex I. Sci. Rep. 6, 31357 (2016).
Molina, J. R. et al. An inhibitor of oxidative phosphorylation exploits cancer vulnerability. Nat. Med. 24, 1036–1046 (2018).
Wittig, I., Braun, H. P. & Schägger, H. Blue native PAGE. Nat. Protocols 1, 418–428 (2006).
Acknowledgements
We thank C. Zamilpa, D. Abeydeera, W. Ladno, O. Sergeeva, D. Williams and T. Olafsen for assistance with PET/CT imaging of the mice. We thank M. Cilluffo, R. McMickle, V. Muhunthan, M. Han and E. Assali for laboratory assistance. We thank the Translational Pathology Core Laboratory at UCLA’s DGSOM for assistance with tumour sample preparation and processing. This research was supported by the NIH National Center for Advancing Translational Science (NCATS) UCLA CTSI Grant number UL1TR001881. D.B.S. was supported by the UCLA CTSI KL2 Translational Science Award grant numbers KL2TR001882 at the UCLA David Geffen School of Medicine, the UCLA Jonsson Comprehensive Cancer Center grant P30 CA016042, the Department of Defense LCRP grant number W81XWH-13-1-0459 and the NIH/NCI R01 CA208642-01. S.T.B. was supported by an NIH T32 training grant HL072752. A.J. was supported by a USHHS Ruth L. Kirschstein Institutional National Research Service Award T32 CA009056. G.A. and A.G. were supported by an NIH/NCI R01 CA208642-01 diversity supplement. J.T.L. was supported by NIH/NCI P30 CA016042. This research was supported by the Joyce and Saul Brandman Fund for Medical Research. We thank the Scott family and the Carrie Strong Foundation as well as B. and D. Goldfarb for their support.
Author information
Authors and Affiliations
Contributions
D.B.S. conceived the study and wrote the manuscript. D.B.S., M.M., S.T.B. and A.J. designed experiments. M.M., S.T.B., A.J. and J.T.L. performed and analysed PET–CT imaging. S.T.B. and D.S. analysed biodistribution data. M.M. and S.T.B. induced tumours in KL mice using Adeno-Cre and/or Lenti-PGK-Cre. S.T.B. and M.M. created KPL cell lines. M.M. and S.T.B. treated mice. M.M., R.L. and A.J. performed transthoracic injections. G.A. performed a portion of the computed tomography scans. M.M. and S.T.B. performed immunohistochemistry staining and analysed data. M.M., J.T.L. and A.J. performed in vitro 18F-BnTP uptake assays. M.M. performed TMRE staining and western blots. M.C.F. is a board-certified anatomic pathologist who performed the pathological analysis. L.S. and O.S. performed and guided respirometry experiments. S.S., C.M.W., A.G. and T.H. performed radiotracer synthesis experiments. D.V.D. and C.M.K. performed biochemical analysis of mitochondria. E.S. and H.C. performed metabolic analysis of lung tumours. S.M.D. contributed resources and critical feedback on the project.
Corresponding author
Ethics declarations
Competing interests
S.M.D. is an advisory board member for EarlyDx Inc., T-Cure Bioscience Inc., Cynvenio Biosystems Inc. and the Johnson and Johnson Lung Cancer Initiative. D.B.S., M.M. and S.S. have filed a provisional patent US 62/901,947 and are listed as inventors for the use of 18F-BnTP to guide use of complex I inhibitors for the treatment of lung cancer.
Additional information
Peer review information Nature thanks Kevin Brindle, Ralph Deberardinis and Jared Rutter for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mitochondrial markers in KL mouse lung tumours.
Whole-cell lysates from lung tumours isolated from KL mice were immunoblotted with the indicated antibodies. Tumours with high levels of the ratio of SP-C to actin (>0.5) were defined as ADCs (blue box), whereas tumours with low SP-C to actin ratios (<0.5) were defined as SCCs (red box). Each lane represents an individual tumour isolated from KL mice. Western blot was done on 20 individual tumours isolated from KL mice from three independent experiments.
