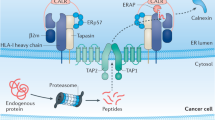
Abstract
The ability of the immune system to eliminate and shape the immunogenicity of tumours defines the process of cancer immunoediting1. Immunotherapies such as those that target immune checkpoint molecules can be used to augment immune-mediated elimination of tumours and have resulted in durable responses in patients with cancer that did not respond to previous treatments. However, only a subset of patients benefit from immunotherapy and more knowledge about what is required for successful treatment is needed2,3,4. Although the role of tumour neoantigen-specific CD8+ T cells in tumour rejection is well established5,6,7,8,9, the roles of other subsets of T cells have received less attention. Here we show that spontaneous and immunotherapy-induced anti-tumour responses require the activity of both tumour-antigen-specific CD8+ and CD4+ T cells, even in tumours that do not express major histocompatibility complex (MHC) class II molecules. In addition, the expression of MHC class II-restricted antigens by tumour cells is required at the site of successful rejection, indicating that activation of CD4+ T cells must also occur in the tumour microenvironment. These findings suggest that MHC class II-restricted neoantigens have a key function in the anti-tumour response that is nonoverlapping with that of MHC class I-restricted neoantigens and therefore needs to be considered when identifying patients who will most benefit from immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
Code for the hmMHC algorithm used to predict presentation of neoantigens by I-Ab can be accessed at https://github.com/artyomovlab/hmmhc.
References
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Larkin, J. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Motzer, R. J. et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N. Engl. J. Med. 373, 1803–1813 (2015).
Borghaei, H. et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N. Engl. J. Med. 373, 1627–1639 (2015).
Lennerz, V. et al. The response of autologous T cells to a human melanoma is dominated by mutated neoantigens. Proc. Natl Acad. Sci. USA 102, 16013–16018 (2005).
Matsushita, H. et al. Cancer exome analysis reveals a T-cell-dependent mechanism of cancer immunoediting. Nature 482, 400–404 (2012).
DuPage, M., Mazumdar, C., Schmidt, L. M., Cheung, A. F. & Jacks, T. Expression of tumour-specific antigens underlies cancer immunoediting. Nature 482, 405–409 (2012).
Robbins, P. F. et al. Mining exomic sequencing data to identify mutated antigens recognized by adoptively transferred tumor-reactive T cells. Nat. Med. 19, 747–752 (2013).
Gubin, M. M. et al. Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature 515, 577–581 (2014).
Wölfel, T. et al. A p16INK4a-insensitive CDK4 mutant targeted by cytolytic T lymphocytes in a human melanoma. Science 269, 1281–1284 (1995).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Strønen, E. et al. Targeting of cancer neoantigens with donor-derived T cell receptor repertoires. Science 352, 1337–1341 (2016).
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Spranger, S. et al. Density of immunogenic antigens does not explain the presence or absence of the T-cell-inflamed tumor microenvironment in melanoma. Proc. Natl Acad. Sci. USA 113, E7759–E7768 (2016).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Hellmann, M. D. et al. Genomic features of response to combination immunotherapy in patients with advanced non-small-cell lung cancer. Cancer Cell 33, 843–852 (2018).
Kreiter, S. et al. Mutant MHC class II epitopes drive therapeutic immune responses to cancer. Nature 520, 692–696 (2015).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Ossendorp, F., Mengedé, E., Camps, M., Filius, R. & Melief, C. J. Specific T helper cell requirement for optimal induction of cytotoxic T lymphocytes against major histocompatibility complex class II negative tumors. J. Exp. Med. 187, 693–702 (1998).
Corthay, A. et al. Primary antitumor immune response mediated by CD4+ T cells. Immunity 22, 371–383 (2005).
Wong, S. B. J., Bos, R. & Sherman, L. A. Tumor-specific CD4+ T cells render the tumor environment permissive for infiltration by low-avidity CD8+ T cells. J. Immunol. 180, 3122–3131 (2008).
Bos, R. & Sherman, L. A. CD4+ T-cell help in the tumor milieu is required for recruitment and cytolytic function of CD8+ T lymphocytes. Cancer Res. 70, 8368–8377 (2010).
