Abstract
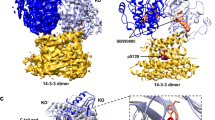
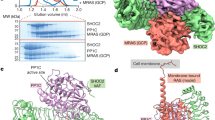
RAF family kinases are RAS-activated switches that initiate signalling through the MAP kinase cascade to control cellular proliferation, differentiation and survival1,2,3. RAF activity is tightly regulated and inappropriate activation is a frequent cause of cancer4,5,6; however, the structural basis for RAF regulation is poorly understood at present. Here we use cryo-electron microscopy to determine autoinhibited and active-state structures of full-length BRAF in complexes with MEK1 and a 14-3-3 dimer. The reconstruction reveals an inactive BRAF–MEK1 complex restrained in a cradle formed by the 14-3-3 dimer, which binds the phosphorylated S365 and S729 sites that flank the BRAF kinase domain. The BRAF cysteine-rich domain occupies a central position that stabilizes this assembly, but the adjacent RAS-binding domain is poorly ordered and peripheral. The 14-3-3 cradle maintains autoinhibition by sequestering the membrane-binding cysteine-rich domain and blocking dimerization of the BRAF kinase domain. In the active state, these inhibitory interactions are released and a single 14-3-3 dimer rearranges to bridge the C-terminal pS729 binding sites of two BRAFs, which drives the formation of an active, back-to-back BRAF dimer. Our structural snapshots provide a foundation for understanding normal RAF regulation and its mutational disruption in cancer and developmental syndromes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Three-dimensional cryo-EM density maps have been deposited in the Electron Microscopy Data Bank (EMDB) with accession codes EMD-0541, EMD-20550, EMD-20552 and EMD-20551. Atomic coordinates corresponding to these cryo-EM reconstructions have been deposited in the Protein Data Bank (PDB) with accession codes 6NYB, 6Q0J, 6Q0T and 6Q0K. Structure factors and atomic coordinates for the BRAF kinase domain–MEK1 crystal structure have been deposited in the PDB with accession code 6PP9.
References
Lavoie, H. & Therrien, M. Regulation of RAF protein kinases in ERK signalling. Nat. Rev. Mol. Cell Biol. 16, 281–298 (2015).
Simanshu, D. K., Nissley, D. V. & McCormick, F. RAS proteins and their regulators in human disease. Cell 170, 17–33 (2017).
Terrell, E. M. & Morrison, D. K. Ras-mediated activation of the Raf family kinases. Cold Spring Harb. Perspect. Med. 9, a033746 (2019).
Holderfield, M., Deuker, M. M., McCormick, F. & McMahon, M. Targeting RAF kinases for cancer therapy: BRAF-mutated melanoma and beyond. Nat. Rev. Cancer 14, 455–467 (2014).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in The Cancer Genome Atlas. Cell 173, 321–337 (2018).
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature 417, 949–954 (2002).
Gardino, A. K., Smerdon, S. J. & Yaffe, M. B. Structural determinants of 14-3-3 binding specificities and regulation of subcellular localization of 14-3-3-ligand complexes: a comparison of the X-ray crystal structures of all human 14-3-3 isoforms. Semin. Cancer Biol. 16, 173–182 (2006).
Ghosh, S. et al. The cysteine-rich region of raf-1 kinase contains zinc, translocates to liposomes, and is adjacent to a segment that binds GTP-ras. J. Biol. Chem. 269, 10000–10007 (1994).
Hekman, M. et al. Associations of B- and C-Raf with cholesterol, phosphatidylserine, and lipid second messengers: preferential binding of Raf to artificial lipid rafts. J. Biol. Chem. 277, 24090–24102 (2002).
Leonard, T. A. & Hurley, J. H. Regulation of protein kinases by lipids. Curr. Opin. Struct. Biol. 21, 785–791 (2011).
Rajakulendran, T., Sahmi, M., Lefrançois, M., Sicheri, F. & Therrien, M. A dimerization-dependent mechanism drives RAF catalytic activation. Nature 461, 542–545 (2009).
Weber, C. K., Slupsky, J. R., Kalmes, H. A. & Rapp, U. R. Active Ras induces heterodimerization of cRaf and BRaf. Cancer Res. 61, 3595–3598 (2001).
