Abstract
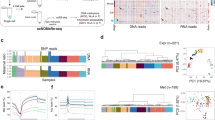
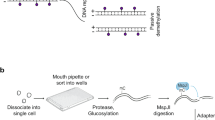
Implantation is a milestone event during mammalian embryogenesis. Implantation failure is a considerable cause of early pregnancy loss in humans1. Owing to the difficulty of obtaining human embryos early after implantation in vivo, it remains unclear how the gene regulatory network and epigenetic mechanisms control the implantation process. Here, by combining an in vitro culture system for the development human embryos after implantation and single-cell multi-omics sequencing technologies, more than 8,000 individual cells from 65 human peri-implantation embryos were systematically analysed. Unsupervised dimensionality reduction and clustering algorithms of the transcriptome data show stepwise implantation routes for the epiblast, primitive endoderm and trophectoderm lineages, suggesting robust preparation for the proper establishment of a mother-to-offspring connection during implantation. Female embryos showed initiation of random X chromosome inactivation based on analysis of parental allele-specific expression of X-chromosome-linked genes during implantation. Notably, using single-cell triple omics sequencing analysis, the re-methylation of the genome in cells from the primitive endoderm lineage was shown to be much slower than in cells of both epiblast and trophectoderm lineages during the implantation process, which indicates that there are distinct re-establishment features in the DNA methylome of the epiblast and primitive endoderm—even though both lineages are derived from the inner cell mass. Collectively, our work provides insights into the complex molecular mechanisms that regulate the implantation of human embryos, and helps to advance future efforts to understanding early embryonic development and reproductive medicine.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The single-cell RNA-sequencing data have been deposited in the GEO (accession number GSE109555). The single-cell MALBAC whole-genome sequencing, full-length RNA-sequencing and Trio-seq2 data have been deposited in the European Genome-phenome Archive (EGA; https://www.ebi.ac.uk/ega/) with accession number EGAS00001003443. The whole-genome sequencing data of the paternal sperm and maternal peripheral blood were from a previous publication (EGAS00001002987). The data deposited and made public are compliant with the regulations of Ministry of Science and Technology of China.
Code availability
Scripts of the main steps of the analysis are provided at https://github.com/WRui/Post_Implantation. Other R scripts associated with graphic presentation are available from the corresponding authors on reasonable request.
References
Koot, Y. E., Teklenburg, G., Salker, M. S., Brosens, J. J. & Macklon, N. S. Molecular aspects of implantation failure. Biochim. Biophys. Acta 1822, 1943–1950 (2012).
Shahbazi, M. N. et al. Self-organization of the human embryo in the absence of maternal tissues. Nat. Cell Biol. 18, 700–708 (2016).
Deglincerti, A. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016).
Gao, S. et al. Tracing the temporal-spatial transcriptome landscapes of the human fetal digestive tract using single-cell RNA-sequencing. Nat. Cell Biol. 20, 721–734 (2018).
Bian, S. et al. Single-cell multiomics sequencing and analyses of human colorectal cancer. Science 362, 1060–1063 (2018).
Dickens, B. M. International Society for Stem Cell Research (ISSCR) Guidelines for the Conduct of Human Embryonic Stem Cell Research (December 2006). Med. Law 27, 179–190 (2008).
National Research Council and Institute of Medicine of the National Acadamies. Final Report of the National Academies’ Human Embryonic Stem Cell Research Advisory Committee and 2010 Amendments to the National Academies’ Guidelines for Human Embryonic Stem Cell Research (National Academies Press, 2010).
Stirparo, G. G. et al. Integrated analysis of single-cell embryo data yields a unified transcriptome signature for the human pre-implantation epiblast. Development 145, dev158501 (2018).
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Nakamura, T. et al. A developmental coordinate of pluripotency among mice, monkeys and humans. Nature 537, 57–62 (2016).
Shahbazi, M. N. et al. Pluripotent state transitions coordinate morphogenesis in mouse and human embryos. Nature 552, 239–243 (2017).
Li, Y. et al. BMP4-directed trophoblast differentiation of human embryonic stem cells is mediated through a ΔNp63+ cytotrophoblast stem cell state. Development 140, 3965–3976 (2013).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Hou, Y. et al. Single-cell triple omics sequencing reveals genetic, epigenetic, and transcriptomic heterogeneity in hepatocellular carcinomas. Cell Res. 26, 304–319 (2016).