Extended Data Fig. 2 Measuring mitochondrial membrane potential in vitro in A549 and L3161C cells.
a, Gating strategy used for the quantification of TMRE signal. The R2 region representing single cells was used for quantification of the TMRE signal. b, Overlay histogram showing shifts in TMRE staining in L3161C cells treated with vehicle, 8 μM oligomycin or 8 μM oligomycin plus 4 μM FCCP. c, TMRE measurements in A549 cells treated with the indicated concentrations of phenformin or FCCP for 3 h (n = 3 biological replicates). d, TMRE measurements in mouse cell line L3161C treated with the indicated concentrations of phenformin or FCCP for 3 h (n = 3 biological replicates). e, Viability of A549 cells treated with the indicated concentrations of phenformin for 3 h (n = 3 biological replicates). f, Uptake of 18F-BnTP probe measured by gamma counter in A549 cells treated with 1 mM phenformin for 3 h (n = 5 biological replicates). g, OCR per cell measured in A549 cells treated acutely with 1 mM phenformin (n = 25 technical replicates). h, OCR per cell measured in mouse cell line L3161C treated acutely with 1 mM phenformin (n = 25 technical replicates). i, TMRE measurements in mouse cell line L3161C treated with vehicle, 8 μM oligomycin, or 8 μM oligomycin with 4 μM FCCP for 3 h (n = 3 biological replicates). j, Uptake of 18F-BnTP probe measured by gamma counter in mouse L3161C cells treated with vehicle, 8 μM oligomycin, or 8 μM oligomycin with 4 μM FCCP for 3 h (n = 6 biological replicates). k, Viability of L3161C cells treated as in j (n = 6 biological replicates). Data are mean ± s.d. Experiments in c–i, were repeated twice with similar results. Experiments in j and k were done once.
Extended Data Fig. 3 Short-term treatment with phenformin does not lead to changes in proliferation or apoptosis.
a, Transverse 18F-BnTP PET–CT overlay (left) of mouse lung (middle) after treatment with phenformin. H&E staining of a lung lobe with an ADC tumour (right). b, Representative slides stained with H&E (left), CC3 (middle) and Ki67 (right), from tumours from KL mice treated with vehicle (top) or phenformin (bottom). Experiment was performed once on slides from n = 5 (vehicle) and n = 6 (phenformin) mouse lungs. c, d, Quantification of staining for Ki67 (c) and CC3 (d) for tumours from KL mice treated with vehicle (n = 5 mice) or phenformin (n = 6 mice). Experiment was performed once. e, Phenformin in lung tumours isolated form KL mice was quantified using liquid chromatography–mass spectroscopy. Tumours were isolated from mice treated with vehicle (n = 6) or 100 mg kg−1 (n = 2) or 200 mg kg−1 phenformin (n = 2) for 5 days. Experiment was performed once. f, Representative 18F-BnTP PET–CT overlay of a tumour formed by transthoracically implanted L3161C lung cells into syngeneic recipient mice. This image is representative of at least 20 PET–CT images. g, H&E slide of a tumour formed as in f. h, Higher magnification image of H&E staining of tumour formed by L3161C mouse cell line as in f. j, Representative slides stained with Ki67 (top), and CC3 (bottom) from tumours formed by transthoracically transplanted L3161C cells that were treated with vehicle, metformin or phenformin. Experiment was performed once on slides from n = 8 (vehicle), n = 5 (metformin) or n = 6 (phenformin) tumours. Data are mean ± s.d. P values determined by unpaired two-tailed t-test.
Extended Data Fig. 4 Expressing ND1 in mouse L3161C lung ADC cell line reduces sensitivity of mitochondrial membrane potential to phenformin in vitro and in vivo.
a, Basal OCR rate per cell for L3161C cells expressing empty vector (pBabe; black) (n = 12 technical replicates) or L3161C cells expressing ND1 (L3161C-ND1; red) (n = 12 technical replicates) treated with 50 μM phenformin for 24 h. b, Basal OCR rate per cell for L3161C-pBabe (black) (n = 12 technical replicates for all conditions, except n = 6 for 250 μM and n = 9 for 500 μM phenformin) and L3161C-ND1 cells (red) (n = 12 technical replicates) treated with the indicated concentrations of phenformin for 24 h. Data are mean ± s.d. c, Waterfall plot of the percentage change in maximum uptake of 18F-BnTP after treatment relative to before treatment for mice transthoracically implanted with L3161C cells expressing empty vector (pBabe; n = 5 mice) or ND1 (n = 5 mice) and treated with 125 mg kg−1 phenformin for 5 days. d, Waterfall plot of the percentage change in maximum uptake of 18F-BnTP in tumours formed by transthoracically implanted L3161C-pBabe (n = 3 mice) or L3161C-ND1 cells (n = 5 mice) treated with vehicle for 5 days. Experiments in a–d were performed once. P values determined by unpaired two-tailed t-test.
Extended Data Fig. 5 Multi-tracer imaging and immunohistochemistry markers in lung tumours from KL mice.
