Abstract
The molecular mechanisms of exon definition and back-splicing are fundamental unanswered questions in pre-mRNA splicing. Here we report cryo-electron microscopy structures of the yeast spliceosomal E complex assembled on introns, providing a view of the earliest event in the splicing cycle that commits pre-mRNAs to splicing. The E complex architecture suggests that the same spliceosome can assemble across an exon, and that it either remodels to span an intron for canonical linear splicing (typically on short exons) or catalyses back-splicing to generate circular RNA (on long exons). The model is supported by our experiments, which show that an E complex assembled on the middle exon of yeast EFM5 or HMRA1 can be chased into circular RNA when the exon is sufficiently long. This simple model unifies intron definition, exon definition, and back-splicing through the same spliceosome in all eukaryotes and should inspire experiments in many other systems to understand the mechanism and regulation of these processes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Zhang, L., Vielle, A., Espinosa, S. & Zhao, R. RNAs in the spliceosome: insight from cryoEM structures. Wiley Interdiscip. Rev. RNA 10, e1523 (2019).
Wan, R., Bai, R., Yan, C., Lei, J. & Shi, Y. Structures of the catalytically activated yeast spliceosome reveal the mechanism of branching. Cell 177, 339–351 (2019).
De Conti, L., Baralle, M. & Buratti, E. Exon and intron definition in pre-mRNA splicing. Wiley Interdiscip. Rev. RNA 4, 49–60 (2013).
Berget, S. M. Exon recognition in vertebrate splicing. J. Biol. Chem. 270, 2411–2414 (1995).
Sharma, S., Kohlstaedt, L. A., Damianov, A., Rio, D. C. & Black, D. L. Polypyrimidine tract binding protein controls the transition from exon definition to an intron defined spliceosome. Nat. Struct. Mol. Biol. 15, 183–191 (2008).
Schneider, M. et al. Exon definition complexes contain the tri-snRNP and can be directly converted into B-like precatalytic splicing complexes. Mol. Cell 38, 223–235 (2010).
Wang, P. L. et al. Circular RNA is expressed across the eukaryotic tree of life. PLoS One 9, e90859 (2014).
Wilusz, J. E. A. A 360° view of circular RNAs: from biogenesis to functions. Wiley Interdiscip. Rev. RNA 9, e1478 (2018).
Starke, S. et al. Exon circularization requires canonical splice signals. Cell Rep. 10, 103–111 (2015).
Séraphin, B., Kretzner, L. & Rosbash, M. A U1 snRNA:pre-mRNA base pairing interaction is required early in yeast spliceosome assembly but does not uniquely define the 5′ cleavage site. EMBO J. 7, 2533–2538 (1988).
Siliciano, P. G. & Guthrie, C. 5′ splice site selection in yeast: genetic alterations in base-pairing with U1 reveal additional requirements. Genes Dev. 2, 1258–1267 (1988).
Ruby, S. W. & Abelson, J. An early hierarchic role of U1 small nuclear ribonucleoprotein in spliceosome assembly. Science 242, 1028–1035 (1988).
Abovich, N. & Rosbash, M. Cross-intron bridging interactions in the yeast commitment complex are conserved in mammals. Cell 89, 403–412 (1997).
Plaschka, C., Lin, P. C., Charenton, C. & Nagai, K. Prespliceosome structure provides insights into spliceosome assembly and regulation. Nature 559, 419–422 (2018).
Bai, R., Wan, R., Yan, C., Lei, J. & Shi, Y. Structures of the fully assembled Saccharomyces cerevisiae spliceosome before activation. Science 360, 1423–1429 (2018).
Lewis, J. D., Izaurralde, E., Jarmolowski, A., McGuigan, C. & Mattaj, I. W. A nuclear cap-binding complex facilitates association of U1 snRNP with the cap-proximal 5′ splice site. Genes Dev. 10, 1683–1698 (1996).
Qiu, Z. R., Chico, L., Chang, J., Shuman, S. & Schwer, B. Genetic interactions of hypomorphic mutations in the m7G cap-binding pocket of yeast nuclear cap binding complex: an essential role for Cbc2 in meiosis via splicing of MER3 pre-mRNA. RNA 18, 1996–2011 (2012).
Puig, O., Gottschalk, A., Fabrizio, P. & Séraphin, B. Interaction of the U1 snRNP with nonconserved intronic sequences affects 5′ splice site selection. Genes Dev. 13, 569–580 (1999).
Lesser, C. F. & Guthrie, C. Mutational analysis of pre-mRNA splicing in Saccharomyces cerevisiae using a sensitive new reporter gene, CUP1. Genetics 133, 851–863 (1993).