Lyon, M. F. Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 190, 372–373 (1961).
Deng, X. et al. Evidence for compensatory upregulation of expressed X-linked genes in mammals, Caenorhabditis elegans and Drosophila melanogaster. Nat. Genet. 43, 1179–1185 (2011).
Yildirim, E., Sadreyev, R. I., Pinter, S. F. & Lee, J. T. X-chromosome hyperactivation in mammals via nonlinear relationships between chromatin states and transcription. Nat. Struct. Mol. Biol. 19, 56–61 (2012).
Giorgetti, L. et al. Predictive polymer modeling reveals coupled fluctuations in chromosome conformation and transcription. Cell 157, 950–963 (2014).
Sangrithi, M. N. & Turner, J. M. A. Mammalian X chromosome dosage compensation: perspectives from the germ line. BioEssays 40, 1800024 (2018).
Brockdorff, N. & Turner, B. M. Dosage compensation in mammals. Cold Spring Harb. Perspect. Biol. 7, a019406 (2015).
Vallot, C. et al. XACT noncoding RNA competes with XIST in the control of X chromosome activity during human early development. Cell Stem Cell 20, 102–111 (2017).
Guo, H. et al. The DNA methylation landscape of human early embryos. Nature 511, 606–610 (2014).
Qiu, W. et al. Spatio-temporal expression of matrix metalloproteinase-26 in human placental trophoblasts and fetal red cells during normal placentation. Biol. Reprod. 72, 954–959 (2005).
Camolotto, S. et al. Expression and transcriptional regulation of individual pregnancy-specific glycoprotein genes in differentiating trophoblast cells. Placenta 31, 312–319 (2010).
Yadgary, L., Wong, E. A. & Uni, Z. Temporal transcriptome analysis of the chicken embryo yolk sac. BMC Genomics 15, 690 (2014).
Gerovska, D. & Araúzo-Bravo, M. J. Does mouse embryo primordial germ cell activation start before implantation as suggested by single-cell transcriptomics dynamics? Mol. Hum. Reprod. 22, 208–225 (2016).
Acknowledgements
We thank D. Xie and J. Zhou for discussions and Y. Hu, X. Fan, D. Liu, Y. Yuan and Y. Cui for technical help in pilot experiments; C. Shan from National Center for Protein Sciences at Peking University for immunofluorescence imaging and analysis. This work was supported by grants from the Beijing Municipal Science and Technology Commission (Z181100001318001), the National Natural Science Foundation of China (81521002, 31625018 and 81730078) and National Basic Research Program of China (2017YFA0102702 and 2018YFA0107601). F.Z. was supported by Young Elite Scientists Sponsorship Program by China Association for Science and Technology (YESS20160129), the Postdoctoral Fellowship of Peking-Tsinghua Center for Life Sciences and the grant from China Postdoctoral Science Foundation (2017M610015 and 2017T100015). F.Z. was a Bayer Postdoc of the Bayer-Peking University Center for Translational Research. This work was supported by the Beijing Advanced Innovation Center for Genomics at Peking University. Part of the computational analyses was performed on the High Performance Computing Platform of the Center for Life Science.
Author information
Authors and Affiliations
Contributions
F.T. and J.Q. conceived the project. F.Z., P.Y., Y.R. and Y.M. performed the experiments including the collection and culture of embryos, collection of single cells and single-cell sequencing data generation with the help of L.W., L.Y., R.L., J.L. and Y.L. R.W. performed computational analyses. F.T., F.Z. and R.W. wrote the manuscript with feedback from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Ali H. Brivanlou, Gist Croft, Insoo Hyun, Celine Vallot and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Lineage and sex identification of human pre- and post-implantation embryos.
