Abstract
Ferroptosis is an iron-dependent form of necrotic cell death marked by oxidative damage to phospholipids1,2. To date, ferroptosis has been thought to be controlled only by the phospholipid hydroperoxide-reducing enzyme glutathione peroxidase 4 (GPX4)3,4 and radical-trapping antioxidants5,6. However, elucidation of the factors that underlie the sensitivity of a given cell type to ferroptosis7 is crucial to understand the pathophysiological role of ferroptosis and how it may be exploited for the treatment of cancer. Although metabolic constraints8 and phospholipid composition9,10 contribute to ferroptosis sensitivity, no cell-autonomous mechanisms have been identified that account for the resistance of cells to ferroptosis. Here we used an expression cloning approach to identify genes in human cancer cells that are able to complement the loss of GPX4. We found that the flavoprotein apoptosis-inducing factor mitochondria-associated 2 (AIFM2) is a previously unrecognized anti-ferroptotic gene. AIFM2, which we renamed ferroptosis suppressor protein 1 (FSP1) and which was initially described as a pro-apoptotic gene11, confers protection against ferroptosis elicited by GPX4 deletion. We further demonstrate that the suppression of ferroptosis by FSP1 is mediated by ubiquinone (also known as coenzyme Q10, CoQ10): the reduced form, ubiquinol, traps lipid peroxyl radicals that mediate lipid peroxidation, whereas FSP1 catalyses the regeneration of CoQ10 using NAD(P)H. Pharmacological targeting of FSP1 strongly synergizes with GPX4 inhibitors to trigger ferroptosis in a number of cancer entities. In conclusion, the FSP1–CoQ10–NAD(P)H pathway exists as a stand-alone parallel system, which co-operates with GPX4 and glutathione to suppress phospholipid peroxidation and ferroptosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Conrad, M., Angeli, J. P., Vandenabeele, P. & Stockwell, B. R. Regulated necrosis: disease relevance and therapeutic opportunities. Nat. Rev. Drug Discov. 15, 348–366 (2016).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Friedmann Angeli, J. P. et al. Inactivation of the ferroptosis regulator Gpx4 triggers acute renal failure in mice. Nat. Cell Biol. 16, 1180–1191 (2014).
Zilka, O. et al. On the mechanism of cytoprotection by ferrostatin-1 and liproxstatin-1 and the role of lipid peroxidation in ferroptotic cell death. ACS Cent. Sci. 3, 232–243 (2017).
Shah, R., Shchepinov, M. S. & Pratt, D. A. Resolving the role of lipoxygenases in the initiation and execution of ferroptosis. ACS Cent. Sci. 4, 387–396 (2018).
Stockwell, B. R. et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell 171, 273–285 (2017).
Tarangelo, A. et al. p53 suppresses metabolic stress-induced ferroptosis in cancer cells. Cell Rep. 22, 569–575 (2018).
Kagan, V. E. et al. Oxidized arachidonic and adrenic PEs navigate cells to ferroptosis. Nat. Chem. Biol. 13, 81–90 (2017).
Doll, S. et al. ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat. Chem. Biol. 13, 91–98 (2017).
Wu, M., Xu, L. G., Li, X., Zhai, Z. & Shu, H. B. AMID, an apoptosis-inducing factor-homologous mitochondrion-associated protein, induces caspase-independent apoptosis. J. Biol. Chem. 277, 25617–25623 (2002).
Ingold, I. et al. Selenium utilization by GPX4 is required to prevent hydroperoxide-induced ferroptosis. Cell 172, 409–422 (2018).
Viswanathan, V. S. et al. Dependency of a therapy-resistant state of cancer cells on a lipid peroxidase pathway. Nature 547, 453–457 (2017).
Hangauer, M. J. et al. Drug-tolerant persister cancer cells are vulnerable to GPX4 inhibition. Nature 551, 247–250 (2017).
Tsoi, J. et al. Multi-stage differentiation defines melanoma subtypes with differential vulnerability to drug-induced iron-dependent oxidative stress. Cancer Cell 33, 890–904 (2018).
Angeli, J. P. F., Shah, R., Pratt, D. A. & Conrad, M. Ferroptosis inhibition: mechanisms and opportunities. Trends Pharmacol. Sci. 38, 489–498 (2017).