Rushworth, L. K., Hindley, A. D., O’Neill, E. & Kolch, W. Regulation and role of Raf-1/B-Raf heterodimerization. Mol. Cell. Biol. 26, 2262–2272 (2006).
Haling, J. R. et al. Structure of the BRAF–MEK complex reveals a kinase activity independent role for BRAF in MAPK signaling. Cancer Cell 26, 402–413 (2014).
Diedrich, B. et al. Discrete cytosolic macromolecular BRAF complexes exhibit distinct activities and composition. EMBO J. 36, 646–663 (2017).
Nassar, N. et al. The 2.2 Å crystal structure of the Ras-binding domain of the serine/threonine kinase c-Raf1 in complex with Rap1A and a GTP analogue. Nature 375, 554–560 (1995).
Mott, H. R. et al. The solution structure of the Raf-1 cysteine-rich domain: a novel ras and phospholipid binding site. Proc. Natl Acad. Sci. USA 93, 8312–8317 (1996).
Wan, P. T. et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 116, 855–867 (2004).
Improta-Brears, T., Ghosh, S. & Bell, R. M. Mutational analysis of Raf-1 cysteine rich domain: requirement for a cluster of basic aminoacids for interaction with phosphatidylserine. Mol. Cell. Biochem. 198, 171–178 (1999).
Travers, T. et al. Molecular recognition of RAS/RAF complex at the membrane: role of RAF cysteine-rich domain. Sci. Rep. 8, 8461 (2018).
Clark, G. J. et al. 14-3-3 ζ negatively regulates Raf-1 activity by interactions with the Raf-1 cysteine-rich domain. J. Biol. Chem. 272, 20990–20993 (1997).
Molzan, M. et al. Impaired binding of 14-3-3 to C-RAF in Noonan syndrome suggests new approaches in diseases with increased Ras signaling. Mol. Cell. Biol. 30, 4698–4711 (2010).
Thevakumaran, N. et al. Crystal structure of a BRAF kinase domain monomer explains basis for allosteric regulation. Nat. Struct. Mol. Biol. 22, 37–43 (2015).
Rauen, K. A. The RASopathies. Annu. Rev. Genomics Hum. Genet. 14, 355–369 (2013).
Fischmann, T. O. et al. Crystal structures of MEK1 binary and ternary complexes with nucleotides and inhibitors. Biochemistry 48, 2661–2674 (2009).
Michaud, N. R., Fabian, J. R., Mathes, K. D. & Morrison, D. K. 14-3-3 is not essential for Raf-1 function: identification of Raf-1 proteins that are biologically activated in a 14-3-3- and Ras-independent manner. Mol. Cell. Biol. 15, 3390–3397 (1995).
Daub, M. et al. The RafC1 cysteine-rich domain contains multiple distinct regulatory epitopes which control Ras-dependent Raf activation. Mol. Cell. Biol. 18, 6698–6710 (1998).
Sarkozy, A. et al. Germline BRAF mutations in Noonan, LEOPARD, and cardiofaciocutaneous syndromes: molecular diversity and associated phenotypic spectrum. Hum. Mutat. 30, 695–702 (2009).
Zhang, B. H. & Guan, K. L. Activation of B-Raf kinase requires phosphorylation of the conserved residues Thr598 and Ser601. EMBO J. 19, 5429–5439 (2000).
Imielinski, M. et al. Oncogenic and sorafenib-sensitive ARAF mutations in lung adenocarcinoma. J. Clin. Invest. 124, 1582–1586 (2014).
Pfister, S. et al. BRAF gene duplication constitutes a mechanism of MAPK pathway activation in low-grade astrocytomas. J. Clin. Invest. 118, 1739–1749 (2008).
Hatzivassiliou, G. et al. Mechanism of MEK inhibition determines efficacy in mutant KRAS- versus BRAF-driven cancers. Nature 501, 232–236 (2013).
Ishii, N. et al. Enhanced inhibition of ERK signaling by a novel allosteric MEK inhibitor, CH5126766, that suppresses feedback reactivation of RAF activity. Cancer Res. 73, 4050–4060 (2013).
Lito, P. et al. Disruption of CRAF-mediated MEK activation is required for effective MEK inhibition in KRAS mutant tumors. Cancer Cell 25, 697–710 (2014).
