Abstract
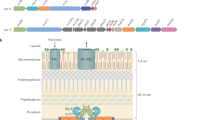
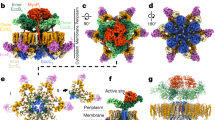
Host infection by pathogenic mycobacteria, such as Mycobacterium tuberculosis, is facilitated by virulence factors that are secreted by type VII secretion systems1. A molecular understanding of the type VII secretion mechanism has been hampered owing to a lack of three-dimensional structures of the fully assembled secretion apparatus. Here we report the cryo-electron microscopy structure of a membrane-embedded core complex of the ESX-3/type VII secretion system from Mycobacterium smegmatis. The core of the ESX-3 secretion machine consists of four protein components—EccB3, EccC3, EccD3 and EccE3, in a 1:1:2:1 stoichiometry—which form two identical protomers. The EccC3 coupling protein comprises a flexible array of four ATPase domains, which are linked to the membrane through a stalk domain. The domain of unknown function (DUF) adjacent to the stalk is identified as an ATPase domain that is essential for secretion. EccB3 is predominantly periplasmatic, but a small segment crosses the membrane and contacts the stalk domain. This suggests that conformational changes in the stalk domain—triggered by substrate binding at the distal end of EccC3 and subsequent ATP hydrolysis in the DUF—could be coupled to substrate secretion to the periplasm. Our results reveal that the architecture of type VII secretion systems differs markedly from that of other known secretion machines2, and provide a structural understanding of these systems that will be useful for the design of antimicrobial strategies that target bacterial virulence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Cryo-EM maps have been deposited in the Electron Microscopy Database under accession codes EMD-10186 (core complex dimer), EMD-10187 (protomer 1), EMD-10189 (conformation 1), EMD-10190 (conformation 2) and EMD-10188 (dimer of EccB3). The model of the ESX-3 core complex, protomer 1, and the fork have been deposited in the Protein Data Bank under PDB accession codes 6SGW, 6SGX, and 6SGY, respectively. The cryo-EM map and model for protomer 2 (discussed but not shown, because it has lower resolution than that of protomer 1) are deposited under accession codes EMD-10191 and 6SGZ. All gel source data are provided in Supplementary Fig. 1, and all other data are available from the corresponding author upon reasonable request.
References
Gröschel, M. I., Sayes, F., Simeone, R., Majlessi, L. & Brosch, R. ESX secretion systems: mycobacterial evolution to counter host immunity. Nat. Rev. Microbiol. 14, 677–691 (2016).
Costa, T. R. et al. Secretion systems in Gram-negative bacteria: structural and mechanistic insights. Nat. Rev. Microbiol. 13, 343–359 (2015).
Maciąg, A., Piazza, A., Riccardi, G. & Milano, A. Transcriptional analysis of ESAT-6 cluster 3 in Mycobacterium smegmatis. BMC Microbiol. 9, 48 (2009).
Rodriguez, G. M., Voskuil, M. I., Gold, B., Schoolnik, G. K. & Smith, I. ideR, an essential gene in Mycobacterium tuberculosis: role of IdeR in iron-dependent gene expression, iron metabolism, and oxidative stress response. Infect. Immun. 70, 3371–3381 (2002).
Siegrist, M. S. et al. Mycobacterial Esx-3 is required for mycobactin-mediated iron acquisition. Proc. Natl Acad. Sci. USA 106, 18792–18797 (2009).
Serafini, A., Boldrin, F., Palù, G. & Manganelli, R. Characterization of a Mycobacterium tuberculosis ESX-3 conditional mutant: essentiality and rescue by iron and zinc. J. Bacteriol. 191, 6340–6344 (2009).
Tufariello, J. M. et al. Separable roles for Mycobacterium tuberculosis ESX-3 effectors in iron acquisition and virulence. Proc. Natl Acad. Sci. USA 113, E348–E357 (2016).
Tinaztepe, E. et al. Role of metal-dependent regulation of ESX-3 secretion in intracellular survival of Mycobacterium tuberculosis. Infect. Immun. 84, 2255–2263 (2016).