Horikoshi, N., Cong, J., Kley, N. & Shenk, T. Isolation of differentially expressed cDNAs from p53-dependent apoptotic cells: activation of the human homologue of the Drosophila peroxidasin gene. Biochem. Biophys. Res. Commun. 261, 864–869 (1999).
Bersuker, K. et al. The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit ferroptosis. Nature https://doi.org/10.1038/s41586-019-1705-2 (2019).
Seiler, A. et al. Glutathione peroxidase 4 senses and translates oxidative stress into 12/15-lipoxygenase dependent- and AIF-mediated cell death. Cell Metab. 8, 237–248 (2008).
Eisenhaber, F. et al. Prediction of lipid posttranslational modifications and localization signals from protein sequences: big- Π, NMT and PTS1. Nucleic Acids Res. 31, 3631–3634 (2003).
Borgese, N., Aggujaro, D., Carrera, P., Pietrini, G. & Bassetti, M. A role for N-myristoylation in protein targeting: NADH-cytochrome b 5 reductase requires myristic acid for association with outer mitochondrial but not ER membranes. J. Cell Biol. 135, 1501–1513 (1996).
Mousnier, A. et al. Fragment-derived inhibitors of human N-myristoyltransferase block capsid assembly and replication of the common cold virus. Nat. Chem. 10, 599–606 (2018).
Elguindy, M. M. & Nakamaru-Ogiso, E. Apoptosis-inducing factor (AIF) and its family member protein, AMID, are rotenone-sensitive NADH:ubiquinone oxidoreductases (NDH-2). J. Biol. Chem. 290, 20815–20826 (2015).
Frei, B., Kim, M. C. & Ames, B. N. Ubiquinol-10 is an effective lipid-soluble antioxidant at physiological concentrations. Proc. Natl Acad. Sci. USA 87, 4879–4883 (1990).
Haidasz, E. A., Van Kessel, A. T. & Pratt, D. A. A continuous visible light spectrophotometric approach to accurately determine the reactivity of radical-trapping antioxidants. J. Org. Chem. 81, 737–744 (2016).
Niki, E. Mechanisms and dynamics of antioxidant action of ubiquinol. Mol. Aspects Med. 18, 63–70 (1997).
Mannes, A. M., Seiler, A., Bosello, V., Maiorino, M. & Conrad, M. Cysteine mutant of mammalian GPx4 rescues cell death induced by disruption of the wild-type selenoenzyme. FASEB J. 25, 2135–2144 (2011).
Shimada, K., Hayano, M., Pagano, N. C. & Stockwell, B. R. Cell-line selectivity improves the predictive power of pharmacogenomic analyses and helps identify NADPH as biomarker for ferroptosis sensitivity. Cell Chem. Biol. 23, 225–235 (2016).
Shimada, K. et al. Global survey of cell death mechanisms reveals metabolic regulation of ferroptosis. Nat. Chem. Biol. 12, 497–503 (2016).
Morré, D. J. & Morré, D. M. Non-mitochondrial coenzyme Q. Biofactors 37, 355–360 (2011).
Nyquist, S. E., Barr, R. & Morré, D. J. Ubiquinone from rat liver Golgi apparatus fractions. Biochim. Biophys. Acta 208, 532–534 (1970).
Rees, M. G. et al. Correlating chemical sensitivity and basal gene expression reveals mechanism of action. Nat. Chem. Biol. 12, 109–116 (2016).
Seashore-Ludlow, B. et al. Harnessing connectivity in a large-scale small-molecule sensitivity dataset. Cancer Discov. 5, 1210–1223 (2015).
Basu, A. et al. An interactive resource to identify cancer genetic and lineage dependencies targeted by small molecules. Cell 154, 1151–1161 (2013).