Karoulia, Z., Gavathiotis, E. & Poulikakos, P. I. New perspectives for targeting RAF kinase in human cancer. Nat. Rev. Cancer 17, 676–691 (2017).
Hymowitz, S. G. & Malek, S. Targeting the MAPK pathway in RAS mutant cancers. Cold Spring Harb. Perspect. Med. 8, a031492 (2018).
Jin, T. et al. RAF inhibitors promote RAS–RAF interaction by allosterically disrupting RAF autoinhibition. Nat. Commun. 8, 1211 (2017).
Zakeri, B. et al. Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. Proc. Natl Acad. Sci. USA 109, E690–E697 (2012).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun Biol. 2, 218 (2019).
Zivanov, J., Nakane, T., Forsberg, B. O., Kimanius, D., Hagen, W. J., Lindahl, E. & Scheres, S. H. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Potterton, E., Briggs, P., Turkenburg, M. & Dodson, E. A graphical user interface to the CCP4 program suite. Acta Crystallogr. D 59, 1131–1137 (2003).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Alexander, W. M., Ficarro, S. B., Adelmant, G. & Marto, J. A. multiplierz v2.0: A Python-based ecosystem for shared access and analysis of native mass spectrometry data. Proteomics 17, 1700091 (2017).
Parikh, J. R. et al. multiplierz: an extensible API based desktop environment for proteomics data analysis. BMC Bioinformatics 10, 364 (2009).
Ficarro, S. B., Alexander, W. M. & Marto, J. A. Mzstudio: A dynamic digital canvas for user-driven interrogation of mass spectrometry data. Proteomes 5, 20 (2017).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Morin, A., Eisenbraun, B., Key, J., Sanschagrin, P. C., Timony, M. A., Ottaviano, M. & Sliz, P. Collaboration gets the most out of software. eLife 2, e01456 (2013).
Acknowledgements
This work was supported in part by the PLGA fund at the Pediatric Brain Tumor Foundation, by Novartis Institutes for Biomedical Research, and by National Institutes of Health (NIH) grants P50 CA165962 (M.J.E.), PO1 CA154303 (M.J.E.) and R50 CA221830 (E.P.). Cryo-EM imaging for the autoinhibited structure was carried out at the University of Massachusetts Medical School Cryo-EM Core Facility, and we thank C. Xu and K. Song for their guidance in image acquisition. Active-state structures were imaged at the Harvard Cryo-EM Center for Structural Biology. Diffraction data for the BRAF/MEK1 kinase complex crystal structure were recorded at beamline 24-ID-C of the Northeast Collaborative Access Team at the Advanced Photon Source, Argonne National Laboratory, which is supported in part by the Department of Energy and NIH grant GM124165. We thank M. Kostic for critical reading and editing of the manuscript.
Author information
Authors and Affiliations
Contributions
E.P. expressed, purified and biochemically characterized all BRAF–MEK1–14-3-3 and BRAF–14-3-3 complexes. E.P. and H.J. prepared the samples and collected electron microscopy data. S.R., together with H.J. and M.J.E., processed the electron microscopy data to obtain the 3D reconstructions. E.P., S.R., B.-W.K., K.L., H.J. and M.J.E. built and analysed cryo-EM models. K.L. and G.G.-D.P. determined the BRAF–MEK1 kinase domain crystal structure. S.B.F. and J.A.M. were responsible for the mass spectrometry experiments. H.S. carried out preliminary expression and purification studies for the complex. H.J. and M.J.E. directed the project, and M.J.E. drafted the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
This work was supported in part by a sponsored research agreement from Novartis Institutes for Biomedical Research (NIBR) and M.J.E. has been a consultant to NIBR. J.A.M. serves on the Scientific Advisory Board of 908 Devices.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Frank Sicheri and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Biochemical characterization of purified BRAF complexes and cryo-EM analysis of the autoinhibited BRAF–MEK1–14-3-3 complex.
