Abstract
Soluble guanylate cyclase (sGC) is the primary sensor of nitric oxide. It has a central role in nitric oxide signalling and has been implicated in many essential physiological processes and disease conditions. The binding of nitric oxide boosts the enzymatic activity of sGC. However, the mechanism by which nitric oxide activates the enzyme is unclear. Here we report the cryo-electron microscopy structures of the human sGCα1β1 heterodimer in different functional states. These structures revealed that the transducer module bridges the nitric oxide sensor module and the catalytic module. Binding of nitric oxide to the β1 haem-nitric oxide and oxygen binding (H-NOX) domain triggers the structural rearrangement of the sensor module and a conformational switch of the transducer module from bending to straightening. The resulting movement of the N termini of the catalytic domains drives structural changes within the catalytic module, which in turn boost the enzymatic activity of sGC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Cryo-EM maps of the haem-unliganded, haem-oxidized, NO-activated and β1(H105C) mutant sGC structures have been deposited in the EMDB under accession numbers EMD-9883, EMD-9884, EMD-9885 and EMD-9886, respectively. Atomic coordinates of the haem-unliganded, haem-oxidized and NO-activated sGC structures have been deposited in the PDB under accession numbers 6JT0, 6JT1 and 6JT2, respectively.
References
Hollenberg, S. M. & Cinel, I. Bench-to-bedside review: nitric oxide in critical illness—update 2008. Crit. Care 13, 218 (2009).
Horst, B. G. & Marletta, M. A. Physiological activation and deactivation of soluble guanylate cyclase. Nitric Oxide 77, 65–74 (2018).
Luiking, Y. C., Engelen, M. P. & Deutz, N. E. Regulation of nitric oxide production in health and disease. Curr. Opin. Clin. Nutr. Metab. Care 13, 97–104 (2010).
Farah, C., Michel, L. Y. M. & Balligand, J. L. Nitric oxide signalling in cardiovascular health and disease. Nat. Rev. Cardiol. 15, 292–316 (2018).
Derbyshire, E. R. & Marletta, M. A. Structure and regulation of soluble guanylate cyclase. Annu. Rev. Biochem. 81, 533–559 (2012).
Deloukas, P. et al. Large-scale association analysis identifies new risk loci for coronary artery disease. Nat. Genet. 45, 25–33 (2013).
Hervé, D. et al. Loss of α1β1 soluble guanylate cyclase, the major nitric oxide receptor, leads to moyamoya and achalasia. Am. J. Hum. Genet. 94, 385–394 (2014).
Wallace, S. et al. Disrupted nitric oxide signaling due to GUCY1A3 mutations increases risk for moyamoya disease, achalasia and hypertension. Clin. Genet. 90, 351–360 (2016).
Sandner, P. From molecules to patients: exploring the therapeutic role of soluble guanylate cyclase stimulators. Biol. Chem. 399, 679–690 (2018).
Koesling, D., Mergia, E. & Russwurm, M. Physiological functions of NO-sensitive guanylyl cyclase isoforms. Curr. Med. Chem. 23, 2653–2665 (2016).
Montfort, W. R., Wales, J. A. & Weichsel, A. Structure and activation of soluble guanylyl cyclase, the nitric oxide sensor. Antioxid. Redox Signal. 26, 107–121 (2017).
Dasgupta, A., Bowman, L., D’Arsigny, C. L. & Archer, S. L. Soluble guanylate cyclase: a new therapeutic target for pulmonary arterial hypertension and chronic thromboembolic pulmonary hypertension. Clin. Pharmacol. Ther. 97, 88–102 (2015).
Zhao, Y. et al. Inhibition of soluble guanylate cyclase by ODQ. Biochemistry 39, 10848–10854 (2000).
Purohit, R., Weichsel, A. & Montfort, W. R. Crystal structure of the Alpha subunit PAS domain from soluble guanylyl cyclase. Protein Sci. 22, 1439–1444 (2013).
Ma, X., Beuve, A. & van den Akker, F. Crystal structure of the signaling helix coiled-coil domain of the β1 subunit of the soluble guanylyl cyclase. BMC Struct. Biol. 10, 2 (2010).