Liu, S. et al. Structure of the yeast spliceosomal postcatalytic P complex. Science 358, 1278–1283 (2017).
Lu, M. et al. Crystal structure of the three tandem FF domains of the transcription elongation regulator CA150. J. Mol. Biol. 393, 397–408 (2009).
Liu, J., Fan, S., Lee, C. J., Greenleaf, A. L. & Zhou, P. Specific interaction of the transcription elongation regulator TCERG1 with RNA polymerase II requires simultaneous phosphorylation at Ser2, Ser5, and Ser7 within the carboxyl-terminal domain repeat. J. Biol. Chem. 288, 10890–10901 (2013).
Li, X. et al. CryoEM structure of Saccharomyces cerevisiae U1 snRNP offers insight into alternative splicing. Nat. Commun. 8, 1035 (2017).
Görnemann, J. et al. Cotranscriptional spliceosome assembly and splicing are independent of the Prp40p WW domain. RNA 17, 2119–2129 (2011).
Ester, C. & Uetz, P. The FF domains of yeast U1 snRNP protein Prp40 mediate interactions with Luc7 and Snu71. BMC Biochem. 9, 29 (2008).
Wiesner, S., Stier, G., Sattler, M. & Macias, M. J. Solution structure and ligand recognition of the WW domain pair of the yeast splicing factor Prp40. J. Mol. Biol. 324, 807–822 (2002).
Jacewicz, A., Chico, L., Smith, P., Schwer, B. & Shuman, S. Structural basis for recognition of intron branchpoint RNA by yeast Msl5 and selective effects of interfacial mutations on splicing of yeast pre-mRNAs. RNA 21, 401–414 (2015).
Kappel, K. & Das, R. Sampling native-like structures of RNA-protein complexes through Rosetta folding and docking. Structure 27, 140–151.e145 (2019).
Howe, K. J., Kane, C. M. & Ares, M., Jr. Perturbation of transcription elongation influences the fidelity of internal exon inclusion in Saccharomyces cerevisiae. RNA 9, 993–1006 (2003).
Campodonico, E. & Schwer, B. ATP-dependent remodeling of the spliceosome: intragenic suppressors of release-defective mutants of Saccharomyces cerevisiae Prp22. Genetics 160, 407–415 (2002).
Liang, D. et al. The output of protein-coding genes shifts to circular RNAs when the pre-mRNA processing machinery is limiting. Mol. Cell. 68, 940–954.e943 (2017).
Ragan, C., Goodall, G. J., Shirokikh, N. E. & Preiss, T. Insights into the biogenesis and potential functions of exonic circular RNA. Sci. Rep. 9, 2048 (2019).
Liang, D. & Wilusz, J. E. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 28, 2233–2247 (2014).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013).
Mokry, M. et al. Accurate SNP and mutation detection by targeted custom microarray-based genomic enrichment of short-fragment sequencing libraries. Nucleic Acids Res. 38, e116 (2010).
Spingola, M., Grate, L., Haussler, D. & Ares, M., Jr. Genome-wide bioinformatic and molecular analysis of introns in Saccharomyces cerevisiae. RNA 5, 221–234 (1999).
Li, X. et al. Comprehensive in vivo RNA-binding site analyses reveal a role of Prp8 in spliceosomal assembly. Nucleic Acids Res. 41, 3805–3818 (2013).
Abelson, J. et al. Conformational dynamics of single pre-mRNA molecules during in vitro splicing. Nat. Struct. Mol. Biol. 17, 504–512 (2010).
Carragher, B. et al. Leginon: an automated system for acquisition of images from vitreous ice specimens. J. Struct. Biol. 132, 33–45 (2000).
Zheng, S. Q., Palovcak, E., Armache, J.-P., Cheng, Y. & Agard, D. A. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Keating, K. S. & Pyle, A. M. RCrane: semi-automated RNA model building. Acta Crystallogr. D Biol. Crystallogr. 68, 985–995 (2012).
Chou, F. C., Sripakdeevong, P., Dibrov, S. M., Hermann, T. & Das, R. Correcting pervasive errors in RNA crystallography through enumerative structure prediction. Nat. Methods 10, 74–76 (2013).
Kappel, K. et al. De novo computational RNA modeling into cryo-EM maps of large ribonucleoprotein complexes. Nat. Methods 15, 947–954 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Wiśniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Grimm, M., Zimniak, T., Kahraman, A. & Herzog, F. xVis: a web server for the schematic visualization and interpretation of crosslink-derived spatial restraints. Nucleic Acids Res. 43, W362–W369 (2015).