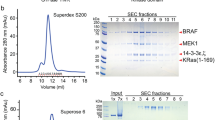
a, The full-length, autoinhibited BRAF–MEK1–14-3-3ε,ζ complex used for cryo-EM structure determination. Left, the elution profile from SEC on a Superdex 200 column; centre, Coomassie-stained SDS–PAGE analysis of elution fractions; right, analysis by size-exclusion chromatography with multi-angle light scattering. A molar mass of 196 kDa was indicated; the calculated molecular weight of the complex is 192 kDa. b, Analysis of co-expression of human 14-3-3 isoforms with BRAF and MEK1 in insect cells. Left, Strep-tagged BRAF, MEK1(S218A/S222A) and the indicated Flag-tagged human 14-3-3 isoforms were co-expressed in Sf9 cells and BRAF–MEK1–14-3-3 complexes were affinity-isolated from clarified lysates with either Strep-TactinXT (left four lanes) or anti-Flag (right four lanes) magnetic beads. Right, parallel gels were blotted with an anti-14-3-3 antibody that recognizes all 14-3-3 isoforms (top blot) or with an anti-Flag antibody (bottom blot). Note that even in the presence of robust overexpression of these human isoforms, BRAF preferentially associated with the endogenous insect-cell 14-3-3 proteins (as seen in the Strep-TactinXT-precipitated lanes of the Coomassie-stained gel). c, Mass-spectrometry-based quantification of selected phosphorylation sites in complexes with wild-type BRAF and with BRAF(S365A) purified for structural analysis. Note that the BRAF(S365A) complex was prepared with MEK1(S218A/S222A), whereas the wild-type BRAF complex used in this analysis contained wild-type MEK1. d, Portion of a representative micrograph used for reconstruction of the autoinhibited BRAF–MEK1–14-3-3 complex. e, Representative 2D class averages for reconstruction of the autoinhibited BRAF–MEK1–14-3-3 complex. Scale bar, 10 nm. f, Fourier shell correlation (FSC) curves for the reconstruction. The horizontal line indicates a correlation of 0.143; the FSC curve for two half-maps (blue) crosses this threshold at a resolution of 4.1Å. A correlation curve for the map versus the atomic model is plotted in red. g, The cryo-EM map of the autoinhibited BRAF–MEK1–14-3-3 complex filtered to 5 Å resolution and contoured at a lower level to reveal weaker density corresponding to the RBD domain. The map surface is coloured by domain as in Fig. 1. Unassigned densities (grey) can be ascribed to the RBD domain and other poorly structured elements as indicated. For gel source data, see Supplementary Fig. 1. Experiments in a and b were repeated at least twice with similar results. Imaging experiments in d and e were repeated four times with similar results.
Extended Data Fig. 2 14-3-3 domain sequence alignment and interactions of the CRD domain in the autoinhibited state.
a, Sequence alignment of insect-cell (S. frugiperda) and human 14-3-3 isoforms. Secondary structure is indicated above the alignment. Identically conserved residues are shaded red. Symbols above the alignment indicate contacts with the BRAF CRD domain (purple squares), kinase domain (blue triangles), and pS365 or pS729 segments (black circles). b, Interactions of the CRD domain. Domains that contact the CRD are shown with a transparent surface and the CRD domain is shown as a purple ribbon with grey spheres representing bound zinc atoms. Sidechains are shown for CRD residues that correspond to 7 (of 11 total) residues identified in an alanine scanning mutagenesis study of the CRAF CRD domain27. Alanine mutations in the corresponding residues increased RAS(G12V)-dependent activation of CRAF. Two mutations in this study fully activated CRAF in the absence of RAS(G12V); the corresponding BRAF residues are F247 and D249. F247 makes hydrophobic contacts with both the kinase C-lobe and the 14-3-3 domain, whereas D249 is positioned to form a salt bridge with R691 in the kinase C-lobe. The remaining four residues are also at sites of interdomain contacts but are not illustrated (T241, K253, Q262 and K267).
Extended Data Fig. 3 Interactions of the 14-3-3 dimer with BRAF in the autoinhibited state.
