Abstract
Metastasis—the disseminated growth of tumours in distant organs—underlies cancer mortality. Breast-to-brain metastasis (B2BM) is a common and disruptive form of cancer and is prevalent in the aggressive basal-like subtype, but is also found at varying frequencies in all cancer subtypes. Previous studies revealed parameters of breast cancer metastasis to the brain, but its preference for this site remains an enigma. Here we show that B2BM cells co-opt a neuronal signalling pathway that was recently implicated in invasive tumour growth, involving activation by glutamate ligands of N-methyl-d-aspartate receptors (NMDARs), which is key in model systems for metastatic colonization of the brain and is associated with poor prognosis. Whereas NMDAR activation is autocrine in some primary tumour types, human and mouse B2BM cells express receptors but secrete insufficient glutamate to induce signalling, which is instead achieved by the formation of pseudo-tripartite synapses between cancer cells and glutamatergic neurons, presenting a rationale for brain metastasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All materials are readily available from the corresponding author upon request or from standard commercial sources, including original IVIS data, Confocal, STED and electron microscopy images, immunohistochemical staining of human or mouse samples and full western blots. There are no restrictions on availability of the materials used in the study.
References
Lambert, A. W., Pattabiraman, D. R. & Weinberg, R. A. Emerging biological principles of metastasis. Cell 168, 670–691 (2017).
Vanharanta, S. & Massagué, J. Origins of metastatic traits. Cancer Cell 24, 410–421 (2013).
Lin, N. U., Amiri-Kordestani, L., Palmieri, D., Liewehr, D. J. & Steeg, P. S. CNS metastases in breast cancer: old challenge, new frontiers. Clin. Cancer Res. 19, 6404–6418 (2013).
Bos, P. D. et al. Genes that mediate breast cancer metastasis to the brain. Nature 459, 1005–1009 (2009).
Sevenich, L. et al. Analysis of tumour- and stroma-supplied proteolytic networks reveals a brain-metastasis-promoting role for cathepsin S. Nat. Cell Biol. 16, 876–888 (2014).
Valiente, M. et al. Serpins promote cancer cell survival and vascular co-option in brain metastasis. Cell 156, 1002–1016 (2014).
Chen, Q. et al. Carcinoma-astrocyte gap junctions promote brain metastasis by cGAMP transfer. Nature 533, 493–498 (2016).
Michael, I. P. et al. ALK7 signaling manifests a homeostatic tissue barrier that is abrogated during tumorigenesis and metastasis. Dev. Cell 49, 409–424 (2019).
Park, E. S. et al. Cross-species hybridization of microarrays for studying tumor transcriptome of brain metastasis. Proc. Natl Acad. Sci. USA 108, 17456–17461 (2011).
Neman, J. et al. Human breast cancer metastases to the brain display GABAergic properties in the neural niche. Proc. Natl Acad. Sci. USA 111, 984–989 (2014).
Li, L. & Hanahan, D. Hijacking the neuronal NMDAR signaling circuit to promote tumor growth and invasion. Cell 153, 86–100 (2013).
Li, L. et al. GKAP acts as a genetic modulator of NMDAR signaling to govern invasive tumor growth. Cancer Cell 33, 736–751 (2018).
Robinson, H. P. C. & Li, L. Autocrine, paracrine and necrotic NMDA receptor signalling in mouse pancreatic neuroendocrine tumour cells. Open Biol. 7, 170221 (2017).
Roche, K. W. et al. Molecular determinants of NMDA receptor internalization. Nat. Neurosci. 4, 794–802 (2001).
Takasu, M. A., Dalva, M. B., Zigmond, R. E. & Greenberg, M. E. Modulation of NMDA receptor-dependent calcium influx and gene expression through EphB receptors. Science 295, 491–495 (2002).
Lavezzari, G., McCallum, J., Lee, R. & Roche, K. W. Differential binding of the AP-2 adaptor complex and PSD-95 to the C-terminus of the NMDA receptor subunit NR2B regulates surface expression. Neuropharmacology 45, 729–737 (2003).