Mehra, A. et al. Mycobacterium tuberculosis type VII secreted effector EsxH targets host ESCRT to impair trafficking. PLoS Pathog. 9, e1003734 (2013).
Portal-Celhay, C. et al. Mycobacterium tuberculosis EsxH inhibits ESCRT-dependent CD4+ T-cell activation. Nat. Microbiol. 2, 16232 (2016).
Mittal, E. et al. Mycobacterium tuberculosis type VII secretion system effectors differentially impact the ESCRT endomembrane damage response. MBio 9, e01765-18 (2018).
Ohol, Y. M. et al. Mycobacterium tuberculosis MycP1 protease plays a dual role in regulation of ESX-1 secretion and virulence. Cell Host Microbe 7, 210–220 (2010).
Brodin, P. et al. Dissection of ESAT-6 system 1 of Mycobacterium tuberculosis and impact on immunogenicity and virulence. Infect. Immun. 74, 88–98 (2006).
Siegrist, M. S. et al. Mycobacterial Esx-3 requires multiple components for iron acquisition. MBio 5, e01073-14 (2014).
Houben, E. N. et al. Composition of the type VII secretion system membrane complex. Mol. Microbiol. 86, 472–484 (2012).
Abdallah, A. M. et al. A specific secretion system mediates PPE41 transport in pathogenic mycobacteria. Mol. Microbiol. 62, 667–679 (2006).
van Winden, V. J. et al. Mycosins are required for the stabilization of the ESX-1 and ESX-5 type VII secretion membrane complexes. MBio 7, e01471-16 (2016).
Gray, T. A. et al. Intercellular communication and conjugation are mediated by ESX secretion systems in mycobacteria. Science 354, 347–350 (2016).
Beckham, K. S. et al. Structure of the mycobacterial ESX-5 type VII secretion system membrane complex by single-particle analysis. Nat. Microbiol. 2, 17047 (2017).
Rosenberg, O. S. et al. Substrates control multimerization and activation of the multi-domain ATPase motor of type VII secretion. Cell 161, 501–512 (2015).
Wagner, J. M. et al. Structures of EccB1 and EccD1 from the core complex of the mycobacterial ESX-1 type VII secretion system. BMC Struct. Biol. 16, 5 (2016).
Solomonson, M. et al. Structure of the mycosin-1 protease from the mycobacterial ESX-1 protein type VII secretion system. J. Biol. Chem. 288, 17782–17790 (2013).
Zhang, X. L. et al. Core component EccB1 of the Mycobacterium tuberculosis type VII secretion system is a periplasmic ATPase. FASEB J. 29, 4804–4814 (2015).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
McGowan, S. et al. X-ray crystal structure of the streptococcal specific phage lysin PlyC. Proc. Natl Acad. Sci. USA 109, 12752–12757 (2012).
Zoltner, M. et al. EssC: domain structures inform on the elusive translocation channel in the type VII secretion system. Biochem. J. 473, 1941–1952 (2016).
Arbing, M. A. et al. Heterologous expression of mycobacterial Esx complexes in Escherichia coli for structural studies is facilitated by the use of maltose binding protein fusions. PLoS ONE 8, e81753 (2013).
Rueden, C. T. et al. ImageJ2: ImageJ for the next generation of scientific image data. BMC Bioinformatics 18, 529 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
de la Rosa-Trevín, J. M. et al. Scipion: a software framework toward integration, reproducibility and validation in 3D electron microscopy. J. Struct. Biol. 195, 93–99 (2016).
Källberg, M. et al. Template-based protein structure modeling using the RaptorX web server. Nat. Protoc. 7, 1511–1522 (2012).
Yang, J. and Zhang, Y. Protein structure and function prediction using I-TASSER. Curr. Protoc. Bioinform. 52, 5.8.1–5.8.15 (2015).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430, 2237–2243 (2018).
Mostosi, P., Schindelin, H., Kollmannsberger, P. & Thorn, A. Automated interpretation of cryo-EM density maps with convolutional neural networks. Preprint at https://www.biorxiv.org/content/10.1101/644476v1 (2019).