Allerston, C. K., von Delft, F. & Gileadi, O. Crystal structures of the catalytic domain of human soluble guanylate cyclase. PLoS ONE 8, e57644 (2013).
Seeger, F. et al. Interfacial residues promote an optimal alignment of the catalytic center in human soluble guanylate cyclase: heterodimerization is required but not sufficient for activity. Biochemistry 53, 2153–2165 (2014).
Campbell, M. G., Underbakke, E. S., Potter, C. S., Carragher, B. & Marletta, M. A. Single-particle EM reveals the higher-order domain architecture of soluble guanylate cyclase. Proc. Natl Acad. Sci. USA 111, 2960–2965 (2014).
Underbakke, E. S. et al. Nitric oxide-induced conformational changes in soluble guanylate cyclase. Structure 22, 602–611 (2014).
Martin, E., Sharina, I., Kots, A. & Murad, F. A constitutively activated mutant of human soluble guanylyl cyclase (sGC): implication for the mechanism of sGC activation. Proc. Natl Acad. Sci. USA 100, 9208–9213 (2003).
Nakane, T., Kimanius, D., Lindahl, E. & Scheres, S. H. Characterisation of molecular motions in cryo-EM single-particle data by multi-body refinement in RELION. eLife 7, e36861 (2018).
Tesmer, J. J. et al. Two-metal-ion catalysis in adenylyl cyclase. Science 285, 756–760 (1999).
Denninger, J. W. et al. Interaction of soluble guanylate cyclase with YC-1: kinetic and resonance Raman studies. Biochemistry 39, 4191–4198 (2000).
Sayed, N., Baskaran, P., Ma, X., van den Akker, F. & Beuve, A. Desensitization of soluble guanylyl cyclase, the NO receptor, by S-nitrosylation. Proc. Natl Acad. Sci. USA 104, 12312–12317 (2007).
Mayer, B. et al. Inactivation of soluble guanylate cyclase by stoichiometric S-nitrosation. Mol. Pharmacol. 75, 886–891 (2009).
Fernhoff, N. B., Derbyshire, E. R., Underbakke, E. S. & Marletta, M. A. Heme-assisted S-nitrosation desensitizes ferric soluble guanylate cyclase to nitric oxide. J. Biol. Chem. 287, 43053–43062 (2012).
Koglin, M. & Behrends, S. A functional domain of the α1 subunit of soluble guanylyl cyclase is necessary for activation of the enzyme by nitric oxide and YC-1 but is not involved in heme binding. J. Biol. Chem. 278, 12590–12597 (2003).
Ruiz-Stewart, I. et al. Guanylyl cyclase is an ATP sensor coupling nitric oxide signaling to cell metabolism. Proc. Natl Acad. Sci. USA 101, 37–42 (2004).
Cary, S. P., Winger, J. A. & Marletta, M. A. Tonic and acute nitric oxide signaling through soluble guanylate cyclase is mediated by nonheme nitric oxide, ATP, and GTP. Proc. Natl Acad. Sci. USA 102, 13064–13069 (2005).
Tsai, A. L., Berka, V., Sharina, I. & Martin, E. Dynamic ligand exchange in soluble guanylyl cyclase (sGC): implications for sGC regulation and desensitization. J. Biol. Chem. 286, 43182–43192 (2011).
Vermehren-Schmaedick, A., Ainsley, J. A., Johnson, W. A., Davies, S. A. & Morton, D. B. Behavioral responses to hypoxia in Drosophila larvae are mediated by atypical soluble guanylyl cyclases. Genetics 186, 183–196 (2010).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Li, N. et al. Structure of a pancreatic ATP-sensitive potassium channel. Cell 168, 101–110 (2017).
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006).
Tang, Q. et al. Structure of the receptor-activated human TRPC6 and TRPC3 ion channels. Cell Res. 28, 746–755 (2018).
Olesen, S. P. et al. Characterization of NS 2028 as a specific inhibitor of soluble guanylyl cyclase. Br. J. Pharmacol. 123, 299–309 (1998).
Brandwein, H. J., Lewicki, J. A., Waldman, S. A. & Murad, F. Effect of GTP analogues on purified soluble guanylate cyclase. J. Biol. Chem. 257, 1309–1311 (1982).