Nakazawa, T. et al. NR2B tyrosine phosphorylation modulates fear learning as well as amygdaloid synaptic plasticity. EMBO J. 25, 2867–2877 (2006).
Matsumura, S. et al. Impairment of CaMKII activation and attenuation of neuropathic pain in mice lacking NR2B phosphorylated at Tyr1472. Eur. J. Neurosci. 32, 798–810 (2010).
Knox, R. et al. NR2B phosphorylation at tyrosine 1472 contributes to brain injury in a rodent model of neonatal hypoxia-ischemia. Stroke 45, 3040–3047 (2014).
Levy, A. D. et al. Noonan syndrome-associated SHP2 dephosphorylates GluN2B to regulate NMDA receptor function. Cell Rep. 24, 1523–1535 (2018).
Ciriello, G. et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell 163, 506–519 (2015).
Fonnum, F., Storm-Mathisen, J. & Divac, I. Biochemical evidence for glutamate as neurotransmitter in corticostriatal and corticothalamic fibres in rat brain. Neuroscience 6, 863–873 (1981).
Briggs, K. J. et al. Paracrine induction of HIF by glutamate in breast cancer: EglN1 senses cysteine. Cell 166, 126–139 (2016).
Takano, T. et al. Glutamate release promotes growth of malignant gliomas. Nat. Med. 7, 1010–1015 (2001).
Buckingham, S. C. et al. Glutamate release by primary brain tumors induces epileptic activity. Nat. Med. 17, 1269–1274 (2011).
Danbolt, N. C. Glutamate uptake. Prog. Neurobiol. 65, 1–105 (2001).
Scheiffele, P., Fan, J., Choih, J., Fetter, R. & Serafini, T. Neuroligin expressed in nonneuronal cells triggers presynaptic development in contacting axons. Cell 101, 657–669 (2000).
Fu, Z., Washbourne, P., Ortinski, P. & Vicini, S. Functional excitatory synapses in HEK293 cells expressing neuroligin and glutamate receptors. J. Neurophysiol. 90, 3950–3957 (2003).
Stogsdill, J. A. et al. Astrocytic neuroligins control astrocyte morphogenesis and synaptogenesis. Nature 551, 192–197 (2017).
Harris, K. M. & Weinberg, R. J. Ultrastructure of synapses in the mammalian brain. Cold Spring Harb. Perspect. Biol. 4, a005587 (2012).
Pacifici, M. & Peruzzi, F. Isolation and culture of rat embryonic neural cells: a quick protocol. J. Vis. Exp. 63, e3965 (2012).
Fellmann, C. et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 5, 1704–1713 (2013).
Lorger, M. & Felding-Habermann, B. Capturing changes in the brain microenvironment during initial steps of breast cancer brain metastasis. Am. J. Pathol. 176, 2958–2971 (2010).
Amit, M., Na’ara, S. & Gil, Z. Mechanisms of cancer dissemination along nerves. Nat. Rev. Cancer 16, 399–408 (2016).
Ikonomidou, C. & Turski, L. Why did NMDA receptor antagonists fail clinical trials for stroke and traumatic brain injury? Lancet Neurol. 1, 383–386 (2002).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Hatzis, C. et al. A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. J. Am. Med. Assoc. 305, 1873–1881 (2011).
Weilinger, N. L. et al. Metabotropic NMDA receptor signaling couples Src family kinases to pannexin-1 during excitotoxicity. Nat. Neurosci. 19, 432–442 (2016).
Cardona, A. et al. TrakEM2 software for neural circuit reconstruction. PLoS One 7, e38011 (2012).
Fellmann, C. et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 5, 1704–1713 (2013).
Vargas-Caballero, M. & Robinson, H. P. C. Fast and slow voltage-dependent dynamics of magnesium block in the NMDA receptor: the asymmetric trapping block model. J. Neurosci. 24, 6171–6180 (2004).