a–d, Cryo-EM density is shown at key sites of interaction that stabilize the autoinhibited complex, and domains are coloured as in Fig. 1. a, A portion of the interface between the CRD and 14-3-3 domain. b, The pS365 segment (CR2) bound in the recognition groove of the 14-3-3 domain. c, Contact between the α8–α9 loop of the 14-3-3 domain and the BRAF kinase domain. d, The C-terminal pS729 segment coordinated in the opposite recognition groove of the 14-3-3 dimer. The map is contoured at the same level in a–d. e, Front and back views of the reconstruction. We observe continuous density connecting the C terminus of the BRAF kinase and the pS729 14-3-3 binding site (inset). f, Comparison of the binding mode of the pS365 segment in the present structure with that in a previously determined crystal structure of an isolated CRAF peptide bound to 14-3-3ζ (PDB ID: 3NKX). The corresponding region of the present structure (the pS365 segment is shown with orange carbon atoms and the 14-3-3 domain is shown in tan) is superimposed on the 3NKX crystal structure (shown in blue and cyan), revealing a close correspondence in conformations of the bound peptides. g, The BRAF CRD is a hot spot for RASopathy mutations, which map to sites of contact between the CRD (purple), kinase (blue) and 14-3-3 domains (tan), and are expected to destabilize the autoinhibited assembly. Sites of RASopathy mutations are shown in stick form and are labelled. RASopathy mutations in the BRAF kinase domain (Q709) and CR2 region (red, S365) are also expected to destabilize these inhibitory intramolecular contacts.
Extended Data Fig. 4 Additional views and analysis of the BRAF and MEK kinase domains in the autoinhibited BRAF–MEK1–14-3-3 complex.
a, Cryo-EM density map in the region of the BRAF active site showing bound ATP-γ-S. b, Cryo-EM density map in the region of the MEK1 active site indicating bound ADP, which is probably hydrolysed from ATP-γ-S. Maps in a and b are contoured at the same level. c, Superposition of the BRAF–MEK1 component of the present autoinhibited cryo-EM structure (green and dark blue) with the previously reported crystal structure of a BRAF and MEK1 kinase domain complex (yellow and light blue; PDB ID: 4MNE). The superposition is based on the MEK component of the structures, and it reveals a relative rotation of BRAF of approximately 5° about the C-lobe contact. d, Superposition of the BRAF kinase domain from the present structure with that of previously isolated BRAF–MEK kinase complex (PDB ID: 4MNE). Note that the present structure (dark blue, with C-helix coloured purple and the activation segment orange) exhibits key features of an autoinhibited state (C-helix out, with an inhibitory turn in the activation segment), whereas the previous structure (light blue) adopts an overall active conformation. e, Detailed view of a portion of the C-lobe contact between BRAF (blue) and MEK1 (green). f, Portions of the BRAF (blue) and MEK1 (green) activation segments interact in an anti-parallel orientation. Activating phosphorylation sites in the MEK1 activation loop are substituted with alanine in this structure (S218A/S222A), but neither residue is positioned appropriately for phosphorylation by BRAF. Note that our discussion of these interactions relies in part on the crystal structures referenced to build the atomic model, as the cryo-EM map in this region does not unambiguously define all sidechain conformations. g–j, Comparison of BRAF kinase domain conformations and relative N- and C-lobe orientations. g, Sulfonamide-containing BRAF inhibitors perturb the inactive conformation of BRAF. The BRAF kinase domain in the present structure (blue ribbon, with C-helix coloured red and the activation segment orange) is superimposed on the structure of the BRAF kinase domain crystallized as a monomer with PLX4720 (grey, PDB ID: 4WO5). The superposition is based on the C-lobes of both kinases, revealing an altered orientation of the N-lobe in the inhibitor-bound structure (a rotation of around 15°). Note also that the inhibitory turn in the activation segment helix is replaced by a short helix in the PLX4720 complex. h, Alternative view of the superposition shown in g, highlighting the axis of rotation (pink arrow) between the N-lobes. i, As in h, but with a representative inhibitor-bound dimeric BRAF structure superimposed (PDB ID: 5CSW, a dabrafenib complex). The rotation axes for N-lobe rotations of dimer structures are shown as green arrows. Note that the orientation of the rotation axis is similar for all of the dimer structures, but almost orthogonal to that of the monomer structure in h. In both h and i, the Cα atoms of K522 are shown as spheres as a point of reference. j, Relative N-lobe rotation of wild-type and BRAF(V600E) crystal structures available in the Protein Data Bank (PDB) are compared with the present nucleotide-bound, autoinhibited structure. As illustrated in h and i, C-lobes of the BRAF kinase domains were superimposed, and the rotation required to bring the kinase N-lobes into register were calculated using PyMOL. With the exception of 4MNE, all structures compared were determined in complex with inhibitors.