Acknowledgements
This work is supported by the Junior Group Leader program of the Rudolf Virchow Center, University of Würzburg and Deutsche Forschungsgemeinschaft (DFG) FR 3746/3-1 to J.P.F.A., the DFG CO 291/5-2 and CO 291/7-1, the German Federal Ministry of Education and Research (BMBF) through the Joint Project Modelling ALS Disease In vitro (MAIV, 01EK1611B) and the VIP+ program NEUROPROTEKT (03VP04260), as well as the m4 Award provided by the Bavarian Ministry of Economic Affairs, Regional Development and Energy (StMWi) to M.C., the Cancer Research UK (CRUK, grants C29637/A20183 and C29637/A21451) to E.W.T., the European Research Council (LipidArrays) to V.O., the Natural Sciences and Engineering Council of Canada and the Canada Foundation for Innovation to D.A.P. and PhD scholarship by DFG GRK2157 to A.K.
Author information
Authors and Affiliations
Contributions
M.C., J.P.F.A. and S.D. conceived the study and wrote the manuscript. M.A. and V.O. performed (oxi)lipidomics analysis and data interpretation. S.D., B.P., E.P., D.W., F.P.F., J.P.F.A., T.V., V.M., I.I., K.B., M. Sato, M.R., T.N.X.d.S. and M.C.d.S. performed in vitro experiments. R.S. and D.A.P. performed functional characterization of recombinant FSP1. S.D., F.P.F., D.A.P., J.P.F.A. and M.C. performed evaluation and interpretation of the in vitro data. M. Sattler, A.M. and G.M.P. expressed and purified recombinant FSP1. C.H.S. provided TNBC cell lines. A.F. and A. Schepers helped to generate the monoclonal antibodies. B.P. and J.W. carried out screening of FSP1 inhibitors and related structure–activity relationship studies. W.S. and A. Schulze performed liquid chromatography–mass spectrometry analysis of ubiquinone content. A.G.G. and E.W.T. studied myristoylation of FSP1. A.K., M. Sauer, F.P.F. and J.P.F.A. performed enhanced microscopy experiments. All authors read and agreed on the content of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Kivanc Birsoy, Navdeep S. Chandel and Brent R. Stockwell for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification and characterization of FSP1 as an anti-ferroptotic protein.
a, Schematic depicting the generation of a lentiviral cDNA-overexpression library using the total mRNA from MCF7 cells. b, Genomic PCRs of the 14 Pfa1 cell clones that remained clones after the removal of false-positive results using human-specific primers to amplify the human cDNAs of GPX4 (571 bp) or AIFM2 (524 bp). The clones 2, 16, 24, 25, 26, 28 and 30 showed positive PCR results for GPX4 (571 bp). Clones 1, 44, 45, 50, 51, 52 and 53 were positive for AIFM2 (524 bp) as indicated by the white arrows. Data are one of n = 3 independent experiments. c, Immunoblot analysis of ACSL4, HA, GPX4 and β-actin expression in Pfa1 cells stably expressing mock or FSP1–HA. Gpx4 deletion was induced by the administration of TAM for the indicated time period. d, Immunoblot analysis of ACSL4, HA, GPX4 and β-actin expression in wild-type and GPX4-knockout HT1080 cells stably expressing mock, GPX4–FSH or FSP1–HA. e, Proliferation of mock and FSP1–HA Pfa1 cells treated with or without TAM. Cell numbers were assessed every 24 h using a Neubauer haemocytometer. Data are mean ± s.d. of n = 3 wells of a 24-well plate from one representative of two independent experiments. f, g, Dose-dependent toxicity of erastin (f) and l-buthionine sulfoximine (BSO; g), which is an inhibitor of γ-glutamyl-cysteine ligase, in Pfa1 cells expressing mock or FSP1–HA. h, i, Dose-dependent toxicity of erastin (h) and BSO (i) in HT1080 cells expressing mock or FSP1–HA. Cell viability was assessed 48 h (f, h) or 72 h (g, i) after treatments using Aquabluer. Data are mean ± s.d. of n = 3 wells of a 96-well plate from one representative of three (f–i) independent experiments. *P < 0.01; two-way ANOVA. j, Measurement of total glutathione levels in Pfa1 mock, FSP1-expressing and FSP1-expressing Gpx4-knockout cells treated with or without BSO. Data are mean ± s.d. of n = 3 wells of a 96-well plate from one representative of three independent experiments. k, Left, determination of NADPH consumption by glutathione reductase as an indirect measure of the GPX4 activity. Phosphatidylcholine lipid hydroperoxide (PCOOH) was used as a GPX4-specific substrate. Right, cell lysates from mock and FSP1–HA Pfa1 cells treated with or without TAM for 48 h were used for the assay as shown by the immunoblot (FSP1, GPX4 and β-actin). FSP1 was detected using the polyclonal antibody (PA5-24562). Data are mean ± s.d. of n = 3 wells of a 6-well plate from one representative of three independent experiments. Immunoblot images (c, d, k) are cropped from the chemiluminescence signal files. For gel source data (c, d, k) showing the overlap of colorimetric and chemiluminescence signals, see Supplementary Fig. 1.