Kim, N.-K. & Robinson, H. P. C. Effects of divalent cations on slow unblock of native NMDA receptors in mouse neocortical pyramidal neurons. Eur. J. Neurosci. 34, 199–212 (2011).
Shchors, K., Massaras, A. & Hanahan, D. Dual targeting of the autophagic regulatory circuitry in gliomas with repurposed drugs elicits cell-lethal autophagy and therapeutic benefit. Cancer Cell 28, 456–471 (2015).
Acknowledgements
We thank: J. Massagué and J. Joyce for, respectively, providing the MDA231 and TS1 parental and derivative B2BM cell lines; J. Massagué, K. Pietras and M. De Palma for comments on the manuscript; S. Grant for suggestions; M. W. Peng, A. F. Dubois, S. Lamy and L. Drori for technical support; J. Scotton for providing mouse glioma samples; W. Tang and all the members of the Hanahan laboratory for discussions. This work was principally supported by grants from the Swiss National Science Foundation and the European Research Council, and by a gift from the Biltema Foundation that was administered by the ISREC Foundation, Lausanne, Switzerland.
Author information
Authors and Affiliations
Contributions
Q.Z., G.C. and D.H. conceptualized the project and designed the experiments; I.P.M. contributed to the experimental design. G.C., S.S. and P.Z. performed the bioinformatic analysis; I.P.M. and Q.Z. generated the knockdown and rescue approach. J.A.G. and I.Z. did the immunohistochemical staining on human breast cancer and brain metastasis samples; H.P.C.R. carried out electrophysiology and calcium imaging. J.W., Q.Z. and B.D.M. performed the STED analysis of mouse brain with brain metastases; G.K. did the electron microscopy analysis; Q.Z. performed in vivo, in vitro and primary culture experiments, data analyses and quantification; Q.Z., G.C., I.P.M. and D.H. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Andres Barria, Adrienne Boire, Brunhilde Felding and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Association of the NMDAR signalling pathway with human breast cancer.
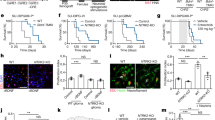
a, mRNA expression of six NMDAR-associated genes (GRIN2A, GRIN2B, GRIN2C, GRIN2D, GRIN1, and DLGAP1) in 1,100 patients with breast cancer from The Cancer Genome Atlas (TCGA). Samples were sorted by NMDAR score (bottom), which combines, for each sample, the Z-scores of normalized expression values of all six genes. b, mRNA expression of four AMPAR-associated genes (GRIA1, GRIA2, GRIA4, and GRIA3) in 1,100 patients with breast cancer from TCGA. Samples were sorted by AMPAR score (bottom), which combines, for each sample, the Z-scores of normalized expression values of all four genes. c, mRNA expression of 5 kainate-associated genes (GRIK2, GRIK4, GRIK3, GRIK1, and GRIK5) in 1,100 patients with breast cancer from TCGA. Samples were sorted by kainate score (bottom), which combines, for each sample, the Z-scores of normalized expression values of all five genes. d, mRNA expression of eight metabotropic glutamate-associated genes (GRM8, GRM1, GRM3, GRM4, GRM7, GRM2, GRM6, and GRM5) in 1,100 patients with breast cancer from TCGA. Samples were sorted by metabotropic score (bottom), which combines, for each sample, the Z-scores of normalized expression values of all eight genes. e, Box plot comparison of mRNA expression of GRIN2A, GRIN2B, GRIN2C, GRIN2D, DLGAP1, and GRIN1 (from left to right) across the five breast cancer PAM50 subtypes. P values were computed using a two-sided ANOVA test. f, Comparison between NMDAR score (x axis) and a previously derived NMDAR expression signature (y axis) across all breast cancer samples. Points are colour coded based on points concentration (warm colour = high concentration, cold colours = low concentration), n = 1,100. g, Box plot comparison of scores from the previously derived NMDAR expression signature across the five PAM50 breast cancer subtypes. h, Kaplan–Meier plots comparing distant-relapse free survival between samples with high GRIN2B expression (top 25%, red line) versus remaining cases (blue line) in luminal B, HER2-enriched, luminal B, normal-like and basal-like breast cancer subtypes (from left to right). The log-rank tests are two sided. In all comparisons, the number of samples for each breast cancer subtype (a–e, g) are: luminal A (n = 467), luminal B (n = 198), HER2-enriched (n = 72), normal-like (n = 28) and basal-like (n = 155). The thick central line of each box plot represents the median number of significant motifs, the bounding box corresponds to the 25th to 75th percentiles, and the whiskers extend up to 1.5 times the interquartile range. These results are based upon data generated by the TCGA Research Network (https://www.cancer.gov/tcga)21.