Zhu, Z. et al. CD4+ T cell help selectively enhances high-avidity tumor antigen-specific CD8+ T cells. J. Immunol. 195, 3482–3489 (2015).
Borst, J., Ahrends, T., Bąbała, N., Melief, C. J. M. & Kastenmüller, W. CD4+ T cell help in cancer immunology and immunotherapy. Nat. Rev. Immunol. 18, 635–647 (2018).
Andreatta, M. et al. An automated benchmarking platform for MHC class II binding prediction methods. Bioinformatics 34, 1522–1528 (2018).
Mittal, P. et al. Tumor-unrelated CD4 T cell help augments CD134 plus CD137 dual costimulation tumor therapy. J. Immunol. 195, 5816–5826 (2015).
Suri, A., Walters, J. J., Rohrs, H. W., Gross, M. L. & Unanue, E. R. First signature of islet β-cell-derived naturally processed peptides selected by diabetogenic class II MHC molecules. J. Immunol. 180, 3849–3856 (2008).
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+ T cells in a patient with epithelial cancer. Science 344, 641–645 (2014).
Linnemann, C. et al. High-throughput epitope discovery reveals frequent recognition of neo-antigens by CD4+ T cells in human melanoma. Nat. Med. 21, 81–85 (2015).
Old, L. J. & Boyse, E. A. Immunology of experimental tumors. Annu. Rev. Med. 15, 167–186 (1964).
Wei, S. C. et al. Distinct cellular mechanisms underlie anti-CTLA-4 and anti-PD-1 checkpoint blockade. Cell 170, 1120–1133 (2017).
Gubin, M. M. et al. High-dimensional analysis delineates myeloid and lymphoid compartment remodeling during successful immune-checkpoint cancer therapy. Cell 175, 1014–1030 (2018).
Marzo, A. L. et al. Tumor-specific CD4+ T cells have a major “post-licensing” role in CTL mediated anti-tumor immunity. J. Immunol. 165, 6047–6055 (2000).
Bennett, S. R. M., Carbone, F. R., Karamalis, F., Miller, J. F. A. P. & Heath, W. R. Induction of a CD8+ cytotoxic T lymphocyte response by cross-priming requires cognate CD4+ T cell help. J. Exp. Med. 186, 65–70 (1997).
Corthay, A., Lundin, K. U., Lorvik, K. B., Hofgaard, P. O. & Bogen, B. Secretion of tumor-specific antigen by myeloma cells is required for cancer immunosurveillance by CD4+ T cells. Cancer Res. 69, 5901–5907 (2009).
Abelin, J. G. et al. Defining HLA-II ligand processing and binding rules with mass spectrometry enhances cancer epitope prediction. Immunity https://doi.org/10.1016/j.immuni.2019.08.012 (2019).
Mamitsuka, H. Predicting peptides that bind to MHC molecules using supervised learning of hidden Markov models. Proteins 33, 460–474 (1998).
Welch, L. R. Hidden Markov models and the Baum–Welch algorithm. IEEE Inf. Theory Soc. Newsl. 53, 10–13 (2003).
Jurtz, V. et al. NetMHCpan-4.0: improved peptide–MHC class I interaction predictions integrating eluted ligand and peptide binding affinity data. J. Immunol. 199, 3360–3368 (2017).
Jensen, K. K. et al. Improved methods for predicting peptide binding affinity to MHC class II molecules. Immunology 154, 394–406 (2018).
Kowalewski, D. J. & Stevanović, S. Biochemical large-scale identification of MHC class I ligands. Methods Mol. Biol. 960, 145–157 (2013).
Tungatt, K. et al. Antibody stabilization of peptide-MHC multimers reveals functional T cells bearing extremely low-affinity TCRs. J. Immunol. 194, 463–474 (2015).