Extended Data Fig. 2 FSP1 expression does not change the phospholipid composition.
Lipidomic profile (only detectable phospholipid species of phosphatidylethanolamine (PE), phosphatidylcholine (PC), phosphatidylglycerol (PG), phosphatidylinositol (PI) and phosphatidylserine (PS), including plasmenyl (O) and plasmanyl (P) lipids) of mock, FSP1–HA and Gpx4-knockout FSP1–HA Pfa1 cells. Data are the mean values of the area of analyte (A) over the internal standard (IS) per total protein (mg) of n = 4 replicates of one experiment performed independently three times. log10-transformation has been applied to better visualize and compare the abundance of the different phospholipid species in the samples. *P < 0.05; multiple t-test with Sidak–Bonferroni correction for multiple comparisons.
Extended Data Fig. 3 FSP1 is a highly specific anti-ferroptotic protein.
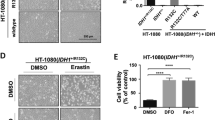
a, Dose-dependent toxicity of phenylarsine oxide, indomethacin, auranofin, ivermectin, sunitinib, obatoclax, mitoxantrone, irinotecan, vinblastine, ABT-263, nocodazole, etoposide, paclitaxel, H2O2 and tert-butyl hydroperoxide (tBOOH) of Pfa1 cells expressing mock or FSP1–HA. Cell viability was assessed 24 h after treatment using Aquabluer. b, Dose-dependent toxicity of TNF and staurosporine of mock and FSP1–HA-expressing Pfa1 cells. Cell viability was assessed 24 h after treatment using Aquabluer. c, Immunoblot analysis (ACSL4, HA, cleaved caspase 3 (clv. Casp3), GPX4 and β-actin) of Pfa1 FSP1–HA cells treated with or without TNF for 6 h. d, Immunoblot analysis of FSP1, ACSL4, p53, p21 and VCP expression in p53 (also known as TP53) wild-type and p53-knockout (CRISPR–CAS9-modified) HT1080 cell lines treated with the MDM2 (MDM2 proto-oncogene) inhibitor Nutlin3 or the cytostatic compound doxorubicin (Doxo). Expression of FSP1 was not altered by Nutlin3 or doxorubicin treatment, whereas the expression of p53 and p21 was strongly induced in HT1080 p53 wild-type cells. Data show one representative of n = 3 independent experiments. e, Flow cytometry analysis of annexin V/propidium iodide staining in Pfa1 cells expressing mock or FSP1–HA treated with or without TNF for 4 h. No difference in the apoptotic activity was observed in cells as visualized in the Alexa Fluor 488/PE–Cy5 channels. Data show one representative experiment of an experiment performed independently twice. f, Immunoblot analysis of AIFM1, ACSL4, GPX4 and β-actin in two different Pfa1 Aifm1-knockout cell clones overexpressing mock or AIFM1. Data show one representative of n = 3 independent experiments. g, Dose-dependent toxicity of RSL3, erastin and BSO in Aifm1-knockout Pfa1 cell clones (1 and 2) overexpressing mock or AIFM1. AIFM1 expression does not affect ferroptosis sensitivity. Data are the mean of n = 3 replicates of a representative experiment performed independently three times. h, Time-dependent lactate dehydrogenase (LDH) release of Pfa1 cells stably expressing mock, FSP1–HA or FSP1(G2A) treated with TAM to induce GPX4 loss. Supernatants were collected from 6-well plates at different time points after TAM induction and assayed for lactate dehydrogenase content in a 96-well plate. i, Wild-type and GPX4-knockout HT1080 cells overexpressing mock, hGPX4–FSH, FSP1–HA or FSP1(G2A)–HA treated with and without 200 nM Lip-1. Cell viability was assessed after 72 h using Aquabluer. Data are the mean ± s.d. of n = 3 wells of a 96-well plate from one representative of three independent experiments (a, b, g–i); *P < 0.01; two-way ANOVA. Immunoblot images (c, d, f) are cropped from the chemiluminescence signal files. For gel source data (c, d, f) showing the overlap of colorimetric and chemiluminescence signals, see Supplementary Fig. 1.