Extended Data Fig. 2 GluN2B–NMDAR signalling is activated in brain metastasis.
a, Representative haematoxylin and eosin-stained images of two paired human primary breast cancers (primary) and brain metastases (brain metastases) are shown. Scale bar, 100 µm. Two independent experiments. b, Quantification of pGluN2B(Y1472) staining in 16 primary breast tumours and 18 brain metastases, including 9 matched primary cancers and brain metastases, along with 7 unmatched primary tumours and 9 unmatched brain metastases. Mean ± s.e.m. Mann–Whitney test (two-tailed) was used. Subtype categorization for those patients are: 4 TNBC, 6 HER2+, 2 luminal and 13 unknown. c–e, Representative images (c) and quantification (d) of immunohistochemical staining of pGluN2B(Y1252) in 12 paired human primary breast cancers (primary) and brain metastases (brain metastases). Mean ± s.e.m. Wilcoxon test analysis (two-tailed). Subtype categorization for these twelve patients: 1 TNBC, 4 HER2+, 2 luminal and 5 unknown. e, Quantification of pGluN2B(Y1252) staining in 16 primary breast tumours and 22 brain metastases, including 12 matched primary cancers and brain metastases, along with 4 unmatched primary tumours and 8 unmatched brain metastases. Mean ± s.e.m. The Mann–Whitney (two-tailed) statistical test was used. Subtype categorization for those patients: 4 TNBC, 6 HER2+, 2 luminal and 13 unknown. Scale bar, 100 µm. f–h, Immunohistochemical staining of GluN2B in 16 human primary breast cancers and 23 brain metastases: representative images (f), Wilcoxon test analysis (two-tailed) of the 15 matched primary cancers and brain metastases (g), and Mann–Whitney (two-tailed) test analysis (h) of all samples. Mean ± s.e.m. Note that it remains to be ascertained whether the few brain metastases lacking GluN2B expression have activated NMDAR signalling via another GluN2 subunit. Scale bar, 100 μm. Subtype categorization for those patients: 4 TNBC, 6 HER2+, 2 luminal and 13 unknown. i–k, Immunohistochemical staining of GluN1 in 16 human primary breast cancers and 16 brain metastases: representative images (i), Wilcoxon test (two-tailed) analysis of the 11 matched primary cancers and brain metastases (j) and Mann–Whitney test (two-tailed) analysis (k) of all samples. Mean ± s.e.m. Scale bar, 100 μm. Note that owing to variations in the presence of cancer in the sections taken from the tissue microarray block for analysis, not all 25 samples were represented in each of the immunostainings performed in Fig. 2 and in this figure. Subtype categorization for those patients: 4 TNBC, 6 HER2+, 2 luminal and 13 unknown. l, Representative immunohistochemical staining of pGluN2B(Y1472), total GluN2B and luciferase in primary tumours, brain metastases, and lung metastases formed by MDA231-BrM cells (day 28–35 after injection). Scale bar, 100 µm. The areas indicated by black boxes are shown in Fig. 2e. m, n, Immunohistochemical staining of pGluN2B(Y1252), total GluN2B, and luciferase (m) and quantification of pGluN2B(Y1252) staining (n) in primary tumours, brain metastases, and lung metastases formed by MDA231-BrM cells (day 28–35 after injection). Scale bar, 100 µm. Insets are magnified 3.2× relative to the main field. Two-tailed Student’s t-test, mean ± s.e.m., n = 3 mice per group.