Acknowledgements
We thank all members of the Schreiber laboratory for discussions and technical support. This work was supported by grants to R.D.S. from the National Cancer Institute of the National Institutes of Health (RO1CA190700), the Parker Institute for Cancer Immunotherapy, the Cancer Research Institute, Janssen Pharmaceutical Company of Johnson and Johnson and the Prostate Cancer Foundation, and by a Stand Up to Cancer-Lustgarten Foundation Pancreatic Cancer Foundation Convergence Dream Team Translational Research Grant. Stand Up to Cancer is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research. E.A. and D.M.L were supported by a postdoctoral training grant (T32 CA00954729) from the National Cancer Institute. D.M.L. and M.M.G. were supported by the Irvington Postdoctoral Fellowship from the Cancer Research Institute. M.D. is a St Baldrick’s Scholar with support from Hope with Hazel and a Pew-Stewart Scholar for Cancer Research supported by the Pew Charitable Trusts. J.P.W. is supported by the National Cancer Institute of the National Institutes of Health Paul Calabresi Career Development Award in Clinical Oncology (K12CA167540). M.M.G. is supported by a Parker Bridge Scholar Award from the Parker Institute for Cancer Immunotherapy. K.W.W. receives support from the National Institutes of Health (R01CA238039). T.J. receives support from a National Institutes of Health Cancer Center Support Grant (P30CA14051) and the Howard Hughes Medical Institute. E.R.U. receives support from the National Institute of Diabetes and Digestive and Kidney Diseases of the National Institutes of Health (AI114551 and DK058177). Aspects of the studies, including ELISPOT, were performed by D. Bender at the Immunomonitoring Laboratory (IML), which is supported by the Andrew M. and Jane N. Bursky Center for Human Immunology and Immunotherapy Programs and the Alvin J. Siteman Comprehensive Cancer Center which, in turn, is supported by a National Cancer Institute of the National Institutes of Health Cancer Center Support Grant (P30CA91842).
Author information
Authors and Affiliations
Contributions
E.A. conceived and designed the experiments, collected the data, performed and interpreted the analyses, and wrote the manuscript. D.M.L. and A.P.M. planned experiments, and collected and analysed data. I.K. conceived of and designed the hmMHC algorithm and performed analyses using it, and wrote the methodological description. M.D. generated the KP9025 sarcoma cell line. A.M.L provided technical assistance and helped to plan experiments using MHC-II tetramers. W.M. and C.F.L. planned, performed and analysed mass spectrometry experiments. E.E. assisted with bioinformatics analyses. A.N.V. assisted with the generation of the CD4+ T cell hybridomas, and helped to design and perform experiments using them. D.R. designed experiments involving multi-colour flow cytometry and collected and analysed the data. J.P.W. provided technical support for MHC-I tetramer staining. M.M.G. assisted in experiment planning. R.F.V.M. collected and analysed data for experiments involving multi-colour flow cytometry. C.D.A., K.C.F.S. and J.M.W. provided technical assistance throughout the study. A.C. collected data. K.W.W. provided mITGB1–MHC-II monomers and provided assistance in experimental design. T.J. provided support in experimental design and data analysis regarding the KP9025 sarcoma line. M.N.A. conceived and designed the hmMHC algorithm and provided bioinformatics support. E.R.U. provided assistance with experimental design. R.D.S. conceived experiments, interpreted data, and wrote the manuscript. All authors contributed to manuscript revision.
Corresponding author
Ethics declarations
Competing interests
R.D.S. is a cofounder, scientific advisory board member, stockholder, and royalty recipient of Jounce Therapeutics and Neon Therapeutics and is a scientific advisory board member for A2 Biotherapeutics, BioLegend, Codiak Biosciences, Constellation Pharmaceuticals, NGM Biopharmaceuticals and Sensei Biotherapeutics. K.W.W. serves on the scientific advisory board of Tscan Therapeutics and Nextechinvest, and receives sponsored research funding from Bristol-Myers Squibb and Novartis; these activities are not related to the findings described in this publication. T.J. is a member of the Board of Directors of Amgen and Thermo Fisher Scientific. He is also a co-founder of Dragonfly Therapeutics and T2 Biosystems, and serves on the Scientific Advisory Board of Dragonfly Therapeutics, SQZ Biotech, and Skyhawk Therapeutics. None of these affiliations represent a conflict of interest with respect to the design or execution of this study or interpretation of data presented in this manuscript. The laboratory of T.J. currently also receives funding from the Johnson & Johnson Lung Cancer Initiative and Calico, but this funding did not support the research described in this manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Lelia Delamarre, Cornelis Melief and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 The hmMHC predictive algorithm and IEDB′18 H2-I-Ab training dataset composition.