Seraphin, B. & Rosbash, M. Identification of functional U1 snRNA-pre-mRNA complexes committed to spliceosome assembly and splicing. Cell 59, 349–358 (1989).
Qin, D., Huang, L., Wlodaver, A., Andrade, J. & Staley, J. P. Sequencing of lariat termini in S. cerevisiae reveals 5′ splice sites, branch points, and novel splicing events. RNA 22, 237–253 (2016).
Li, Z. & Brow, D. A. A rapid assay for quantitative detection of specific RNAs. Nucleic Acids Res. 21, 4645–4646 (1993).
Kozlowski, L. P. & Bujnicki, J. M. MetaDisorder: a meta-server for the prediction of intrinsic disorder in proteins. BMC Bioinformatics 13, 111 (2012).
Acknowledgements
This work was supported by NIH grants GM126157 and GM130673 (R.Z.); GM071940 and AI094386 (Z.H.Z.); and GM122579, GM121487, and CA219847 (R.D.). S.E. is a Howard Hughes Medical Institute Gilliam Fellow. K.K. was supported by an NSF GRFP award and a Stanford Graduate Fellowship. We acknowledge the use of instruments at the Electron Imaging Center for Nanomachines (supported by UCLA and by grants from the NIH (1S10OD018111, 1U24GM116792) and NSF (DBI-1338135 and DMR-1548924)) as well as the CU Anschutz School of Medicine Cryo-EM and proteomics core facilities (partially supported by the School of Medicine and the University of Colorado Cancer Center Support Grant P30CA046934). Molecular graphics and analyses were performed with the UCSF Chimera and ChimeraX, developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, with support from NIGMS P41-GM103311 (Chimera, ChimeraX) and NIH R01-GM129325 (ChimeraX). We also thank M. Ares, D. Black, and D. Brow for comments on early versions of the manuscript.
Author information
Authors and Affiliations
Contributions
X.L. and S.L. contributed equally to the work and are listed alphabetically in the author list. R.Z. and Z.H.Z. conceived the project; X.L. prepared and optimized the sample; X.L., L.Z., S.E. and S.S. performed biochemical analyses; S.L. and Y.C. recorded and processed the EM data; A.I., R.C.H. and K.C.H. performed mass spectrometry analyses; S.L. built the atomic models; K.K. and R.D. built the partial U1 snRNA model and the minimal exon model in the A complex; R.Z., S.L., X.L., and Z.H.Z. analysed and interpreted the models; S.L., X.L. and R.Z. prepared the illustrations; R.Z., S.L. and Z.H.Z. wrote the paper; and all authors contributed to the editing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 In vitro assembly and purification of the ACT1 complex.
a, Schematic representation of the ACT1 pre-mRNA tagged with three MS2-binding sites (M3–ACT1) used for E complex assembly and purification. Boxes represent exon 1 (E1) and truncated exon 2 (E2). The 5′ SS (GU) and BPS (UACUAAC) are also shown. The red line represents the DNA oligo complementary to a region 5 nt upstream of the BPS for the RNase H cleavage experiment. b, RNA components of the assembled E complex (with or without DNA oligo and RNase H treatment) after proteinase K digestion are shown on a denaturing urea gel or native agarose gel. These results demonstrate that RNase treatment cleaved M3–ACT1 into two fragments. Note that the sizes of RNA on the native gel do not match their linear length, possibly owing to the existence of secondary structures. This experiment was repeated two additional times with similar results.
Extended Data Fig. 2 The cryo-EM structural determination process for the ACT1 complex.
a, Representative drift-corrected cryo-EM micrograph (out of 11,283 micrographs) of the E complex assembled on the ACT1 pre-mRNA. A representative particle is shown in a white dotted circle. b, Representative 2D class averages of the ACT1 complex obtained in RELION. This experiment was repeated one additional time with similar results. c, Data processing workflow. For processing above the red dashed line, the particle images were binned to a pixel size of 2.72 Å. The rest of the processing was performed with a pixel size of 1.36 Å. The masks used in data processing are outlined with red solid lines (see Methods). d, Angular distribution of all particles used for the final 3.2 Å map of the ACT1 complex. e, FSC as a function of spatial frequency demonstrating the resolution of the final reconstruction of the ACT1 complex. f, Resmap local resolution estimation. g, FSC coefficients as a functional of spatial frequency between model and cryo-EM density maps. The generally similar appearances between the FSC curves obtained with half maps with (red) and without (blue) model refinement indicate that the refinement of the atomic coordinates did not suffer from severe over-fitting.