Extended Data Fig. 3 NMDAR-mediated calcium signalling and single channel currents in B2BM cells.
To test for functional NMDARs, fluorescence imaging and patch-clamp recordings were carried out in 2D cultures of TS1-BrM cells, 14 fields of view (301 × 301 μm) from 10 different culture dishes. Total of 151 responding cells out of 1,070 imaged cells (14.11%). Data from bath addition of 50–100 μl of 100 mM potassium glutamate, 50–200 μl of 10 mM NMDA, local perfusion-pipette puffing of 100 μM potassium glutamate or 500 μM NMDA, all of which produced calcium responses. a–c, Representative calcium imaging experiments. a, TS1-BrM cells were loaded with Oregon Green BAPTA-1 AM fluorescent calcium indicator, and imaged with a sCMOS camera at 5 frames per second. A subset of cells in the field of view (13/75) showed calcium elevation in response to the addition of glutamate to the bath (50 μl of 10 mM glutamate, final concentration 125 μM). b, Calcium signals from four cells as indicated in a; the arrow indicates glutamate application. c, Average signal over the 13 agonist-responsive cells during application of glutamate (blue trace) or preceding control application of the same volume of Ringer solution (red trace). d, Response to application of 50 μl of 10 mM NMDA (final concentration 125 μM), averaged over 6 responding cells in a field of view containing 30 cells (blue trace), compared to an example response averaged over 52 cells to the same amount of NMDA, but in the presence of 100 μM APV (2-amino-5-phosphonopentanoic acid; a selective NMDA receptor antagonist) and 2 mM Mg2+ (red trace). Overall, calcium transients stimulated either by NMDA or glutamate were detectable in about 14% of cells (151 of 1,070 cells from 10 culture dishes). e–g, Puff application (pipette at left) of NMDA (200 μm, grey bar) activates inward single-channel currents recorded through the pipette at right (e, f), with amplitudes of around 4.5 pA at a holding potential of −90 mV (g), equivalent to a chord conductance of 50 pS, characteristic of GluN2A- or GluN2B-containing NMDARs. Single-channel currents of 4.5 pA were detectable in low-noise whole-cell recordings on the application of glutamate (100 μM) or NMDA (200 μM) in 45% (9/20) of recorded cells. h, Resting membrane potentials were measured by whole-cell current-clamp using pipettes filled with cytoplasm-like high-potassium solutions (box plot, −52 ± 10.6 mV (mean ± s.e.m., n = 9 cells). For comparison, the voltage dependence of GluN2A and GluN2B inward current in physiological (1 mM) magnesium is superimposed41,42, indicating that membrane potentials are sufficiently depolarized to overcome a substantial fraction of voltage-dependent magnesium block of NMDARs.
Extended Data Fig. 4 Brain metastases are not inducing the neuronal apoptosis anticipated for autocrine secretion of glutamate by cancer cells.
a, Western blot analysis of the xCT and vGlut2 glutamate transporters in pairs of human and mouse parental breast cell lines and their corresponding brain metastatic derivatives. Protein levels in B2BM cells are quantified relative to cognate parental cells, after normalization to GAPDH. Three independent experiments. GAPDH was run on separate gels from xCT and vGlut2 as sample processing controls. b, Immunohistochemical-(DAB)-based staining to detect apoptotic cells in mouse brains with metastases formed by MDA231-BrM cells (day 28–35 after injection). DNAase I treatment on brain tissue sections was used as a TUNEL+ control. Images shown are representative of an analysis of more than 20 brain metastases from 4 mouse brains, 2 sections per mouse brain. Scale bar, 100 µm. c, Immunofluorescent staining for cleaved-caspase 3 (green), luciferase (cancer cells; magenta) and NeuN (neurons; red) in mouse brain with metastases formed by MDA231-BrM cells (day 28–35 after injection). Images shown are representative of an analysis of >20 brain metastases from 4 mouse brains, 2 sections/mouse brain. Scale bar, 100 µm. d, Immunofluorescence staining for TUNEL (green), NeuN (red), GFAP (glioma cells; magenta) in a brain harbouring a glioma that arose in a genetically engineered mouse model43. Images shown are representative of an analysis of more than six gliomas from six mouse brains, one section per mouse brain. As previously reported24, glioma cells secrete glutamate—which causes neuronal apoptosis—in contrast to B2BM cells, which do not. Scale bar, 50 µm.