Extended Data Fig. 4 FSP1 protects against unrestrained lipid peroxidation in a COQ2-dependent manner.
a, Enhanced resolution confocal microscopy images demonstrating different localizations of FSP1–GFP and the FSP1(G2A)–GFP mutant in HT1080 cells. DAPI (yellow), GFP (green), endoplasmic reticulum or Golgi tracker (magenta). Scale bars, 20 nm. Data show one representative of n = 3 independently performed experiments. b, Formation of 5-hydro(pero)xyeicosatetraenoic acid (5-H(P)ETE) (multiple reaction monitoring (MRM): 319 → 115), 12-H(P)ETE (MRM: 319 → 179) and 15-H(P)ETE (MRM: 319 → 219) in either mock (black) or FSP1–HA-overexpressing (red) Pfa1 cells treated with 0.2 μM RSL3 and 40 μM arachidonic acid. Hydroperoxides were analysed as their alcohols following reduction with PPh3 (triphenylphosphane) in methanol. Data are the mean of biological triplicates from one representative of n = 3 independently performed experiments. c, Dose-dependent rescue of three independent COQ2-knockout HT1080 cell clones (56, 61 and 68) by supplementation of the cell culture medium with uridine, CoQ10 or decyl-ubiquinone. Cell viability was assed using the Aquabluer assay 48 h after treatment. Data are mean ± s.d. of n = 3 wells of a 96-well plate performed once. d, Immunoblot analysis of FSP1 and β-actin in HT1080 parental (left) and HT1080 COQ2-knockout (56) (right) cells overexpressing FSP1–GFP, FSP1(G2A)–GFP or GFP. Immunoblot images are cropped from the chemiluminescence signal files. For gel source data showing the uncropped chemiluminescence signals, see Supplementary Fig. 1. e, SDS gels showing the different purification steps of recombinant FSP1 from bacterial cell lysates. Left, SDS gel of protein extracts after initial nickel affinity chromatography (E1), the SUMO-tag was cleaved in the eluate by addition of the SUMO protease (dtUD1) and a second round of nickel affinity chromatography was performed to remove the cleaved SUMO-tag as well as uncleaved SUMO–FSP1 and SUMO protease (E2). The flow-through fraction was collected (second nickel). The SUMO–FSP1 fusion protein is visible around 55 kDa and FSP1 at 40.5 kDa. Right, SDS gel showing different fractions containing FSP1 40.5 kDa (A8–A12, B1–B7 and C3–C4) from size-exclusion chromatography of FSP1 after the second nickel-affinity chromatography. Fractions C3 and C4 were used for subsequent assays. One representative of at least three independent experiments.
Extended Data Fig. 5 FSP1 protects against lipid peroxidation by reducing radical-trapping antioxidants.
a, b, Co-autoxidations of STY-BODIPY (1 μM) (a) and the polyunsaturated lipids of chicken egg phosphatidylcholine liposomes (1 mM). The increase in fluorescence of oxidized STY-BODIPY is monitored over the course of the autoxidation, which is initiated using C9-HN (0.2 mM) (b). c, Representative autoxidations inhibited by 50 nM FSP1 (green), 8 μM NADH (purple), 16 μM NADH (orange), 50 nM FSP1 and 8 μM NADH (black) or 50 nM FSP1 and 16 μM NADH (blue) (c). d–j, Analogous representative of inhibited autoxidations to which CoQ10 (d, e), α-tocopherol (α-tocopherol) and CoQ10 (h, i), or α-tocopherol (j) was added. f, g, Recombinant NQO1 failed to suppress autoxidations in a similar manner compared to FSP1 (f, g). k, 1-Palmitoyl-2-linoleoyl-phosphatidylcholine hydroperoxide (PLPC-OOH) produced from the autoxidation of soy lecithin liposomes (13.3 mM), inhibited by FSP1 alone, or in the presence of either 10 μM CoQ10 or 10 μM α-tocopherol and 10 μM CoQ10. PLPC-OOH was measured 0, 60, 120 and 180 min after autoxidation was induced using liquid chromatography–mass spectrometry (MRM: 790 → 184). Data show one of n = 3 representative experiments.