a, An example of a fully connected HMM with three hidden states, and emissions corresponding to amino acids. b–d, Composition of IEDB dataset (MHC full ligand export downloaded on 25 November 2018) represented as number of peptides per binding category and measurement type (b, c) and binding category and peptide length (d). Strong binders: IC50 ≤ 50 nM; binders: 50 nM < IC50 ≤ 500 nM; weak binders: 500 nM < IC50 ≤ 5,000 nM; non-binders: all remaining peptides.
Extended Data Fig. 2 Performance of hmMHC compared to netMHCII-2.3 and netMHCIIpan-3.2.
a, hmMHC (orange shapes) underwent 10× cross-validation. In each of the ten cross-validation partitions, on average there were 4,412 binders in the training set, and 771 binders and 77,086 random natural peptides in the validation set. Performance was compared in terms of AUROC to the performance of netMHCII-2.3 (blue triangles) and netMHCIIpan-3.2 (purple triangles) applied on the same validation sets. For hmMHC, performance for different numbers of hidden states is shown. For netMHCII-2.3 and netMHCIIpan-3.2, performance is shown for both predicted affinity and percentile rank (PR). b, Receiver operating characteristic (ROC) curves showing the performance of hmMHC on the H2-I-Ab dataset compared to existing predictors. ROC curves of all peptides and per specific peptide length for every cross-validation partition are shown. c, Illustration of percentile rank for strong binder classification calibrated on random natural peptides. Red lines indicate the percentile ranks of peptides screened for CD4+ T cell reactivity.
Extended Data Fig. 3 mITGB1 is a major MHC-II-restricted neoantigen in T3 sarcomas.
a, b, T3 MHC-II neoantigen predictions for all expressed mutations were made using hmMHC (a) and netMHCII-2.3 (b) (netMHCIIpan-3.2 predictions yielded very similar results, data not shown). The predictions are shown as −log10odds predictor value or logIC50 (smaller values indicate higher likelihood of being presented by I-Ab) and expression level (FPKM). Strong binders are defined as mutations residing in the second percentile of I-Ab binding predictions for random natural peptides for each algorithm (−log10odds ≤ 26.21 or logIC50 ≤ 343.8 nM). The N710Y mutation in ITGB1 met the strong binder threshold in the hmMHC predictions but not in the netMHCII-2.3 predictions. Red dots indicate all mutations that were screened for CD4+ T cell reactivity. Green line denotes high-expression cut-off (FPKM = 89.1). Blue line indicates strong binder cut off for each algorithm. c, Two million T3 sarcoma cells were injected subcutaneously into syngeneic mice and CD4+ TILs were isolated on day 12. IFNγ ELISPOT was performed using naive splenocytes pulsed with 2 μg ml−1 of the indicated peptides. Data are shown as mean of three independent experiments ± s.e.m. d, Gating strategy for pI-Ab tetramer staining of whole TILs. e, Quantification of mITGB1–tetramer and CLIP–tetramer staining of CD4+ T cells from whole T3 TILs 12 days after transplantation. Data are shown as mean ± s.e.m. per cent tetramer-positive cells of CD4+ cells from three independent experiments. f, Syngeneic 129S6 mice were injected subcutaneously with 2 × 106 T3 sarcoma cells and TIL-derived CD4+ T cells were collected 12 days after transplantation. CD4+ T cells were stimulated with naive splenocytes pulsed with 2 μg ml−1 OVA323–339 control or mITGB1697–724 peptide for a flow-based multi-cytokine array. Representative data from one of two independent experiments using pools of five tumours each are shown as average of technical triplicate wells from three pooled tumours.
Extended Data Fig. 4 T3 TIL-derived CD4+ T cell hybridomas are reactive against mITGB1.
CTLL assay of T3 TIL-derived CD4+ T cell hybridoma lines stimulated with naive splenocytes pulsed with 2 μg ml−1 of the individual indicated peptides. Representative data from one of three independent experiments are shown as average cpm from technical duplicate wells.