Artz, J. D., Toader, V., Zavorin, S. I., Bennett, B. M. & Thatcher, G. R. In vitro activation of soluble guanylyl cyclase and nitric oxide release: a comparison of NO donors and NO mimetics. Biochemistry 40, 9256–9264 (2001).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Jurcik, A. et al. CAVER Analyst 2.0: analysis and visualization of channels and tunnels in protein structures and molecular dynamics trajectories. Bioinformatics 34, 3586–3588 (2018).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Acknowledgements
We thank all members of the Chen laboratory for their help; J. Han for sharing the human sGC cDNA; and D. B. Morton for sharing Drosophila and mouse sGC cDNAs. Cryo-EM data collection was supported by the Electron Microscopy Laboratory and Cryo-EM Platform of Peking University with the assistance of X. Li, Z. Guo, B. Shao, X. Pei and G. Wang. Part of the structural computation was also performed on the Computing Platform of the Center for Life Science and High-performance Computing Platform of Peking University. The work was supported by grants from the Ministry of Science and Technology of China (National Key R&D Program of China, 2016YFA0502004 to L.C.), National Natural Science Foundation of China (91857000, 31622021, 31821091 and 31870833 to L.C.), Beijing Natural Science Foundation (5192009 to L.C.), Young Thousand Talents Program of China (to L.C.) and the China Postdoctoral Science Foundation (2016M600856, 2017T100014 and 2019M650324, and 2019T120014 to J.-X.W.). J.-X.W. is supported by the Boya Postdoctoral Fellowship of Peking University.
Author information
Authors and Affiliations
Contributions
L.C. initiated the project and screened expression constructs. R.L. purified protein and prepared the cryo-EM sample. Y.K. and R.L. collected the cryo-EM data with the help of J.-X.W. Y.K. processed the cryo-EM data with the help of L.C. L.C. built and refined the atomic model. Y.K. did the enzymatic activity assay. R.L. did the disulfide bond cross-linking experiment. All authors contributed to manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Mark Gladwin, Jesus Tejero, Focco van den Akker and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Biochemical characterization of the human α1β1 sGC heterodimer protein and single-particle cryo-EM data processing procedure for sGC in the inactive (haem-oxidized) state.
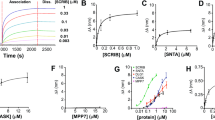
a, Size-exclusion chromatography of sGC on a superdex 200 column. The fractions indicated by dashed lines were pooled for cryo-EM sample preparation. b, SDS–PAGE of the size-exclusion chromatography fractions labelled in a. Arrows show the positions of the α1 and β1 subunits. For gel source data, see Supplementary Fig. 1. c, UV-vis spectra of purified sGC before (red) and after (black) treatment with the haem oxidant NS2028. The positions of the Soret peaks are indicated by arrowheads. a–c, The experiments were repeated independently three times with similar results. d, A representative raw micrograph of sGC in the inactive (haem-oxidized) state. e, Representative 2D class averages of sGC in the inactive (haem-oxidized) state. f, The angular distribution for the consensus refinement of the inactive (haem-oxidized) state is indicated by the sizes of spheres. g, The cryo-EM data processing workflow for sGC in the inactive (haem-oxidized) state.
Extended Data Fig. 2 Conformational heterogeneity and local density quality of sGC in the inactive (haem-oxidized) state.
a, Gold-standard FSC curves of haem-oxidized sGC after correction for masking effects. Resolution estimations were based on the criterion of the FSC 0.143 cutoff. b, Local resolution distribution of the composite map of sGC in the inactive (haem-oxidized) state. c, Histogram of the eigenvectors that contribute to the variance. The top eigenvector is highlighted in grey. d, Histogram of the amplitudes along the top eigenvectors shows monomodal distribution. Particle populations with amplitudes less than −3 or greater than 3 are indicated as red and blue arrows, respectively. e, A 4 Å low-pass filtered map reconstructed from particles indicated as red and blue arrows in d. N-lobes were used for alignment. f, Representative cryo-EM densities of fragments from each individual domain. g, Representative cryo-EM densities of several key residues involving the interactions between β1 H-NOX and adjacent domains in the inactive (haem-oxidized) state. h, The cryo-EM map of the sGC in the haem-oxidized state. The putative linkers between the H-NOX and PAS domains are shown in grey. The B-factor of the map was adjusted to −100 Å2 during the post-processing procedure to visualize features with high flexibility. i, Cryo-EM maps of the catalytic module in the haem-oxidized state (cyan) and the haem-unliganded state (purple). The density of the putative phosphate groups is shown in grey.