Extended Data Fig. 5 Additional analysis of the crystal structure of the autoinhibited BRAF–MEK1 kinase domain complex.
a, Crystallographic data collection and refinement statistics for the structure of the BRAF kinase domain (BRAFKD) in complex with MEK1(S218A/S222A), AMP-PNP and MEK inhibitor GDC-0623. Data were recorded from a single crystal. b, The crystal structure of the autoinhibited BRAF–MEK1 kinase domain complex is superimposed on the corresponding region of the autoinhibited cryo-EM structure. c, ATP-analogue AMP-PNP is extensively coordinated in the autoinhibited state. Hydrogen bonds from coordinating residues are indicated by dashed lines. d, MEK1 residue E102 in the β3–αC loop is positioned to form a hydrogen bond with a ribose hydroxyl of the nucleotide bound in the BRAF active site. e, Rare but recurrent oncogenic mutations in MEK1 map to the region of the N-terminal helix. A small, in-frame deletion of two residues (E102, I103) in the β3–αC loop maps to the region of the interface between BRAF and MEK1. f, RASopathy mutations in BRAF illustrated in the inactive conformation of the kinase domain. As with oncogenic mutations in many of the same residues, RASopathy-associated mutations will perturb nucleotide binding and/or the stability of the inhibitory turn. Notably, residues E501 and T599 form a hydrogen bond (dashed line) that appears to contribute to the stability of the inhibitory turn.
Extended Data Fig. 6 RAF sequence alignment.
Human ARAF, BRAF and CRAF sequences are aligned and identically conserved residues are shaded red. Secondary structure elements are indicated above the alignment. Symbols above the alignment indicate residues that, in the autoinhibited structure, lie at the interface with MEK1 (cyan circles), the CRD/14-3-3 interface (violet squares), the CRD/kinase domain interface (blue triangles), and the 14-3-3/kinase domain interface (yellow stars).
Extended Data Fig. 7 Purification and characterization of wild-type and BRAF(S365A) complexes.
a, BRAF(S365A) was co-expressed with MEK1(S218A/S222A) in insect cells, purified by serial Ni-NTA agarose and StrepTrapHP affinity chromatography, and subjected to SEC on Superose 6 column. The SEC elution trace is shown on the left with a Coomassie-stained SDS–PAGE gel of elution fractions on the right. A parallel gel was blotted with an antibody against pS729 (bottom right). BRAF activity in each fraction was measured in a MEK phosphorylation assay (top right; see Methods for assay details.). b, c, Side-by-side comparison of wild-type BRAF complexes isolated from insect cells without (b) and with (c) co-expression of MEK1(S218A/S222A). Complexes were purified by serial Ni-NTA agarose and StrepTrapHP affinity chromatography and subjected to SEC on Superose 6. The SEC elution traces are shown on the left with Coomassie-stained SDS–PAGE gels of elution fractions on the right. BRAF activity in each fraction was measured in a MEK phosphorylation assay as described above (top right). Note that co-expression of MEK1 markedly decreases the void peak and enables the isolation of a late-eluting peak (around 15 ml) with little MEK-phosphorylation activity that corresponds to the autoinhibited BRAF–MEK1–14-3-3 monomer complex (c, fractions B8–C3). d, Wild-type BRAF was expressed in mammalian HEK293 cells, purified by serial Ni-NTA agarose and StrepTrapHP affinity chromatography, and subjected to SEC on Superdex 200. e, Elution fractions from the wild-type BRAF–14-3-3 SEC run in d are analysed by SDS–PAGE and western blotting, revealing that BRAF co-purifies with endogenous human 14-3-3 proteins. Fractions were also blotted for total BRAF (anti-StrepII), pS365 and pS729. f, Mass spectrometry analysis of trypsin and Lys-C protease digests of peak fractions of the BRAF–14-3-3 complex from HEK293 cells revealed multiple peptide sequences that mapped uniquely to six of the seven human 14-3-3 isoforms. The δ and α isoforms are phosphorylation variants of ζ and β, respectively. For gel source data, see Supplementary Fig. 1. SEC experiments were repeated at least three times (a–e), activity assays twice (a) and once (b, c), and blotting twice (e) with similar results.