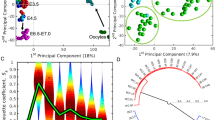
Related to Fig. 1. a–d, Immunofluorescence images of human embryos at different developmental stages (n = 3). Scale bars, 40 μm (a), 30 μm (b), 50 μm (c), 40 μm (d). e, Cell numbers of both sexes at each embryonic day. In total, 7,636 single cells were included. Day (D)6, n = 1,029 (filtered, n = 296; female, n = 207; male, n = 526). Day 8, n = 2,260 (filtered, n = 183; female, n = 1,288; male, n = 789). Day 10, n = 2,516 (filtered, n = 806; female, n = 1,022; male, n = 688). Day 12, n = 1,183 (filtered, n = 329; female, n = 281; male, n = 573). Day 14, n = 648 (filtered, n = 111; female, n = 0; male, n = 537). f, The number of expressed genes in each individual cell for different developmental stages. On average, 7,100 expressed genes and 139,792 transcripts were detected in each individual cell. Black lines indicate median values, the boxes range from the 25th to 75th percentiles and the whiskers correspond to 1.5× the IQR. In total, 5,911 single cells were included (day 6, n = 733; day 8, n = 2,077; day 10, n = 1,710; day 12, n = 854; day 14, n = 537). g, The number of cells and embryos of each cell lineage at distinct stages after filtering. E, embryo. h, The average mean levels of genes located on chromosome X (pink colour) and Y (blue colour) for each embryo. Mean expression ratios between X and Y are shown in green. Cells are ordered by sex and embryonic day. In total, 23 embryos with a sex expression ratio above two are defined as female embryos (2,798 cells from 4 stages), the remaining 25 embryos were male embryos (3,113 cells from 5 stages). The sex of the embryos highlighted in orange was confirmed by single-cell whole-genome sequencing. i, j, The unsupervised t-SNE plot of all cells at five representative stages, revealing a developmental path and cell lineage identification. i, In total, 5,911 cells were included (day 6, n = 733; day 8, n = 2,077; day 10, n = 1,710; day 12, n = 854; day 14, n = 537). j, Cells were identified as EPI, PE, TE and ysTE cells. Cells (dots) are coloured according to embryo stage and original lineage identity. EPI, n = 330; PE, n = 179; TE, n = 5,363; ysTE, n = 39. Clusters were assigned to indicate cell lineages using known lineage-specific markers. k, Lineage identification was further confirmed by lineage score analysis. The ggtern plot shows the lineage scores for each individual cell, calculated using previously published lineage-specific genes9. The cells are coloured according to their lineage identity.
Extended Data Fig. 2 Identification of gene-expression patterns during implantation.
Related to Fig. 2. a–d, Cluster annotation based on the expression levels of conventional lineage markers. Each dot represents a single cell, and the cells are coloured based on the expression levels (log2(TPM + 1)) of several known gene markers for specific lineages. For example, the EPI population expressed the key pluripotency markers POU5F1 (encoding OCT4), NANOG and SOX2. a–c, Colours from yellow to red represent expression levels from low to high. d, The violin plots show the expression of OTX2 in three main lineages; by setting scale = width, all violins have the same maximum width. In total, 5,911 cells were included; EPI, n = 330; PE, n = 179; TE, n = 5,363; ysTE, n = 39. e, The expression levels of conventional lineage markers at single-cell levels. The cells that clustered away from the other three main cell clusters showed high expression of TE markers (for example, GATA3) and low expression of EPI (for example, SOX2) and PE (for example, GATA4) markers. Colours from blue to red represent expression levels from low to high.
Extended Data Fig. 3 Human lineage-specific gene-expression patterns on previously reported monkey data.
The projection of signature genes for each lineage onto the monkey-related populations10 around implantation stages, including DPPA5, IFITM1 and MEG3 in EPI and derivatives (Gast1, Gast2a and Gast2b), GPX2, APOA1 and APOE in PE (also known as the hypoblast), and SLC7A2, TEAD1 and KRT7 in TE and derivatives (extra-embryonic mesenchyme, EXMC).