Extended Data Fig. 6 Development of FSP1-specific inhibitors as ferroptosis sensitizer.
a, b, Dose-dependent toxicity of iFSP1in FSP1-overexpressing cells (Pfa1 (a); HT1080 (b)) with or without GPX4 loss. Treatment with the ferroptosis inhibitor Lip-1 (150 nM) protected GPX4-knockout cells from iFSP1-induced ferroptosis. iFSP1 is only toxic to cells that depend solely (no GPX4 expression detectable) on FSP1 function. c, Efficacy of iFSP1 and structurally related analogues; half-maximal effective concentration (EC50) values (mean ± s.d.) of iFSP1 (1) and its derivatives (2–14) calculated from experiments performed at least twice in triplicate are shown in the table with the corresponding chemical structures depicted below. Based on commercially available analogues a preliminary structure–activity relationship study revealed that substitution of the amino position (R1, R2) showed broad tolerability of aliphatic groups and that lipophilic substituents of the phenyl group at the 3 position (R3) in the ortho and meta positions were well tolerated. d, Immunoblot analysis of FSP1 and VCP in parental as well as HT1080 FSP1-overexpressing and FSP1-knockout HT1080 cells. An in-house-generated monoclonal antibody against human FSP1 was used. Immunoblot images are cropped from the chemiluminescence signal files. For gel source data showing the overlap of colorimetric and chemiluminescence signals, see Supplementary Fig. 1. e, Dose-dependent toxicity of RSL3 in a panel of genetically engineered (FSP1-knockout) human cancer cell lines (NCl-H1437, NCl-H1437 FSP1 KO, U-373, U-373 FSP1 KO, MDA-MB-436, MDA-MB-436 FSP1 KO, SW620, SW620 FSP1 KO, MDA-MB-435S, MDA-MB-435S FSP1 KO, A549 and A549 FSP1 KO) treated with or without FSP1 inhibitor (iFSP1) and Lip-1. f, Dose-dependent toxicity of RSL3 in a panel of genetically modified (mouse (mFSP1) and human (hFSP1) FSP1 overexpression) human cancer cell lines (IMR5/75 mock, IMR5/75 hFSP1, 786-O mock, 786-O hFSP1, LOX-IMVI mock, LOX-IMVI hFSP1, HLF mock, HLF hFSP1, U-138 mock and U-138 mFSP1) treated with or without iFSP1 and Lip-1. Data show the mean ± s.d. of n = 3 wells of a 96-well plate from one representative of three (a–c) or two (e, f) independent experiments; *P < 0.0001; two-way ANOVA.
Extended Data Fig. 7 FSP1 is expressed in a wide range of cancer cell lines.
a, Immunoblot analysis of the expression of key ferroptosis players including ACSL4, FSP1, GPX4 and XCT (SLC7A11) in a panel of cancer cell lines from different origins. In addition, genetically modified cancer cell lines in which FSP1 is knocked out (MDA-436-MB FSP1 KO, NCl-H1437 FSP1 KO, U-373 FSP1 KO, MDA-MB-435S FSP1 KO, A549 FSP1 KO and SW620 FSP1 KO) as well as cell lines with lentiviral overexpression of FSP1 (IMR5/75 hFSP1, 786-O hFSP1, LOX-IMVI hFSP1 and HLF hFSP1) are shown. VCP or β-actin served as loading control. MDA-MB-231 was used as reference to compare expression levels in between independent blots. Data show one representative of two independent experiments. Immunoblot images are cropped from the chemiluminescence signal files. For gel source data showing the overlap of colorimetric and chemiluminescence signals, see Supplementary Fig. 1.
Extended Data Fig. 8 iFSP1 sensitizes cancer cell lines from different origins to RSL3-induced ferroptosis.