Extended Data Fig. 3 The Cryo-EM structural determination process for the UBC4 complex.
a, Representative drift-corrected cryo-EM micrograph (out of 8,997 micrographs) of the E complex assembled on the UBC4 pre-mRNA. A representative particle is shown in a white dotted circle. b, Representative 2D class averages of the UBC4 complex obtained in RELION. c, Data processing workflow. For processing above the red dashed line, the particle images were binned to a pixel size of 2.72 Å. The rest of the processing was performed with a pixel size of 1.36 Å. The masks used in data processing are outlined with red solid lines (see Methods). d, Angular distribution of all particles used for the final 3.6 Å map of the UBC4 complex. e, FSC as a function of spatial frequency demonstrating the resolution of the final reconstruction of the UBC4 complex. f, Resmap local resolution estimation. g, FSC coefficients as a functional of spatial frequency between model and cryo-EM density maps. The generally similar appearances between the FSC curves obtained with half maps with (red) and without (blue) model refinement indicate that the refinement of the atomic coordinates did not suffer from severe over-fitting.
Extended Data Fig. 4 Representative cryo-EM density maps of the E complex.
a–i, Densities for the UBC4 complex; j, density for the ACT1 complex. Cryo-EM density maps are shown as follows. a, Selected regions of U1 snRNA. b, C-terminal region of Prp39. c, N-terminal domain of Snu71. d, Pre-mRNA and U1 snRNA duplex. e, U1C ZnF domain. f, Luc7 ZnF2 domain. g, Tandem FF domains of Prp40 (the known structure of tandem FF domains from CA150 is also shown with the characteristic boomerang shape). h, RRM2 domain of Nam8. i, NCBP1 and NCBP2. j, Weak density in the ACT1 complex that is assigned as the putative BBP–Mud2 heterodimer. The A complex is also shown, with U1 snRNP in the same orientation as the ACT1 complex and U2 snRNP located in similar positions as the BBP–Mud2 heterodimer with respect to U1 snRNP. The map of the ACT1 complex was low-pass filtered to 40 Å.
Extended Data Fig. 5 Structural and biochemical characterization of the ACT1 and UBC4 complexes.
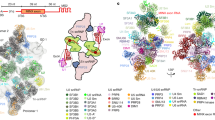
a, Comparison of the ribbon models of the ACT1 complex, the UBC4 complex, and U1 snRNP from other previously determined structures (the U1 snRNP, A, and pre-B complexes). Labels with shading indicate protein or RNA components that differ between the ACT1 and UBC4 complexes. These components and the RRM2 domain of Nam8 are also absent from previously determined structures. Note that U1–70K is shifted towards NCBP2 in the UBC4 complex. b, Purified E complex does not contain U2 snRNA. A native polyacrylamide gel shows the solution hybridization58 result of total cellular RNA or RNA from purified E complex hybridized with fluorescent probes specific for U1 and U2 snRNAs. This experiment was repeated one additional time with similar results.
Extended Data Fig. 6 Secondary structures in the region between the 5′ SS and BPS in the wild-type and mutant ACT1 and UBC4 pre-mRNAs.
a, Secondary structures predicted by RNAstructure 6.0 (https://rna.urmc.rochester.edu/RNAstructureWeb/). b, Sequence between the 5′ SS and BPS (underlined) of ACT1. Red nucleotides were mutated to A (other than the one A, which was mutated to G) in mutant ACT1 to disrupt predicted secondary structures.
Extended Data Fig. 7 Protein interactions in the UBC4 complex.
a, DSSO crosslinking and mass spectrometry analyses of the UBC4 complex. Each blue line indicates a crosslink between a pair of Lys residues. Note that BBP–Mud2 are crosslinked to Luc7, Prp40, Snu56, and Snu71. b, Co-purification assays probing the interaction between Snu71 (or Prp40) and Luc7. Various combinations of protein A–TEV–Prp40, protein A–TEV–Snu71, and CBP-tagged Luc7 or Luc7ΔCC (with coiled-coil domain (residues 123–190) deleted) were co-overexpressed in yeast (only Snu71 is protein A tagged in the Snu71 + Prp40 lanes), purified using IgG resin, eluted through TEV cleavage, analysed on SDS–PAGE, and visualized using western blot with an anti-CBP antibody to detect Luc7 (top) and Ponceau S stain to show Snu71 or Prp40 (middle). Western blot using the same anti-CBP antibody was used to demonstrate Luc7 expression levels in cell lysates (bottom). The faint band around 26 kD in all lanes of the middle gel is TEV. This experiment was repeated one additional time with similar results. c, The linker (residues 73–131) between the WW and FF domains of Prp40 is predicted to be disordered using program MetaDisorderMD259.