Kessner, D., Chambers, M., Burke, R., Agus, D. & Mallick, P. ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24, 2534–2536 (2008).
Götze, M. et al. Automated assignment of MS/MS cleavable cross-links in protein 3D-structure analysis. J. Am. Soc. Mass Spectrom. 26, 83–97 (2015).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Fitzpatrick, A. W. P. et al. Structure of the MacAB–TolC ABC-type tripartite multidrug efflux pump. Nat. Microbiol. 2, 17070 (2017).
Acknowledgements
High-resolution electron microscopy data were collected at The Netherlands Center for Electron Nanoscopy (NeCEN) with assistance from C. Diebolder, at the Rudolf Virchow Center Würzburg with the assistance of C. Kraft and T. Rasmussen, and at the Centro Nacional de Biotecnología (CNB-CSIC) with the assistance of R. Arranz and F. J. Chichón. The work at the CNB-CSIC was supported by iNEXT (PID2907 to access CNB-CSIC) grant number 653706, funded by the Horizon 2020 program of the European Union. S. Lamer acquired mass spectrometry data (AG Schlosser, University of Würzburg). We thank M. Steigedal (Norwegian University of Science and Technology) and R. Manganelli (University of Padova) for the M. smegmatis ESX-3 knockout strains; and D. Eisenberg and M. Arbing (University of California, Los Angeles) for the pMAPLe3 overexpression plasmid. Plasmid pMyNT was a gift from A. Parret and M. Wilmanns (Addgene plasmid 42191). We thank C. Fernández Rodríguez (CNIO) for suggestions during processing, and C. Kisker and H. Schindelin (Rudolf Virchow Center, Würzburg) for critical proofreading. This work was supported by the Elite Network of Bavaria (N-BM-2013-246 to S.G.) and the State Research Agency, Spanish Ministry of Science, Innovation and Universities (MCIU/AEI to O.L.) co-funded by the European Regional Development Fund (ERDF) (SAF2017-82632-P to O.L.), by projects Y2018/BIO4747 and P2018/NMT4443 from the Autonomous Region of Madrid and co-funded by the European Social Fund and the European Regional Development Fund (to O.L.). The CNIO is supported by the National Institute of Health Carlos III.
Author information
Authors and Affiliations
Contributions
N.F., A.R.-C., O.L. and S.G. designed the research; N.F., A.R.-C., G.D., B.B., S.G., M.W. and N.M. carried out the research; N.F. performed the biochemistry, sample preparation, grid screening and built the model together with S.G.; A.R.-C. performed the image processing and reconstructions; N.F., A.R.-C., G.D., B.B., O.L. and S.G. analysed data; G.D. and J.M.S. performed the crosslinking mass spectrometry experiments; A.S. performed the iBAQ measurements; R.F.-L. helped with electron microscopy data processing; and N.F., A.R.-C., O.L. and S.G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Purification and cryo-EM of ESX-3 protein complexes.
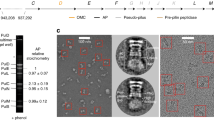
a, Overview of the constructs tested for co-expression studies. The illustrated gene cassettes were cloned into plasmid pMyNT and expressed under the control of the acetamidase promoter (Ace). Ace RBS, acetamidase ribosome-binding site. From top to bottom: the ESX-3 gene cluster (pMyNT:ESX-3), the ESX-3 gene cluster without substrate genes (pMyNT:ESX-3ΔS), without genes encoding substrates and cytosolic proteins (pMyNT:ESX-3ΔSC), only the ESX-3 membrane proteins (pMyNT:Mini), the membrane proteins EccB3, EccC3, EccD3, EccE3 but not MycP3 (pMyNT:MiniΔMycP3). b, Western blot of TeDM-solubilized membrane extracts containing ESX-3EccC3-His6 complexes expressed in M. smegmatis mc2155 from constructs listed in a and separated by BN–PAGE. Anti-His6-tag antibodies were used to detect the His-tagged ESX-3 complexes. The results represent three independent experiments. c, The ESX-3 core complex was expressed in wild-type M. smegmatis mc2155 from plasmid pMyNT:ESX-3 encoding the entire ESX-3 gene cluster and purified; analysis by BN–PAGE (left) and SDS–PAGE (right) is shown. The results represent three independent experiments. d, Size-exclusion chromatogram (Superose-6 Increase 10/300 GL) after the TeDM-to-Amphipol A8-35 exchange of the ESX-3 core complex. The positions of peak 1 (P1) and peak 2 (P2) are indicated with arrows. The results represent three independent experiments. e, Overview micrograph of the higher oligomeric ESX-3 core complex (>1.2 MDa; P1). Some particles are highlighted by circles. Scale bar, 50 nm. f, Representative 2D class average of the larger ESX-3 species. The membrane region is indicated with a bracket. g, Overview micrograph of the 900-kDa ESX-3 core complex (P2). Some particles are highlighted by circles. Scale bar, 50 nm. h, Several 2D averages of the ESX-3 core complex after focused centring on the flexible fork. A flexible attachment of the fork to the membrane region can be observed.