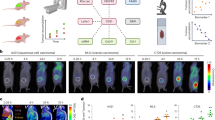
a, Representative PET and computed tomography (CT) images of three KL mice imaged with 18F-BnTP (top) and 18F-FDG (bottom) on sequential days. H, heart; T, tumour. Arrows and circles denote tumours. b, Whole lung slides stained with H&E, TOM20, GLUT1, or CK5 plus TTF1 from three mice. Scale bars, 5 mm. c, Representative higher magnification images of the tumours circled in b stained with H&E, TOM20, GLUT1, CK5 plus TTF1 as indicated. Scale bars, 25 μm. Data are representative of three independent mouse experiments.
Extended Data Fig. 6 PET–CT and biochemical analysis of KL tumours.
a, Crystal structure of complex I (PDB accession 5lc5), with NDUFS1 and NDUFV1 subunits in red, and FeS clusters in yellow.b–d, PET–CT images from three KL mice that were imaged on sequential days with 18F-BnTP (top) and 18F-FDG (bottom). Tumours are circled. Maximum uptake value for each tumour after normalization to maximum uptake of the heart is indicated. e, Western blot analysis from lung nodules that were isolated from mice imaged in b–d. Two lung tumours from mouse 5372 (imaged in b) are shown—T1 in blue (low 18F-FDG and GLUT1 levels; high 18F-BnTP, NDUFS1 and NDUFV1 levels); and T2 in red (high 18F-FDG and GLUT1 levels; low 18F-BnTP, NDUFS1 and NDUFV1 levels). Experiments in b–d are representative of three independent mouse experiments. Experiment in e was performed once.
Extended Data Fig. 7 Levels of NDUFS1 and NDUFV1 in KL tumours.
Whole-cell lysates from lung tumours isolated from KL mice were immunoblotted with the indicated antibodies. This western blot was done on 20 individual tumours isolated from KL mice from three independent experiments.
Extended Data Fig. 8 Sensitivity of mouse and human lung cancer cell lines to complex I inhibitors phenformin and IACS-010759.
a, TMRE measurement as determined by flow cytometry comparing mouse ADC (n = 3 biological replicates) and mouse SCC (n = 3 biological replicates) cell lines. b, Cell viability of mouse ADC (n = 3 biological replicates) and mouse SCC (n = 3 biological replicates) cells was measured in the presence of indicated concentrations of phenformin for 48 h. c, Cell viability of human ADC (A549; n = 3 biological replicates) and human SCC (RH2; n = 3 biological replicates) cells was measured in the presence of indicated concentrations of phenformin for 48 h. d, Cell viability of human ADC (A549; n = 3 biological replicates) and human SCC (RH2; n = 3 biological replicates) cells was measured in the presence of indicated concentrations of IACS-010759 for 48 h. Data are mean ± s.d. P values determined by unpaired two-tailed t-test or one-way ANOVA (for b–d). Experiments were repeated twice with similar results.
Extended Data Fig. 9 Characteristics of tumours from KL mice treated with vehicle or IACS-010759.
a, Uptake of 18F-BnTP in tumours from KL mice before the start of treatment with vehicle or 15 mg kg−1 IACS-010759. Each dot represents a tumour; n = 44 (vehicle), n = 66 (IACS) tumours. b, c, H&E staining images from lung sections from KL mice treated with vehicle (b) or 15 mg kg−1 IACS-010759 (c) for 12 days, with tumours delineated by red lines. Quantification of these data is shown in Fig. 4l. Experiment was performed once.
Extended Data Fig. 10 Intra-tumoral heterogeneity in KL mice.
Higher magnification images of tumour shown in Fig. 4n, o with GLUT1 staining (left) and CK5 and TTF1 staining (right). Areas corresponding to ADC and SCC are indicated, with rectangular boxes corresponding to magnified images shown in Fig. 4n. Data are representative of three independent mouse experiments.
Supplementary information
Supplementary Data
This file contains the source data gel scans.
Rights and permissions
About this article
Cite this article
Momcilovic, M., Jones, A., Bailey, S.T. et al. In vivo imaging of mitochondrial membrane potential in non-small-cell lung cancer. Nature 575, 380–384 (2019). https://doi.org/10.1038/s41586-019-1715-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1715-0
This article is cited by
-
Widespread stable noncanonical peptides identified by integrated analyses of ribosome profiling and ORF features
Nature Communications (2024)
-
Active targeting schemes for nano-drug delivery systems in osteosarcoma therapeutics
Journal of Nanobiotechnology (2023)
-
Positron emission tomography imaging of the sodium iodide symporter senses real-time energy stress in vivo
Cancer & Metabolism (2023)
-
TRPM8 promotes hepatocellular carcinoma progression by inducing SNORA55 mediated nuclear-mitochondrial communication
Cancer Gene Therapy (2023)
-
Diversity of mitochondrial networks in lung cancer imaged
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.