Abstract
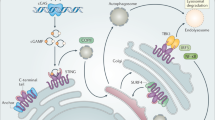
The cyclic GMP–AMP synthase (cGAS)–STING pathway is a central component of the cell-autonomous innate immune system in animals1,2. The cGAS protein is a sensor of cytosolic viral DNA and, upon sensing DNA, it produces a cyclic GMP–AMP (cGAMP) signalling molecule that binds to the STING protein and activates the immune response3,4,5. The production of cGAMP has also been detected in bacteria6, and has been shown, in Vibrio cholerae, to activate a phospholipase that degrades the inner bacterial membrane7. However, the biological role of cGAMP signalling in bacteria remains unknown. Here we show that cGAMP signalling is part of an antiphage defence system that is common in bacteria. This system is composed of a four-gene operon that encodes the bacterial cGAS and the associated phospholipase, as well as two enzymes with the eukaryotic-like domains E1, E2 and JAB. We show that this operon confers resistance against a wide variety of phages. Phage infection triggers the production of cGAMP, which—in turn—activates the phospholipase, leading to a loss of membrane integrity and to cell death before completion of phage reproduction. Diverged versions of this system appear in more than 10% of prokaryotic genomes, and we show that variants with effectors other than phospholipase also protect against phage infection. Our results suggest that the eukaryotic cGAS–STING antiviral pathway has ancient evolutionary roots that stem from microbial defences against phages.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Data that support the findings of this study are available within the article and its Extended Data and Supplementary Tables. GenBank accessions, locus tags and nucleotide ranges of the CBASSs appear in the Methods. IMG gene and genome ID number, contig ID and system and effector classification appear in Supplementary Table 1. Primer sequences for the CBASSs are available in Supplementary Table 2. Any other relevant data are available from the corresponding authors upon reasonable request.
References
Kranzusch, P. J. et al. Ancient origin of cGAS-STING reveals mechanism of universal 2′,3′ cGAMP signaling. Mol. Cell 59, 891–903 (2015).
Margolis, S. R., Wilson, S. C. & Vance, R. E. Evolutionary origins of cGAS-STING signaling. Trends Immunol. 38, 733–743 (2017).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Ablasser, A. et al. cGAS produces a 2′-5′-linked cyclic dinucleotide second messenger that activates STING. Nature 498, 380–384 (2013).
Ishikawa, H., Ma, Z. & Barber, G. N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Davies, B. W., Bogard, R. W., Young, T. S. & Mekalanos, J. J. Coordinated regulation of accessory genetic elements produces cyclic di-nucleotides for V. cholerae virulence. Cell 149, 358–370 (2012).
Severin, G. B. et al. Direct activation of a phospholipase by cyclic GMP-AMP in El Tor Vibrio cholerae. Proc. Natl Acad. Sci. USA 115, E6048–E6055 (2018).
Makarova, K. S., Wolf, Y. I., Snir, S. & Koonin, E. V. Defense islands in bacterial and archaeal genomes and prediction of novel defense systems. J. Bacteriol. 193, 6039–6056 (2011).
Goldfarb, T. et al. BREX is a novel phage resistance system widespread in microbial genomes. EMBO J. 34, 169–183 (2015).
Ofir, G. et al. DISARM is a widespread bacterial defence system with broad anti-phage activities. Nat. Microbiol. 3, 90–98 (2018).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Iyer, L. M., Burroughs, A. M. & Aravind, L. The prokaryotic antecedents of the ubiquitin-signaling system and the early evolution of ubiquitin-like β-grasp domains. Genome Biol. 7, R60 (2006).
Kato, K., Ishii, R., Hirano, S., Ishitani, R. & Nureki, O. Structural basis for the catalytic mechanism of DncV, bacterial homolog of cyclic GMP-AMP synthase. Structure 23, 843–850 (2015).
Molineux, I. J. Host–parasite interactions: recent developments in the genetics of abortive phage infections. New Biol. 3, 230–236 (1991).
Walker, J. T. & Walker, D. H. Mutations in coliphage P1 affecting host cell lysis. J. Virol. 35, 519–530 (1980).
Whiteley, A. T. et al. Bacterial cGAS-like enzymes synthesize diverse nucleotide signals. Nature 567, 194–199 (2019).
Snyder, L. Phage-exclusion enzymes: a bonanza of biochemical and cell biology reagents? Mol. Microbiol. 15, 415–420 (1995).
Burroughs, A. M., Zhang, D., Schäffer, D. E., Iyer, L. M. & Aravind, L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Res. 43, 10633–10654 (2015).
Joshua-Tor, L. & Hannon, G. J. Ancestral roles of small RNAs: an Ago-centric perspective. Cold Spring Harb. Perspect. Biol. 3, a003772 (2011).
Swarts, D. C. et al. DNA-guided DNA interference by a prokaryotic argonaute. Nature 507, 258–261 (2014).
Olovnikov, I., Chan, K., Sachidanandam, R., Newman, D. K. & Aravin, A. A. Bacterial argonaute samples the transcriptome to identify foreign DNA. Mol. Cell 51, 594–605 (2013).