Extended Data Fig. 3 Single particle cryo-EM data processing of the sGC sample in the NO-activated state.
a, Representative raw micrograph of sGC in the NO-activated state. b, Representative 2D class averages of sGC in the NO-activated state. c, The angular distribution for the consensus refinement of the NO-activated state is indicated by the sizes of the spheres. d, The cryo-EM data processing workflow for sGC in the NO-activated state. e, Gold-standard FSC curves (after correction for the masking effects) of NO-activated sGC. Resolution estimations are based on the criterion of FSC 0.143 cutoff. f, Local resolution distribution of the composite map of sGC in the NO-activated state.
Extended Data Fig. 4 Conformational heterogeneity and local density quality of sGC in the NO-activated state.
a, Histogram of the eigenvectors that contribute to the variance. The top eigenvector is highlighted in grey. b, Histogram of the amplitudes along the top eigenvectors shows monomodal distribution. Particle populations with amplitudes less than −2 or greater than 2 are indicated with red and blue arrows, respectively. c, 4 Å low-pass filtered maps reconstructed from particles indicated as red and blue arrows in b. N-lobes were used for alignment. d, Representative cryo-EM densities of fragments from each individual domain. e, Representative cryo-EM densities of several key residues involving the interactions between β1 H-NOX and adjacent domains in the NO-activated state. f, Cryo-EM map of the catalytic module in the NO-activated state (yellow). The putative density of the α1 C terminus is shown in grey, αQ of the α1 subunit is shown in pink, and the αO–βK fragment of the β1 subunit is shown in cyan. The B-factor of the map was adjusted to −100 Å2 in post-processing to visualize features with high flexibility. g, The side view of the cryo-EM map of the β1(H105C) mutant sGC. h, Gold-standard FSC curves (after correction for masking effects) of the β1(H105C) mutant sGC. Resolution estimations were based on the criterion of FSC 0.143 cutoff. i, Cryo-EM map of β1(H105C) mutant sGC (pink) and NO-activated sGC (cyan). The haem density is shown in yellow. The map of NO-activated sGC was low-pass filtered to 6.8 Å. j, The locations of cysteine residues that are involved in sGC desensitization (α1 C244, β1 C78 and β1 C122) are indicated with their Cα atoms shown as red spheres. Because the loop containing α1 C244 is disordered in the NO-activated state, only the termini of the loop (α1 L235 and α1 Y252) containing α1 C244 are labelled.
Extended Data Fig. 5 Sequence alignment of the sGC α-subunit.
The sequences of the Homo sapiens α1 subunit, H. sapiens α2 subunit, Danio rerio α1 subunit, D. rerio α2 subunit, and M. sexta α1 subunit are aligned. Conserved residues are coloured in grey. Residues that are mutated to cysteines for oxidative cross-linking are indicated with a black box. Mutations for activity assay are indicated with a red box. The residues corresponding to H105 in the β1 subunit are indicated with a yellow box. Secondary structural elements are indicated as follows: arrows, β-sheets; cylinders, α-helices; lines, loops. Unmodelled residues are shown as dashed lines. The colours of arrows and cylinders are as in Fig. 1a.
Extended Data Fig. 6 Sequence alignment of the sGC β1 subunit.
The sequences of the H. sapiens β1 subunit, D. rerio β1 subunit, and M. sexta β1 subunit are aligned. Conserved residues are coloured in grey. Residues that are mutated to cysteines for oxidative cross-linking are indicated with a black box. Mutations for activity assay are indicated with a red box. Secondary structural elements are indicated as follows: arrows, β-sheets; cyinders, α-helices; lines, loops. Unmodelled residues are shown as dashed lines. The colours of arrows and cylinders are as in Fig. 1a.