Extended Data Fig. 8 Cryo-EM imaging of dimeric, active-state BRAF complexes and mass-spectrometry-based measurement of phosphorylation stoichiometry in BRAF–14-3-3 and BRAF–MEK1–14-3-3 complexes.
a, Representative 2D class averages for the BRAF(S365A)–MEK1–14-3-3 complex. Scale bar, 10 nm. b, FSC curves for the BRAF(S365A)–MEK1–14-3-3 reconstructions presented in Fig. 4. c, SEC (Superose 6) traces for the indicated affinity-isolated BRAF complexes, prepared using insect or mammalian cells as described in Extended Data Fig. 7 and Methods. SEC experiments were repeated at least three times with similar results. d–f, Per cent phosphorylation of selected BRAF sites in successive elution fractions is plotted for each sample analysed in c. Fractional phosphorylation of these sites was measured using a mass-spectrometry-based assay (see Methods). d, Wild-type BRAF–14-3-3 complex produced by insect cells. e, Wild-type BRAF–14-3-3 complex produced by mammalian cells. f, BRAF–MEK1–14-3-3 complex produced in insect cells, prepared by co-expression of wild-type BRAF and MEK1(S218A/S222A). In d–f, note the high fractional phosphorylation of S729 in all samples, and the negligible phosphorylation of activation segment sites T599 and S602. g, Representative 2D-class averages for wild-type BRAF–14-3-3 complexes prepared from mammalian cells with (top) and without (bottom) the addition of BRAF inhibitor GDC-0879 (1 μM). Both samples yielded class averages indicative of the same particle architecture, but those of the drug-treated sample revealed better-defined secondary structure. Scale bar, 10 nm. h, FSC curve for the wild-type BRAF–14-3-3 reconstruction. i, Single-particle reconstruction of the wild-type BRAF–14-3-3 complex produced in mammalian cells treated with GDC-0879. The reconstruction reveals a back-to-back BRAF kinase domain dimer with a 14-3-3 dimer bridging between its C-terminal pS729 tails. Comparison of these front and back views reveals the highly asymmetric position of the 14-3-3 dimer with respect to the dimerized kinase domains. Imaging experiments in a and g were repeated twice with independent preparations, and gave similar results.
Extended Data Fig. 9 Structural snapshots outline a model for RAF activation.
The RBD domain is exposed in the context of the autoinhibited BRAF–MEK1–14-3-3 monomer complex, enabling high-affinity binding to farnesylated, GTP-loaded RAS at the plasma membrane. We propose that ‘extraction’ of the CRD domain upon binding to prenylated RAS at the membrane is a key step in RAF activation. Without the stabilizing interactions of the CRD domain, the 14-3-3 domain can release from the BRAF kinase domain and pS365 segment to form an ‘open’ monomer. We expect the RAF–MEK kinase module of the open monomer to maintain its inactive, ATP-bound conformation as observed in the crystal structure described here. Finally, the 14-3-3 domain can rearrange to bind the C-terminal pS729 sites of two open RAF molecules, driving formation of the active, back-to-back RAF dimer. As illustrated here, the stoichiometry of 14-3-3 binding changes upon activation, but we do not exclude the possibility that a second 14-3-3 dimer remains associated with the complex, for example by bridging the pS365 segments. KD, RAF kinase domain; red circles (pSer) represent the pS365 and pS729 14-3-3 binding segments.
Supplementary information
Supplementary Figure 1
Uncropped western-blot scans with molecular weight markers.
Rights and permissions
About this article
Cite this article
Park, E., Rawson, S., Li, K. et al. Architecture of autoinhibited and active BRAF–MEK1–14-3-3 complexes. Nature 575, 545–550 (2019). https://doi.org/10.1038/s41586-019-1660-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1660-y
This article is cited by
-
Roles and mechanisms of aberrant alternative splicing in melanoma — implications for targeted therapy and immunotherapy resistance
Cancer Cell International (2024)
-
Live-cell target engagement of allosteric MEKi on MEK–RAF/KSR–14-3-3 complexes
Nature Chemical Biology (2024)
-
The CNK–HYP scaffolding complex promotes RAF activation by enhancing KSR–MEK interaction
Nature Structural & Molecular Biology (2024)
-
BRAF — a tumour-agnostic drug target with lineage-specific dependencies
Nature Reviews Clinical Oncology (2024)
-
Membrane lipids drive formation of KRAS4b-RAF1 RBDCRD nanoclusters on the membrane
Communications Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.