Extended Data Fig. 5 Interactions between B2BM cells and neurons.
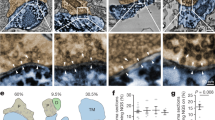
a–c, Immunofluorescence staining of luciferase (B2BM; blue), vGlut2 (red) and pGluN2B(Y1252) (green) in a mouse brain metastasis formed by MDA231-BrM cells (day 28–35 after injection), imaged by STED super-resolution microscopy. a, b, Tilted 3D images of the area circumscribed by the solid white-bordered box in Fig. 4a. Arrow, vGlut2+ in presynaptic neuron; arrowhead, pGluN2B(Y1252)+ in a blue-labelled B2BM cell, potentially in close apposition to a postsynaptic neuron. c, Tilted 3D image of the area circumscribed by the dotted white-bordered box in Fig. 4a, depicting a normal synapse revealed by close association between vGlut2+ puncta in a presynaptic neuron (arrow) with pGluN2B+ puncta in a postsynaptic neuron (arrowhead). Side length of each square in 3D view, 400 nm (a, b) and 300 nm (c). Images are representative of an analysis of nine brain metastases from three mouse brains, two sections per mouse brain. See Supplementary Videos further describing the images shown in a–c. d, e, Immunofluorescence staining of luciferase (green) and synaptobrevin 1 (red) in a mouse brain metastasis formed by MDA231-BrM cells (day 28–35 after injection), imaged by STED super-resolution microscopy. d, A merged image (left) and individual fluorescent images (middle and right). Scale bar, 10 µm. e, A tilted 3D image at two magnifications, highlighting a synaptobrevin 1+ bouton chain (red) localized proximal to luciferase+ tumour cells (green). Images are representative of an analysis of nine brain metastases from three mouse brains, two sections per mouse brain. Side length for each square in 3D view, 10 µm. f, g, Immunofluorescence staining for postsynaptic proteins PSD-95 (red, f), and neuroligin 2 (red, g) along with luciferase (green) in mouse brains with metastases formed by MDA231-BrM cells (days 28–35 after injection; scale bar, 10 µm). The area circumscribed by the white box is shown at higher magnification (left; scale bar, 1 µm). Images are representative of an analysis of more than 20 brain metastases from 4 mouse brains, 2 sections per mouse brain.
Extended Data Fig. 6 NMDAR signalling in B2BM cells is activated in co-culture with neurons.