Akira, S. & Takeda, K. Toll-like receptor signalling. Nat. Rev. Immunol. 4, 499–511 (2004).
Zhou, A. et al. Interferon action and apoptosis are defective in mice devoid of 2′,5′-oligoadenylate-dependent RNase L. EMBO J. 16, 6355–6363 (1997).
Kazlauskiene, M., Kostiuk, G., Venclovas, Č., Tamulaitis, G. & Siksnys, V. A cyclic oligonucleotide signaling pathway in type III CRISPR-Cas systems. Science 357, 605–609 (2017).
Margulis, L. Archaeal–eubacterial mergers in the origin of Eukarya: phylogenetic classification of life. Proc. Natl Acad. Sci. USA 93, 1071–1076 (1996).
Chen, I. A. et al. IMG/M v.5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 47, D666–D677 (2019).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Madeira, F. et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 47, W636–W641 (2019).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430, 2237–2243 (2018).
Berman, H., Henrick, K. & Nakamura, H. Announcing the worldwide Protein Data Bank. Nat. Struct. Biol. 10, 980 (2003).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47, D427–D432 (2019).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 26, 1641–1650 (2009).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Lam, V. et al. Resorufin butyrate as a soluble and monomeric high-throughput substrate for a triglyceride lipase. J. Biomol. Screen. 17, 245–251 (2012).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015).
Acknowledgements
We thank A. Leavitt and S. Sharir for assistance in DNA extraction, library preparation and sequencing, A. Bernheim for assistance in data visualization, and members of the Sorek laboratory for fruitful discussions. This study was supported in part by the Israel Science Foundation (personal grant 1360/16), the European Research Council (grant ERC-CoG 681203), the Ernest and Bonnie Beutler Research Program of Excellence in Genomic Medicine, and the Knell Family Center for Microbiology. A.M. was supported by a fellowship from the Ariane de Rothschild Women Doctoral Program.
Author information
Authors and Affiliations
Contributions
D.C., S.M. and G.A. led the study and performed all experiments unless otherwise indicated. A.M. performed the computational analyses that appear in Figs. 1 and 3. Y.O.-S. performed the microscopy analysis that appears in Extended Data Fig. 7. G.S. assisted with the plaque assays that appear in Fig. 1 and Extended Data Figs. 1 and 3. A.K. and S.D. performed the computational analyses that led to Extended Data Fig. 8. R.S. supervised the study and wrote the paper together with the team.
Corresponding authors
Ethics declarations
Competing interests
R.S. is a scientific cofounder and consultant of BiomX Ltd, Pantheon Ltd and Ecophage Ltd.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Zhijian ‘James’ Chen, Karen Maxwell and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Fold antiphage defence conferred by four-gene defence systems against various phages.
The four-gene operon from either V. cholerae El Tor or E. coli TW11681 was cloned into E. coli MG1655 (Methods). Fold antiphage defence, as measured by plaque assays, is shown. The fold defence was calculated as the ratio between the efficiency of plating of the phage on the operon-lacking control strain and the efficiency of plating on the operon-containing strain (Fig. 2b, Methods). Bar graph represents average of three independent replicates, with individual data points overlaid. Points that fall below the x axis (for SECphi17, SECphi18 and SECphi27) denote values lower than 1.
Extended Data Fig. 2 Transformation efficiency assays.
Transformation efficiency of plasmid pRSFDuet-1 into strains that contain the four-gene operon derived from E. coli TW11681 or from V. cholerae El Tor, presented as a fraction of the transformation efficiency to E. coli MG1655 carrying an empty vector instead of the four-gene operon. Bar graph represents average of three independent replicates, with individual data points overlaid.
Extended Data Fig. 3 Efficiency of plating of coliphages on defence systems with whole-gene deletions or point mutations.
The efficiency of plating of phages infecting strains with the wild-type E.-coli-derived four-gene, deletion strains and strains with point mutations. Data represent plaque-forming units per millilitre; bar graphs represent average of three independent replicates, with individual data points overlaid. Empty vector represents a control E. coli MG1655 strain that lacks the system and has an empty vector instead. a, Infection with the phage T4. b, Infection with the phage T5. c, Infection with the phage T6. d, Infection with the phage λ-vir.
Extended Data Fig. 4 Efficiency of plating of phage P1 on a double-deletion strain.
The efficiency of plating is shown of phage P1 infecting strains with the wild-type E.-coli-derived four-gene system, strains with individual genes deleted and a strain with two genes deleted. Data represent plaque-forming units per millilitre; bar graphs represent average of three independent replicates, with individual data points overlaid. Empty vector represents a control E. coli MG1655 strain that lacks the system and has an empty vector instead.
Extended Data Fig. 5 The bacterial CBASS functions through abortive infection.
a, Growth curves in liquid culture for CBASS-containing and CBASS-lacking (empty vector) bacteria infected by phage SECphi18 at 25 °C. Bacteria were infected at time = 0 at an MOI of 0.02 or 2. Three independent replicates for each MOI are shown, and each curve shows an individual replicate. b, Growth curves in liquid culture for cells containing a minimal CBASS comprising phospholipase–cGAS (capV-dncV) only. Bacteria were infected at time = 0 at an MOI of 2 by phage P1. Three independent replicates for each MOI are shown, and each curve shows an individual replicate.