Extended Data Fig. 2 Image processing and classification strategy for the cryo-EM data of the ESX-3 core complex.
a, Tools in Relion and cryoSPARC were used to clean a preliminary dataset and classify the images. Of the particles, 85% corresponded to complexes that comprised protomers and one fork, and these were grouped and refined, whereas the single protomer did not exhibit the fork (EccB3). This analysis suggests that EccB3 is involved in dimerization. b, A larger dataset was collected for high-resolution analysis. After removing bad particles, using 2D and 3D classification methods, a clean dataset was further classified and refined to the core ESX-3 dimer, and the monomer was obtained after particle subtraction. The structure of the fork was resolved after density subtraction and refinement of the new generated particles.
Extended Data Fig. 3 Cryo-EM maps of the ESX-3 core complex.
a–c, Two views of the ESX-3 dimer (a), the ESX-3 monomer (b) and the fork region (EccB3) (c). d, One view of each of the two conformations of the flexible tips of the arms, corresponding to EccC3, obtained after 3D classification. The density of each cryo-EM map is rendered at a low threshold and represented as a transparent density. Within each density, the same cryo-EM map is represented at higher threshold to highlight the structural details at higher resolution in the map.
Extended Data Fig. 4 Estimation of resolution and local resolution EM maps.
a, Left, FSC plot and average resolution estimation for the core structure of the ESX-3 core complex (dimeric structure). Right, local-resolution map and colour scale. b, As in a for the structure of protomer 1. c, As in a for the structure obtained for the fork region. d, e, Plot of the correlation between the model and the map for the ESX-3 dimer (d) and the ESX-3 protomer 1 (e).
Extended Data Fig. 5 Estimation of resolution and local resolution EM maps for the flexible arms.
a, FSC plot and estimation of average resolution for the two conformations (conformation 1 and 2) obtained for the complex after recentring and processing around the flexible tips, corresponding to EccC3. b, Local-resolution maps and colour scale for conformation 1, represented at two different thresholds. At lower threshold density the whole visible tip region is represented but the higher-resolution details of EccC3 domain DI are not visible. Representations at higher threshold reveal these details at the expense of the regions of lower resolution. c, As in b for conformation 2.
Extended Data Fig. 6 Details of the cryo-EM density and atomic models.
a, Representative regions in the cryo-EM density of protomer 1 showing details of high resolution and the atomic model for several domains in the structure. b, In the cryo-EM map of the ESX-3 dimer there is partial density for one TMH of EccC3 for each protomer. The two TMHs were fitted as polyalanine models into the density using Phenix Refine39. c, Plot revealing that density for the two EccC3 TMHs is well defined only at one end of the helix. Notably, the density for one of the EccC3 TMHs is better defined in the structure of the subtracted monomer, and this is represented in the panels showing the monomer structure in Figs. 2, 3.
Extended Data Fig. 7 Validation experiments.