a, Schematic of cancer cell–primary neuron co-culture system and experimental design. b, Immunofluorescence staining for MAP2 (neuronal marker, green) and GFAP (astrocyte marker, red) in primary cultures of cortical neurons, revealing the preponderance of neurons. The area circumscribed by the white box is shown at higher magnification (left). Scale bars, 1 µm (left), 10 µm (right). Three independent experiments. c, Representative bright-field (BF) and fluorescent (mKate+) images (left) and quantification (right) of MDA231 parental and MDA231-BrM cells co-cultured with primary cortical neurons for 14 days. Two-tailed Student’s t-test, mean ± s.e.m., n = 3 biological replicates over three independent experiments. Scale bar, 100 µm. d, Representative bright-field and fluorescent images (left) and quantification (right) of TS1 parental and TS1-BrM cells co-cultured with primary cortical neurons for 14 days. Data are mean ± s.e.m. Student’s t-test (two-tailed) was used. n = 3 biological replicates over three independent experiments. e–g, Quantification of MDA231 parental and MDA231-BrM cell numbers after 14 days of culture in poly-d-lysine-coated plates in three different conditions: in complete neuronal culture medium (e); in conditioned medium from primary cortical neuron cultures (f); and in a Boyden chamber with neurons in the top chamber and cancer cells in the bottom chamber (g). Mean ± s.e.m. Student’s t-test (two-tailed) was used. n = 3 biological replicates over three independent experiments. The panels below each bar graph show representative images of the cancer cells at the end of the assay, revealed by mKate fluorescence. h, Schematic inducible miR-E-based shRNA knock-down system carrying three distinct miR-E-based shRNA sequences that bind to different regions of a targeted mRNA (top vector), and tet-on inducible vector for GRIN2B open reading frame (ORF) used for rescue experiments (bottom vector). TRE, tet-on inducible promoter; rtTA, reverse tetracycline transactivator. i, Knockdown of GRIN2B in cultured MDA231-BrM cells with tet-on inducible shRNAs, as assessed by western blotting after DOX treatment (1 µg ml−1) for two days. Three independent experiments. The numbers above the blots indicate levels of pGluN2B(Y1252) and total GluN2B protein normalized to GAPDH. GAPDH was run in the same gel as pGluN2B(Y1252), and run on separate gels from GluN2B as sample processing controls. j, Representative bright-field and fluorescent images (bottom) and quantification (top) of MDA231-BrM cells transfected with inducible shCtrl and shGRIN2B (1 µg ml−1), co-cultured with primary cortical neurons for 14 days. Two-tailed Student’s t-test; mean ± s.e.m., n = 3 biological replicates over three independent experiments. Scale bar, 100 µm. k, Rescue expression of GluN2B in cultured MDA231-BrM cells with tet-on inducible shRNAs along with a GRIN2B cDNA, as assessed by western blotting after DOX treatment (1 µg ml−1) for two days. Three independent experiments. GAPDH was run in the same gel as GluN2B. l, Representative fluorescent (mKate+) images (left), and quantification (right) of MDA231-BrM shCtrl, shGRIN2B and GluN2B-rescued cells co-cultured with primary cortical neurons for 14 days. Two-tailed Student’s t-test; mean ± s.e.m., n = 3 biological replicates over three independent experiments. Scale bar, 100 µm. m, Cell proliferation in shCTRL and shGRIN2B MDA231-BrM cells as determined by MTT assays, starting with 5,000 or 10,000 cells per well, after 72 h in culture. Two-way ANOVA; mean ± s.e.m., n = 3 independent experiments.
Extended Data Fig. 7 In vivo assessment of the functional importance of GluN2B–NMDAR signalling in B2BM cells.
a, b, Representative immunofluorescent images of FMRP (green), luciferase (red) and DAPI (blue) in brain metastases formed by shCtrl (control shRNA) and shGRIN2B MDA231 cells (a), and quantification of mean FMRP fluorescence in Luciferase+ tumour-cell clusters (b). Two-tailed Student’s t-test, mean ± s.e.m.; n = 25 for shCtrl and n = 32 shGRIN2B group, all from 3 mice per group. c, Weight of orthotopic breast tumours formed by MDA231-BrM cells transfected with inducible shCtrl or shGRIN2B inoculated into the fourth mammary fat pads of female mice. The mice were fed food containing DOX to induce the shRNAs concomitantly with mammary fat pad injection. Two-tailed Student’s t-test, mean ± s.e.m.; n = 9 mice for the shCtrl group, and 10 mice for shGRIN2B group. d, Bioluminescence imaging (BLI) and quantification of lung metastatic lesions formed by MDA231-BrM cells transfected with inducible shCtrl or shGRIN2B inoculated intravenously. DOX food was added to induce the shRNAs concomitantly with intravenous injection. Two-tailed Student’s t-test; mean ± s.e.m., n = 5 mice per group, two independent experiments. e, BLI and quantification of brain metastatic lesions formed by MDA231-BrM cells transfected with inducible shCtrl or shGRIN2B, or additionally with rescue expression of a GRIN2B cDNA. DOX food was supplied concomitantly with ICD injection to induce shRNA expression. Tumour burden was assessed four weeks later by BLI. Two independent experiments. Two-tailed Student’s t-test, mean ± s.e.m., n = 10 mice for shCtrl group, n = 9 mice for shGRIN2B group and n = 9 mice for the rescue group.