Extended Data Fig. 6 Cell sorting of infected cells stained with propidium iodide.
Cells containing the CBASS derived from E. coli TW11681, and control cells containing an empty vector, were stained with propidium iodide, a fluorescent DNA-binding agent that penetrates cells that have impaired membrane integrity. Cells were infected by phage P1 (MOI of 2) and sorted on the basis of propidium-iodide fluorescence intensity (y axis); the x axis represents forward scatter. a, Uninfected cells that lack the CBASS. b, Uninfected cells that contain the CBASS. c, Cells that lack the CBASS, 40 min after infection. d, Cells that contain the CBASS, 40 min after infection. A large population of cells with high propidium-iodide fluorescence intensity is observed. Data from a representative replicate of two independent replicates are shown.
Extended Data Fig. 7 Microscopy of infected cells.
a–c, Phase contrast and overlay images are shown, of membrane stain (red) and DAPI (blue) images captured at 20 min (a), 40 min (b) and 60 min (c) after infection with phage P1 at an MOI of 2. The two columns on the left show E. coli MG1655 cells containing the CBASS derived from E. coli TW11681. The two columns on the right show E.coli MG1655 cells containing the CBASS derived from E. coli with a single point mutation that inactivates the CapV phospholipase (CapV(S60A)). Cell shape is deformed after 40 min in cells containing the CBASS, but not in cells in which the CBASS is mutated. After 60 min, phage-mediated cell lysis is observed in cells in which the CBASS is mutated. Representative images from a single replicate out of two independent replicates are shown.
Extended Data Fig. 8 A two-gene CBASS protects Bacillus against phage infection.
a, Domain organization of a two-gene operon found in the B. cereus VD146 genome. Locus tags of the depicted genes are indicated below each gene. b, The two-gene operon from B. cereus VD146 was cloned and genomically integrated into B. subtilis BEST7003, which naturally lacks this system. The efficiency of plating of phage SBSphiC infecting the CBASS-lacking and CBASS-containing strains, as well as strains in which one of the two genes was deleted, is shown. Bar graph represents average of three independent replicates, with individual data points overlaid. c, Growth curves in liquid culture for B. subtilis containing the B. cereus two-gene CBASS, or CBASS-lacking B. subtilis that contains an empty vector instead, infected by phage SBSphiC. Bacteria were infected at time = 0 at an MOI of 0.2 or 2. Three independent replicates for each MOI are shown, and each curve shows an individual replicate.
Extended Data Fig. 9 Domain analysis and homology-based structure prediction of a bacterial TIR–STING protein.
a, Schematics of HHpred29 homology-based search results of the Prevotella corporis TIR–STING protein (Supplementary Table 1). b, Phyre235 secondary structure prediction of the TIR domain in the P. corporis TIR–STING protein, compared to the solved crystal structure of the human TIR domain protein MyD88 (PDB accession 2Z5V_A). c, Phyre235 secondary structure prediction of the STING domain in the P. corporis TIR–STING protein, compared to the solved crystal structure of the human STING protein (PDB accession 5BQX_A). Black, identical residues; grey, similar residues. Secondary structure prediction for the bacterial protein appears above the alignment; secondary structure of solved human domain appears below the alignment. d, Structural alignment of human TIR domain protein MYD88 and the modelled bacterial TIR domain. e, Structural alignment of human STING domain and the modelled bacterial STING domain. In d, e, blue and red represent the structure of the human protein and the model of the bacterial domain structure, respectively.
Supplementary information
41586_2019_1605_MOESM1_ESM.xlsx
Supplementary Tables Supplementary Table 1: CBASS systems analyzed in this study includes the Joint Genome Institute (JGI) gene and genome ID number, genome name, contig ID, system type, and the protein effector type. Supplementary Table 2: Primers used in this study includes the primer name, sequence, and description.
Rights and permissions
About this article
Cite this article
Cohen, D., Melamed, S., Millman, A. et al. Cyclic GMP–AMP signalling protects bacteria against viral infection. Nature 574, 691–695 (2019). https://doi.org/10.1038/s41586-019-1605-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1605-5
This article is cited by
-
cGAS-STING, inflammasomes and pyroptosis: an overview of crosstalk mechanism of activation and regulation
Cell Communication and Signaling (2024)
-
Phage defence system CBASS is regulated by a prokaryotic E2 enzyme that imitates the ubiquitin pathway
Nature Microbiology (2024)
-
Conservation and similarity of bacterial and eukaryotic innate immunity
Nature Reviews Microbiology (2024)
-
Phages overcome bacterial immunity via diverse anti-defence proteins
Nature (2024)
-
The Vibrio cholerae CBASS phage defence system modulates resistance and killing by antifolate antibiotics
Nature Microbiology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.