Extended Data Fig. 5 The mITGB1 epitope is presented on I-Ab.
a, T3 CD4+ T cell hybridomas were stimulated with 2 μg ml−1 mITGB1(710Y) or wild-type ITGB1(710N) peptide-pulsed splenocytes. Activation was measured by CTLL assay. Representative data from three independent hybridoma lines are shown as average of technical replicate wells. b, Mapping of the mITGB1 MHC-II binding core was performed using the CD4+ T cell hybridoma line 41 stimulated with naive splenocytes pulsed with 2 μg ml−1 of overlapping peptides covering mITGB1697–724. Red denotes the T3-specific mutant amino acid at position p1 of the minimal epitope; underlining denotes the validated binding core. Green amino acids represent random residue substitutions used to specifically define valines at residues 715 and 718 as the p6 and p9 MHC-II binding positions and the complete MHC-II binding core. Representative data from two independent experiments are shown as the average of technical triplicate wells. c, MHC-II I-Ab staining of parental T3 cells, IFNγ-stimulated T3 cells and T3 cells transduced with a vector encoding CIITA (T3.CIITA). Representative data from one of three independent experiments are shown. d, Mirror plot showing match between MS/MS spectra of the 14-mer peptide sequence encompassing the N710Y site of mITGB1 eluted from T3.CIITA cells (positive axis) and a corresponding synthetic peptide (negative axis). Labelled m/z values reflect those experimentally observed for the endogenous peptide, with peaks representing b ions highlighted in blue and y ions in red.
Extended Data Fig. 6 mITGB1 CD4+ T cells are required for tumour rejection in response to ICT.
a, Comparison of total number of expressed missense mutations between ten different MCA-induced sarcomas and KP9025 cells. Mutations were defined by whole-exome sequencing and RNA sequencing, and mutational load is shown on a per cell basis. b, Comparison of predicted neoantigen MHC-I affinity values between KP9025 and MCA-induced sarcoma F244 for H-2Db (top) and H-2Kb (bottom). KP9025 cells were not predicted to express any MHC-I neoantigens. c, Rag2−/− mice were subcutaneously injected with 1 × 106 KP.mLAMA4, KP.mITGB1, KP.mLAMA4.mITGB1 or KP.mSB2.SIINFEKL cells. Representative data from one of two independent experiments are presented as tumour diameters from individual mice (n = 5 mice per group for KP.mLAMA4, KP.mITGB1 and KP.mLAMA4.mITGB1 and n = 3 mice for KP.mSB2.SIINFEKL group per experiment). d, Wild-type syngeneic 129S4 mice were injected subcutaneously with 1 × 106 KP.mLAMA4, KP.mITGB1 or KP.mLAMA4.mITGB1 cells and treated with anti-PD-1 (top) or anti-CTLA single agent ICT (bottom) on days 3, 6, and 9 after transplantation. Representative data from one of three independent experiments are shown as tumour diameters from individual mice (n = 5 in all groups per experiment). e, Survival curves for experiments in d and Fig. 2e (n = 15 in all groups).
Extended Data Fig. 7 Outgrowth of nonimmunogenic sarcoma cells expressing MHC-I neoantigens is not a result of cancer immunoediting.