Experiments to validate that the ESX-3 core complex extracted from actively secreting mycobacteria is the same as the complex for which the structure was solved. We investigated the expression of the ESX-3 complex in the mycobacterial membrane, the stoichiometry of the extracted and purified complex and the orientation of the complex in the membrane. a, Overview of the constructs used in secretion assays and for the validation of the structure of the ESX-3 core complex. Plasmid pMyNT:ESX-3 encodes the ESX-3 gene cluster under acetamidase promoter control, and pMyNT:ESX-3i under control of the native IdeR promotor. The expression of the latter construct is induced in iron-depleted culture medium. b, BN–PAGE and western blot analyses (EccC3-His) indicate the presence of the 900-kDa ESX-3 core complex in the membrane of secretion-competent cells that contain the ESX-3 gene cluster under control of either the acetamidase promoter or the native IdeR promoter (see a). The gel represents two independent experiments. c, Comparison of size-exclusion chromatography profiles of ESX-3 core complexes purified from secretion-competent cells transformed with plasmids pMyNT:ESX-3 and pMyNT:ESX-3i (both expressed in M. smegmatis ΔESX-3; see a), with the ESX-3 core complex purified from the minimal expression construct (pMyNT:Mini) expressed in wild-type M. smegmatis, which was used to determine the cryo EM structure (Extended Data Fig. 1a). The chromatograms are representative of three independent purifications. d, ESX-3 core complexes purified from secretion-competent M. smegmatis ΔESX-3 transformed with either pMyNT:ESX-3 or pMyNT:ESX-3i, in the presence and absence of β-mercaptoethanol. A disulfide bridge is found in the EccB3 protein in complexes purified from secretion-competent cells. The two positions for EccB3 in SDS–PAGE are indicated. In the structure of the ESX-3 core complex, EccB3 is periplasmic and contributes to the assembly between protomers. We observe a different mobility for EccB3 in the ESX-3 core complex purified from secretion-competent cells when the SDS–PAGE is performed in the presence or absence of β-mercaptoethanol. This is compatible with an intramolecular disulfide bridge in EccB3, as expected in the oxidative environment of the periplasm. The gel represents two independent experiments. e, Determination of the stoichiometry of ESX-3 complexes in secretion-competent M. smegmatis ΔESX-3 transformed with either pMyNT:ESX-3 or pMyNT:ESX-3i. Oriole staining of the SDS–PAGE bands—in order to estimate the ratio of subunits obtained—showed that EccD3 is over-represented, in agreement with the observed 1:1:2:1 stoichiometry in the structure of the ESX-3 core complex. SDS–PAGE shows that the density of the bands of EccD3 is higher than those of EccE3, despite both proteins having a similar molecular mass. The intensities of the bands were integrated and the relative ratio of subunits was estimated, taking into account the differences in molecular mass of the proteins. In the bar chart, the bars are the reported means and the standard deviations are shown as dots. The results indicated a stoichiometry of 1:1:2:1 (EccB3:EccC3:EccD3:EccE3). The gel shows three technical replicates and is representative of two independent experiments. f, Analysis of the composition of the protein complex subunit using iBAQ. The size of the dots correlates with the number of identified unique peptides. The iBAQ–MS method was, in our experiments, not sufficiently sensitive to define the stoichiometry of ESX-3. We used the ESX-3 complex from our structural studies as an internal control because the stoichiometry of the ESX-3 structure was fully determined as 1:1:2:1 at the high resolution of our cryo-EM maps. The iBAQ method estimates a 1:1:1:1 stoichiometry for the purified complex that we use to resolve the structure by cryo-EM, thus indicating that the method does not report the correct stoichiometry for ESX-3 in these experiments. When the stoichiometry of the native ESX-3 core complex in secretion-competent cells was estimated by the iBAQ method used previously for the ESX-5 core complex19, a 1:1:1:1 ratio was also obtained (data not shown). g, Analysis of purified EccD3 using size-exclusion chromatography (Superose-6 Increase 10/300 GL column) coupled with multi-angle light scattering. The lines show the deconvolution of the contributions of the EccD3–DDM protein–detergent complex (dot-dashed line), the protein (dashed line) and the detergent micelles (dotted line) to the total mass of the complex. This experiment demonstrates that EccD3 dimerizes as observed in structure of the ESX-3 core complex. The experiment is representative of two technical replicates. h, Analysis of the derivatives of the ESX-3 core complex by size-exclusion chromatography. Removing the periplasmic fork of EccB3 or EccE3 leads to dissociation towards single protomers, showing that both components are essential for the stability of the ESX-3 complex. The elution peak of the ESX-3 core complex dimer is indicated. The chromatograms are representative of two independent purifications. i, Isolated mycobacterial membranes from wild-type M. smegmatis containing the ‘high yield’ minimal expression construct (pMyNT:Mini) and extraction of ESX-3 core complexes after crosslinking using DSS. ESX-3 complexes were analysed by BN–PAGE and western blot analyses (EccD3–StrepII). As a control, complexes formed by EccD3 expressed alone were also analysed simultaneously. Only ESX-3 assembled into large complexes, including those of higher molecular mass (higher MW species) than the ESX-3 dimer. The gel is representative of two independent experiments.