Dose-dependent toxicity of RSL3 in a panel of human cancer cell lines from different origins (breast, lung, pancreas, brain, liver, kidney, skin and intestine) treated with or without iFSP1 and Lip-1. Data are the mean ± s.d. of n = 3 wells of a 96-well plate from one representative of two independent experiments.
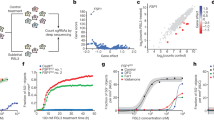
Extended Data Fig. 9 FSP1 expression directly correlates with resistance to ferroptosis and its inhibition selectively sensitizes cells to ferroptosis.
a, Correlation of a panel of 860 cancer cell lines32,33,34. The sensitivity to RSL3, ML162 and ML210 was correlated with gene expression. Genes were plotted according to their Pearson correlation score. FSP1 was the highest ranking gene that correlated with resistance to RSL3 (P = 0.392), ML162 (P = 0.424) and ML210 (P = 0.398). b, Dot plot depicting the correlation of the dependency of a cell on GPX4 (CERES score of −1 means full dependency based on CRISPR–Cas9 knockout screen) and the expression level of FSP1 in a panel of 559 different cancer cell lines (DepMap; https://depmap.org/portal/). Cell lines with high expression of FSP1 were found to be less dependent on GPX4 (Pearson correlation score of 0.366, P = 3.38 × 10−19). c, Dose-dependent toxicity of RSL3 in a panel of human lung cancer cells (NCl-H1437, NCl-H1299, NCl-H1573, NCl-H2126, NCl-H520 and NCl-H661) treated with or without the FSP1 inhibitor iFSP1 (5 μM). Co-treatment of RSL3 and iFSP1 increased the ferroptotic response of all cell lines except in NCl-H1437 cells. d, Dose-dependent toxicity of different cytotoxic compounds (erastin, BSO, RSL3, vinblastine, etoposide, phenylarsine oxide (PAO), mitoxantrone, irinotecan, nocodazole and cisplatin) in Pfa1 mock and FSP1-overexpressing cells treated with or without iFSP1. The protective effect of FSP1 overexpression is lost upon iFSP1 (5 μM) treatment. Data are the mean ± s.d. of n = 3 wells of a 96-well plate from one representative of two independent experiments (c, d).
Supplementary information
Supplementary information
This file contains all uncropped blot information and is referenced by each figure legend of the cropped blots
Supplementary Methods
This file contains the material and methods section including information about the synthesis of a coumarin-quinone conjugate ((2,4,5-trimethyl-3,6-dioxocyclohexa-1,4-dien-1-yl)methyl 7-(diethylamino)-2-oxo-2H-chromene-3-carboxylate) and C9-Hyponitrite ((E)-1,2-bis((2-methyldecan-2-yl)oxy)diazene)
Supplementary Video | FSP1 protects cells from RSL3 induced ferroptosis
Live cell microscopy showing an overlay of Hoechst (blue; Ex 385 nm @ 5%; ET-DAPI filter), propidium iodide (red; Ex 550 @10% nm; ET-CY3 filter) and phase contrast images taken in a 7 min interval. At the start of the experiment, Pfa1 cells overexpressing Mock (left) or FSP1 (right) were treated with 250 nM RSL3 to trigger ferroptosis. Nuclei of living cells are stained blue (Hoechst), while nuclei of dead cells appear red, because propidium iodide can enter their perforated membranes
Source data
Rights and permissions
About this article
Cite this article
Doll, S., Freitas, F.P., Shah, R. et al. FSP1 is a glutathione-independent ferroptosis suppressor. Nature 575, 693–698 (2019). https://doi.org/10.1038/s41586-019-1707-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1707-0
This article is cited by
-
Role of ferroptosis in chronic kidney disease
Cell Communication and Signaling (2024)
-
Role of ferroptosis and ferroptosis-related long non'coding RNA in breast cancer
Cellular & Molecular Biology Letters (2024)
-
Harnessing ferroptosis for enhanced sarcoma treatment: mechanisms, progress and prospects
Experimental Hematology & Oncology (2024)
-
Ferroptosis in early brain injury after subarachnoid hemorrhage: review of literature
Chinese Neurosurgical Journal (2024)
-
Synergistic suppression of ovarian cancer by combining NRF2 and GPX4 inhibitors: in vitro and in vivo evidence
Journal of Ovarian Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.