a, Rag2−/− or wild-type 129S4 mice were injected with 1 × 106 KP9025 or KP.mLAMA4 cells and treated with anti-PD-1, anti-CTLA or anti-PD-1 + anti-CTLA4 on days 3, 6 and 9 after injection. Tumours were removed once they reached a maximum diameter of 20 mm in any direction and sarcoma cell lines were established ex vivo. Cell lines were stimulated with IFNγ to upregulate MHC-I and subsequently used to stimulate the mLAMA4-specific CD8+ 74.14 T cell clone. Secretion of IFNγ by T cells was measured using enzyme-linked immunosorbent assay (ELISA). Representative data from two independent experiments are represented as the average of two independent tumour samples in each group. b, Wild-type 129S4 mice were injected with 1 × 106 KP.mSB2.SIINFEKL cells and treated with anti-PD-1 + anti-CTLA4 combination ICT on days 3, 6 and 9 after injection. Tumours were removed as in a. Established ex vivo cell lines were cloned by limiting dilution and parental KP.mSB2.SIINFEKL cells or individual clones from outgrown tumours were used to stimulate the mSB2-specific C3 CD8+ T cell clone; production of IFNγ was quantified using ELISA. Representative data from four independent experiments are presented as average IFNγ concentration of eight individual clones ± s.e.m. Significance was determine using an unpaired, two-tailed t-test. c, Cell surface staining of SIINFEKL-H-2-Kb expressed by unstimulated or IFNγ-stimulated parental KP.mSB2.SIINFEKL cells or individual clones described in b. A representative histogram is shown. d, Quantification of mean ± s.e.m. SIINFEKL-H-2-Kb mean fluorescence intensity (MFI) from eight individual clones in c. NS, not significant. e, Survival curves of wild-type 129S4 mice injected subcutaneously with 1 × 106 KP.mSB2.SIINFEKL.mITGB1 cells. Mice were treated with control monoclonal antibodies or anti-PD-1 + anti-CTLA4 combination ICT on days 3, 6 and 9 after injection. n = 10 mice per group from two independent experiments. ****P = 1.5 × 10−5, Mantel–Cox test.
Extended Data Fig. 8 mITGB1-specific CD4+ T cells display an activated TH1 phenotype.
a, Whole TILs from KP.mLAMA4.mITGB1 tumours 12 days after transplantation were stained with mITGB1-I-Ab tetramers. Populations were previously gated on viable CD11b−CD4+ cells. Representative data from one of two independent experiments with five pooled tumours each are shown. b, mITGB1-I-Ab tetramer-negative and tetramer-positive cells described in a were analysed for expression of T-BET and FOXP3. Representative plots are shown. c, Quantification of two independent experiments in b as average per cent of tetramer-negative and tetramer-positive cells staining positive for the indicated protein. Tumour-bearing mice were treated with control monoclonal antibodies or anti-CTLA4 on days 3, 6, and 9 after transplantation where indicated. d, mITGB1-I-Ab tetramer-positive and tetramer-negative cells in a were analysed for expression of PD-1. Representative plots are shown. e, Quantification of two independent experiments described in d shown as average per cent of tetramer-negative and tetramer-positive cells staining positive for PD-1. f, mITGB1-I-Ab tetramer-positive cells in a were analysed for expression of the indicated proteins. Representative histograms from one of two independent experiments using pools of five tumours each are shown.
Extended Data Fig. 9 CD4+ T cell help is required at the tumour site during primary and memory responses.
a, Rag2−/− mice were simultaneously injected with 1 × 106 KP.mLAMA4 and KP.mLAMA4.mITGB1 cells on opposite flanks. Representative data from one of two independent experiments are shown as individual tumour diameter (n = 3 in each experiment). b, Wild-type 129S4 mice were injected with 1 × 106 KP.mITGB1 cells and were treated with anti-PD-1 + anti-CTLA4 combination ICT on days 3, 6, and 9 after injection. Representative data from one of two individual experiments are shown as individual tumour diameters (n = 5 in all experiments). c, Wild-type 129S4 mice were simultaneously injected with 1 × 106 KP.mLAMA4 and KP.mLAMA4.mITGB1 cells on opposite flanks and treated as in b. Representative data from one of two individual experiments are shown as individual tumour diameters (n = 5 in all experiments). d, Wild-type 129S6 mice were injected subcutaneously with 2 × 106 T3 sarcoma cells and were treated with anti-PD-1 + anti-CTLA4 combination ICT on days 3, 6, and 9 after injection. Following tumour rejection and a 30-day recovery