Extended Data Fig. 8 Building the ESX-3 hexamer using the ESX-5 negative-stain structure as a template.
a, Modelling of the ESX-3 secretion machine. The negative-stain map of the hexameric ESX-5 core complex (EMDB-3596) was used to generate a model for the ESX-3 secretion machine19. For this, the rigid core of the ESX-3 dimer (without the flexible arm and fork, shown in red) was fitted into ESX-5 (shown as white transparent density) using the contour information of the negative-stain structure. Subsequently, another two dimers were fitted in the remaining subunits to form a hexamer. The atomic structure of each ESX-3 dimer fits approximately the dimensions of ESX-5, as shown in the top and side views of ESX-5. Because the EMDB-3596 reconstruction was generated assuming a perfect six-fold symmetry, all subunits in the ESX-5 complex are identical, whereas the ESX-3 hexamer is based on a trimer of dimers.
Extended Data Fig. 9 The structure of the EccC3/EccB3 contact is reminiscent of ABC type F transporters.
a, Side-by-side comparison between the structure of the DUF and stalk domains of EccC3 interacting with EccB3 in the structure of the ESX-3 core complex, and the MacB ABC transporter (PDB ID: 5NIL)46. The open (ATP unbound) state of MacB is shown. ATP binding and hydrolysis trigger long-range conformational changes in the stalk domains and the periplasmic domains of the MacB dimer, which close and open the transmembrane channel as well as the periplasmic exit.
Extended Data Fig. 10 Comparison between the negative-stain structure of ESX-5 and the cryo-EM structure of ESX-3.
a, Side and top views of the ESX-5 structure, which has been solved previously (EMDB-3596)19. The structure was obtained after applying six-fold symmetry. The use of staining agents limits the obtainable information to the external shape of the complex, while secondary structure elements and internal details are not resolved in negative-stain structures. EccC5 and EccB5 are not visualized in the structure, possibly owing to their flexibility and the use of six-fold symmetry. b, Side and top views of the ESX-3 model (this work).
Supplementary information
Supplementary Tables 1-3
Supplementary Table 1 (Primers and oligonucleotides used for cloning), Supplementary Table 2 (Cryo-EM data collection, refinement and validation statistics) and Supplementary Table 3 (Correlation Coefficients (CC) after model refinement and fitting of homology models).
Supplementary Figure 1
This file contains the uncropped gels presented in the manuscript.
Rights and permissions
About this article
Cite this article
Famelis, N., Rivera-Calzada, A., Degliesposti, G. et al. Architecture of the mycobacterial type VII secretion system. Nature 576, 321–325 (2019). https://doi.org/10.1038/s41586-019-1633-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1633-1
This article is cited by
-
Assembly of a unique membrane complex in type VI secretion systems of Bacteroidota
Nature Communications (2024)
-
Cerium Nanoparticles Can Enhance Humoral and Cellular Immune Responses of Multi-epitope Vaccine Candidates
BioNanoScience (2024)
-
Typ-VII-Sekretionssysteme: Ziele gegen Tuberkulose
BIOspektrum (2022)
-
Type VII secretion systems: structure, functions and transport models
Nature Reviews Microbiology (2021)
-
Toxin secretion and trafficking by Mycobacterium tuberculosis
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.