Extended Data Fig. 7 Domain–domain interfaces of sGC in the inactive state.
a, Side view of soluble guanylate cyclase in the inactive state, highlighting key interfaces (grey rectangles). Each domain is coloured as in Fig. 1a. The surface of sGC is shown in transparency. b, The interface between the α1 H-NOX domain and the PAS domains boxed in a. c, The interface between the PAS domains boxed in a. d, The interface between β1 H-NOX and adjacent domains boxed in a. e, A 180° rotated view compared to d. f, The structure of the transducer module boxed in a. The side chains of α1 L425 and β1 L365 that are in close proximity are shown as spheres. g, The interface between the transducer module and the catalytic module boxed in a. h, A 90° rotated top view compared to g. i, End-point activity of the less-Cys construct (sGCLC, α1LC + β1LC) compared to the wild-type sGC with CGFP. Mean ± s.d., n = 3 biologically independent samples. j, SDS–PAGE of the in vitro disulfide bond cross-linking experiment of α1LC (L275C) with β1LC(A316C) mutants under reducing and non-reducing conditions. The in-gel GFP fluorescence of the α1 subunit is shown in black on a white background. The position of cross-linked heterodimer is indicated with a red asterisk. Oxidative cross-linking happened only when the cysteine mutants, α1(L275C) and β1(A316C), were present in both subunits simultaneously. The experiments were repeated independently three times with similar results. For gel source data, see Supplementary Fig. 1. k, SDS–PAGE of the in vitro disulfide bond cross-linking experiment of α1LC(L425C) with β1LC(L365C) under reducing and non-reducing conditions. Oxidative cross-linking happened only when the cysteine mutants, α1(L425C) and β1(L365C), were present in both subunit simultaneously. For gel source data, see Supplementary Fig. 1. The experiments were repeated independently three times with similar results. l, SDS–PAGE of the in vitro disulfide bond cross-linking experiment of α1LC(L275C) with β1LC(L365C) and α1LC(L425C) with β1LC(A316C) under reducing and non-reducing conditions. In contrast to α1(L275C) with β1(A316C) and α1(L425C) with β1(L365C), α1(L275C) did not crosslink with β1(L365C), and α1(L425C) did not crosslink with β1(A316C), owing to their long spatial distance. For gel sourcfe data, see Supplementary Fig. 1. The experiments were repeated independently twice with similar results.
Extended Data Fig. 8 Domain–domain interfaces of sGC in the NO-activated state.
a, Superposition of the NO-bound β1 H-NOX domain structure (purple) onto the inactive state structure (grey) by alignment of the αF helices. The steric clashes between the side chains of the NO-bound β1 H-NOX domain (purple sphere) and the side chains of the PAS and CC domains of the inactive state (grey sphere) are marked by red circles if their atom-to-atom distances are smaller than 2.2 Å. The arrow indicates the positional change of the Cα atoms of β1 N62 induced by NO binding. b, The interface between β1 H-NOX and adjacent domains of sGC in the NO-activated state. c, A 180° rotated view compared to b. d, Soret peaks of the sGC mutants show markedly decreased NO activation. The experiments were repeated independently twice with similar results. e, The transducer module in the NO-activated state, coloured as in Fig. 1a. f, Top view of the structural comparison of the catalytic module between the inactive state (grey) and the NO-activated state (coloured). The GMPCPP molecule is shown as sticks. The Cα atoms of α1 P459 and β1 P399 are shown as spheres.
Extended Data Fig. 9 Structural comparisons of each domain.