period, tumour-experienced mice were rechallenged with 2 × 106 T3 cells in the presence of control monoclonal antibody or CD4-depleting antibody, or with irrelevant sarcoma cells. Representative data from one of two independent experiments are shown as average tumour diameter ± s.e.m. (n = 5 in all groups per experiment). e, Wild-type 129S4 mice were injected subcutaneously with 1 × 106 KP.mLAMA4.mITGB1 cells followed by surgical resection 10 days after transplantation. After a 30-day recovery period, tumour-experienced mice were rechallenged with 1 × 106 KP9025, KP.mLAMA4.mITGB1, or KP.mLAMA4 cells. Representative data from one of two independent experiments are shown as average tumour diameter ± s.e.m. (n = 5 in all groups per experiment). ****P = 2 × 10−6 by two-way ANOVA with multiple comparisons and Bonferroni correction. f, Quantification of data from three independent experiments in Fig. 5c is shown as average number of spots ± s.e.m. (left) and average number of mITGB1-specific CD4+ cells ± s.e.m. (right). **P = 0.003, ****P = 7.2 × 10−5 (unpaired, two-tailed t-test). g, CD45+Ly6G−MHCII+CD64+CD25−CD11b+F4/80+ macrophages in TILs from mice bearing the indicated contralateral tumours were analysed for expression of iNOS 11 days after tumour transplant. Representative data from four independent experiments are shown. h, Quantification of iNOS+ macrophages from experiments in f as a per cent of total CD45+ cells. Data are shown as average ± s.e.m. of four independent experiments. *P = 0.03 by unpaired, two-tailed t-test. i, CD45+Ly6G−MHCII+CD64+CD25−CD11b+F4/80+ macrophages from the indicated contralateral tumours described were isolated 11 days after transplantation and analysed for expression of iNOS. Representative plots from two independent experiments are shown. j, Quantification of iNOS+ macrophages from two independent experiments in h is shown as average per cent of total CD45+ cells.
Extended Data Fig. 10 Gating strategies for multi-colour flow cytometry.
Gating strategies for multi-colour flow cytometry analysis of tumour-infiltrating macrophage (a) and T cell (b) populations.
Supplementary information
41586_2019_1671_MOESM1_ESM.pdf
Supplementary Table 1 Supplementary Table 1: Characteristics of the screened predicted MHC class II neoantigens. Sequences of mutant peptides identified in T3 sarcomas used in the screening experiments
41586_2019_1671_MOESM3_ESM.pdf
Supplementary Table 2 Supplementary Table 2: Expressed mutations in the KP9025 sarcoma cell line. The four non-synonymous point mutations expressed in the KP9025 sarcoma cell line are shown as the corresponding amino acid substitution
41586_2019_1671_MOESM4_ESM.xlsx
Supplementary Data 1 Supplementary Data 1: T3 nucleotide variant calls. This file contains all the single nucleotide variants determined to generate missense mutations for the methylcholanthrene (MCA)-induced T3 sarcoma cell line as detected by cDNA capture sequencing
41586_2019_1671_MOESM5_ESM.xls
Supplementary Data 2 Supplementary Data 2: KP9025 nucleotide variant calls. This file contains all the single nucleotide variants determined to generate missense mutations for the oncogene driven KP9025 sarcoma cell line as detected by cDNA capture sequencing
Source data
Rights and permissions
About this article
Cite this article
Alspach, E., Lussier, D.M., Miceli, A.P. et al. MHC-II neoantigens shape tumour immunity and response to immunotherapy. Nature 574, 696–701 (2019). https://doi.org/10.1038/s41586-019-1671-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1671-8
This article is cited by
-
Regulatory T cell-mediated immunosuppression orchestrated by cancer: towards an immuno-genomic paradigm for precision medicine
Nature Reviews Clinical Oncology (2024)
-
Discovery of tumor-reactive T cell receptors by massively parallel library synthesis and screening
Nature Biotechnology (2024)
-
Class II HLA-DRB4 is a predictive biomarker for survival following immunotherapy in metastatic non-small cell lung cancer
Scientific Reports (2024)
-
The effect of human leukocyte antigen genotype on survival in advanced prostate cancer treated with primary androgen deprivation therapy: the KYUCOG-1401-A study
Prostate Cancer and Prostatic Diseases (2024)
-
Soluble lymphocyte activation gene-3 (sLAG3) and CD4/CD8 ratio dynamics as predictive biomarkers in patients undergoing immune checkpoint blockade for solid malignancies
British Journal of Cancer (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.