Extended Data Fig. 8 Functional analysis of GluN2B expression in B2BM cells.
a, Knockdown of GluN2B expression in cultured mouse TS1-BrM cells with tet-on inducible shRNAs, as assessed by western blotting two days after DOX treatment in vitro (1 µg ml−1). Three independent experiments. The numbers above the blot indicate levels of GluN2B normalized to GAPDH. GAPDH was run in the same gel as GluN2B. b, Weight of primary orthotopic breast tumours formed by TS1-BrM cells transfected with inducible shCtrl or shGrin2b, and inoculated into the fourth mammary fat pad of female mice. Mice were fed with food containing DOX to induce the shRNAs concomitantly with mammary fat pad injection. Two-tailed Student’s t-test; mean ± s.e.m., n = 6 mice for shCtrl group, and n = 8 mice for shGrin2b group. c–e, In vivo BLI (c) and quantification of brain (d) and thoracic cavity (e) photon flux in mice bearing TS-BrM cells transfected with inducible shCtrl or shGrin2b four weeks after intracardiac injection. DOX food was supplied concomitantly with ICD injection to induce shRNA expression. Two-tailed Student’s t-test; mean ± s.e.m., n = 9 mice per group, two independent experiments. f, Ex vivo BLI and quantification of excised brains with metastatic lesions formed by TS1-BrM cells transfected with inducible shCtrl or shGrin2b. Mice were fed food containing DOX concomitantly with ICD injection to induce shRNA expression. Two-tailed Student’s t-test; mean ± s.e.m., n = 9 mice per group, two independent experiments.
Supplementary information
Supplementary Figure 1
Raw images of western blots.
Supplementary Table 1
Patient information for the breast cancer TMA.
Supplementary Video 1
Pseudo-tripartite synapse in a brain metastasis lesion. Blue = B2BM cancer cell, revealed by luciferase+; red = presynaptic neuron, revealed by vGlut2+ immunostaining; green = pGluN2B 1252+ in B2BM and presumptive neuronal processes in close proximity to the B2BM cancer cells.
Supplementary Video 2
A normal neuronal synapse. Red = presynaptic neuron, revealed by vGlut2+ immunostaining; green = postsynaptic neuron, revealed by pGluN2B 1252+.
Rights and permissions
About this article
Cite this article
Zeng, Q., Michael, I.P., Zhang, P. et al. Synaptic proximity enables NMDAR signalling to promote brain metastasis. Nature 573, 526–531 (2019). https://doi.org/10.1038/s41586-019-1576-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1576-6
This article is cited by
-
A phase Ib/II randomized, open-label drug repurposing trial of glutamate signaling inhibitors in combination with chemoradiotherapy in patients with newly diagnosed glioblastoma: the GLUGLIO trial protocol
BMC Cancer (2024)
-
Leveraging next-generation materials for cancer neuroscience therapies in the central nervous system
Nature Reviews Materials (2024)
-
How cancer hijacks the nervous system to grow and spread
Nature (2024)
-
Cell-intrinsic and microenvironmental determinants of metastatic colonization
Nature Cell Biology (2024)
-
Inflammatory Factor IL1α Induces Aberrant Astrocyte Proliferation in Spinal Cord Injury Through the Grin2c/Ca2+/CaMK2b Pathway
Neuroscience Bulletin (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.