Extended Data Fig. 8 Computational, biochemical, and structural characterization of the EDC.
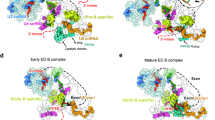
a, The minimal length of RNA needed to connect the upstream branch point (BP) and downstream 5′ SS in the A complex is modelled using the Rosetta RNP-denovo method. The A complex (PDB ID 6G90) is shown in grey. The pre-mRNA is shown in green. The upstream branch point and downstream 5′ SS are shown as purple space-filling models. Twenty-eight nucleotides are sufficient to connect the upstream branch point and downstream 5′ SS (not including the branch point and 5′ SS themselves) without any chain break or clashes. b, Schematics of wild-type and mutant DYN2 pre-mRNA (mutated nucleotides shown in red), IEI, and untagged IEI used for the EDC assembly and in vivo exon definition experiments. Stem-loops represent the MS2 binding sites, and the red line represents the DNA oligonucleotide used for RNase H cleavage. c, SDS–PAGE shows protein components of complexes assembled on wild-type and IEI substrates (lanes 1, 2), on wild-type in the presence of competing untagged IEI (lane 3), and on IEI after RNase H treatment in the absence and presence of the DNA oligo (lanes 4, 5). This experiment was repeated one additional time with similar results. d, RNA components of the same complexes as in lanes 4, 5 of c, confirming that RNase H treatment in the presence of the oligonucleotide cleaves the pre-mRNA. The smaller cleaved fragment (61 nucleotides) is difficult to see because EtBr stains short single-stranded RNA with low efficiency. This experiment was repeated two additional times with similar results. e, Mass spectrometry analyses of spliceosome assembled on the IEI and wild-type DYN2 pre-mRNA indicate that the two complexes have the same components in similar quantities with the exception of NCBP1 and 2, which are absent from the IEI complex. f, 2D classification of negative-stain TEM images of the E complex assembled on DYN2 IEI pre-mRNA. This experiment was repeated one additional time with similar results.
Extended Data Fig. 9 Characterization of circRNAs.
a, Sanger sequencing confirmed that the PCR products in Fig. 5a were derived from T-branches and circRNAs of EFM5 and HMRA1. Solidus, site where two ends of exon 2 are ligated; vertical line, site where the 5′ SS of intron 2 is ligated to the BP of intron 1. The 5′ SS and BPS are shown in bold. The BPS contains deletions (shown as -) due to errors caused by reverse transcriptase reading through the branch. b, RT–PCR was carried out on RNA extracted from wild-type yeast cells with or without RNaseR treatment using primers indicated in the schematic diagrams below the gel, indicating that RNase R treatment eliminates linear RNAs. This experiment was repeated four additional times with similar results. c, Protein and RNA components of E complex assembled on EFM5 IEI-101–M3 pre-mRNA. d, RT–PCR of RNA extracted from BY4742 yeast strain carrying indicated HRMA1 plasmids, with or without RNaseR treatment, using primers shown in the schematic diagrams below the gel. Numbers 246 and 62 designate exon lengths. Lanes 1–3 indicate that all constructs were transcribed (endogenous HMRA1 pre-mRNA level is too low to be detected as indicated in lane 3). The HMRA1 middle exon was slightly modified to create a circRNA primer binding site so that only the modified exogenous (for example, IEI-246 in lane 5) but not wild-type HMRA1 circRNA (IEI-246 WT in lane 4) could be detected. e, IEI-246–M3 RNA or E complex assembled on IEI-246–M3 was incubated with wild-type or U1-depleted yeast extract in the absence or presence of 30-fold excess competing IEI-246 wild-type RNA. CircRNA products were monitored using RT–PCR as in d. Experiments in c–e were repeated one additional time with similar results.
Supplementary information
Supplementary Figure
This file contains gel source data scans.
Rights and permissions
About this article
Cite this article
Li, X., Liu, S., Zhang, L. et al. A unified mechanism for intron and exon definition and back-splicing. Nature 573, 375–380 (2019). https://doi.org/10.1038/s41586-019-1523-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1523-6
This article is cited by
-
Computational approaches and challenges in the analysis of circRNA data
BMC Genomics (2024)
-
Circular RNAs and their roles in idiopathic pulmonary fibrosis
Respiratory Research (2024)
-
Utility of oligonucleotide in upregulating circular RNA production in a cellular model
Scientific Reports (2024)
-
Optimal design of synthetic circular RNAs
Experimental & Molecular Medicine (2024)
-
An axonemal intron splicing program sustains Plasmodium male development
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.