a, Structural comparison of the full-length human sGC between the haem-oxidized state (coloured) and the haem-unliganded state (grey). b, Structural comparison between the α1 H-NOX domain (pink) and the β1 H-NOX domain (blue) in the inactive state. Both α1 H-NOX and β1 H-NOX share common structural features with prokaryotic H-NOX domains, which are composed of both N-terminal and C-terminal subdomains. The N-terminal αA helix of the α1 subunit that occupies the haem binding pocket is shown in red. The haem molecule of the β1 H-NOX domain is shown as a yellow stick. The approximate boundaries of N-terminal and C-terminal subdomains are indicated by dashed lines. c, Structural comparison between the human α1 PAS domain (red) and the M. sexta Ms α PAS domain (grey, PDB ID:4GJ4). d, Structural comparison between the human β1 PAS domain (blue) and the M. sexta Ms α PAS domain (grey, PDB ID:4GJ4). e, Structural comparison between the catalytic module of the full-length sGC in the inactive state (coloured) and the isolated catalytic domain heterodimer (grey, PDB ID: 4NI2). The β1 subunit was used for structural alignment. f, 90° rotated view compared to e. g, Structural comparison between the catalytic module of the full-length sGC in the inactive state (coloured) and the catalytic domain of the active adenylate cyclase (grey, PDB ID: 1CJU, chain A&B). The β1 subunit was used for structural alignment. The residues of sGC that are within 2.2 Å of the substrate are considered as sterical clashes and shown as red spheres. h, Structural comparison between the NO-activated state (purple) and the inactive state (grey) of the human β1 H-NOX domain. The N-terminal subdomain was used for alignment and the movements are indicated as red arrows. i, Structural comparison between the catalytic module of the full-length sGC in the NO-activated (colored) and the isolated catalytic domain heterodimer (grey, PDB ID: 4NI2). The β1 subunit was used for structural alignment. A inter-domain rotational conformational change is observed. j, A 90° rotated view compared to i. k, Cutaway views of the sGC catalytic module in the inactive state and the NO-activated state. The catalytic module is shown in surface representation colored by electrostatic potential calculated in Pymol. The pockets inside the catalytic module are indicated by arrows. GMPCPP molecule is shown as sticks. l, Structural comparison of the catalytic core between the active adenylate cyclase (grey, PDB ID: 1CJU, chain A and B) and sGC in the NO-activated state (coloured). m, Structural comparison between the NO-activated state (coloured) and the inactive state (grey) of the human α1 and β1 PAS heterodimer.
Supplementary information
Supplementary Figure 1
This file contains the uncropped images of SDS-PAGE gels with size marker indicated.
Supplementary Notes
This file contains detailed analysis of domain-domain interactions observed in the structures and relevant discussions.
Supplementary Video 1 | Flexibility of sGC in the inactive state along the first and second eigenvectors.
Repositioning of reconstructed densities of sGC in the inactive state along the first and the second eigenvectors reveals the flexibility of the CC domain.
Supplementary Video 2 | Flexibility of sGC in the NO-activated state along the first and second eigenvectors.
Repositioning of reconstructed densities of sGC in the NO-activated state along the first and the second eigenvectors reveals the flexibility of the CC domain.
Supplementary Video 3 | Structure of sGC in the inactive state.
The cryo-EM map and the domain organization of sGC in the inactive state.
Supplementary Video 4 | Structure of sGC in the NO-activated state.
The cryo-EM map and the domain organization of sGC in the NO-activated state.
Supplementary Video 5 | Conformational changes of sGC during NO activation.
When the PAS domains are aligned, the transducer and catalytic modules have a swing-like motion during the NO-activation. The transducer module switches from a highly bent conformation to a straightened conformation, which leads to the structural reorganization of the catalytic module to adopt a catalytically competent conformation.
Rights and permissions
About this article
Cite this article
Kang, Y., Liu, R., Wu, JX. et al. Structural insights into the mechanism of human soluble guanylate cyclase. Nature 574, 206–210 (2019). https://doi.org/10.1038/s41586-019-1584-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1584-6
This article is cited by
-
The 10th International Conference on cGMP 2022: recent trends in cGMP research and development—meeting report
Naunyn-Schmiedeberg's Archives of Pharmacology (2023)
-
Sex differences and related estrogenic effects in heart failure with preserved ejection fraction
Heart Failure Reviews (2022)
-
Activation mechanism of human soluble guanylate cyclase by stimulators and activators
Nature Communications (2021)
-
Soluble guanylate cyclase signalling mediates etoposide resistance in progressing small cell lung cancer
Nature Communications (2021)
-
Novel therapeutics for the treatment of hypertension and its associated complications: peptide- and nonpeptide-based strategies
Hypertension Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.