Extended Data Fig. 4 Lineage-specific gene-expression dynamics during implantation.
a, Principal component analysis for each lineage. Only cells that came from embryos that contained all three major lineages were used. In total, 3,145 single cells were included: EPI, n = 282; PE, n = 138; TE, n = 2,725. b, The developmental trajectory of each lineage based on Monocle2. Only cells that came from embryos that contained all three major lineages were used. In total, 3,145 single cells were used. EPI, n = 282; PE, n = 138; TE, n = 2,725. c–e, t-SNE analysis of EPI cells at all four stages revealed three clusters. c, Clusters are shown for each of the days. d, We found that the main reason one minor cluster (in grey) of these three clusters was separated might be owing to the differences in the number of expressed genes, although all individual cells from these clusters passed our quality control in the first procedure of data processing. We could not exclude that this was caused by low transcriptional activity of this cluster of cells or technical limitations in our system. e, We therefore removed this cluster of cells and focused on the differentially expressed genes between those two main clusters (C1 and C2). c–e, In total, 282 single cells were included. f, The results showed that the differentially expressed genes in the population were basically consistent with that in the internal EPI stages, reflecting that the differences in gene-expression characteristics between the two clusters are mainly due to the diversity in developmental stages (days 6 and 8 compared with days 10 and 12), indicating that EPI subgroups mainly reflected the developmental stage-specific differences. The gene list related to f is provided in Supplementary Table 5. g, Principal component analysis for TE at different developmental stages. The sublineages of TE emerge around day 10. Day 6 TE, n = 667 cells; day 10 TE, n = 1,443 cells; days 12 and 14 TE, n = 792 cells (day 12) and 533 cells (day 14).
Extended Data Fig. 5 Differential gene-expression features of the two TE clusters.
a, TE cells from day-12 and day-14 embryos were divided into two clusters. In total, 1,325 single cells were included. Day 12, n = 792 cells; day 14, n = 533 cells. b, Differential gene-expression features of the two TE clusters.
Extended Data Fig. 6 Representative CNV patterns during implantation.
a, Heat map shows large-scale CNVs in individual cells (rows) from a day-6 embryo based on single-cell RNA-sequencing data. The majority of cells from this embryo contained a whole-chromosomal duplication of chromosome 17, and a portion of the cells had a whole-chromosomal deletion of chromosome 7. b, CNVs confirmed by single-cell whole-genome sequencing. Related to a. c, Representative CNV-chimeric embryo. d, Heat map showing large-scale CNVs in individual cells (rows) from a day-8 embryo based on single-cell RNA-sequencing data. The majority of cells from this embryo contained a whole-chromosomal deletion of chromosome 22, and a portion of the cells had whole-chromosomal deletion of chromosome 13. Cells also show different chromosome X patterns at the transcriptome level. e, CNVs confirmed by single-cell whole-genome sequencing. Related to d. f, g, Heat maps show large-scale CNVs in individual cells (rows) from two day-12 embryos based on single-cell RNA-sequencing data.
Extended Data Fig. 7 Dynamics of chromosome X dosage during implantation.
Related to Fig. 3. a, The y axis represents the ratio of total expression levels of genes located on chromosome 1 and the same number of genes located on other autosomes (ChrA). Black lines indicate median values, the boxes range from the 25th to 75th percentiles and the whiskers correspond to 1.5× the IQR. In total, 3,184 single cells were included. Day 6, n = 387 (male); day 8, n = 1,525 (female, n = 1,147; male, n = 378); day 10, n = 1,021 (female, n = 917; male, n = 104); day 12, n = 251 (female, n = 104; male, n = 147) (Supplementary Table 1). b, The expression ratios of genes located on chromosome 1 or chromosome X to other autosomes in the different embryos that we sequenced. In total, 3,184 single cells were included: female, n = 2,168 single cells from 13 embryos; male, n = 1,016 single cells from 8 embryos. The statistical test was a two-sided t-test. c, The ratio of chromosome X to other autosomes in adult human digestive tract was used as a control. P, patient. P1, n = 763 cells; P2, n = 700 cells. The statistical test was a two-sided t-test. d, XIST expression in cells with different sexes. In total, 3,184 single cells were included; female, n = 2,168; male, n = 1,016. e, The distribution of XIST expression levels across different developmental stages for male and female embryos. In total, 3,184 single cells were included: day 6, n = 387 (male); day 8, n = 1,525 (female, n = 1,147; male, n = 378); day 10, n = 1,021 (female, n = 917; male, n = 104); day 12, n = 251 (female, n = 104; male, n = 147). f, XIST expression in different lineages for male and female embryos. In total, 3,184 single cells were included: female, n = 2,168; male, n = 1,016 (see details in Supplementary Table 1). g, The expression levels of XIST and XACT for different lineages. EPI, n = 282 cells; PE, n = 138 cells; TE, n = 2,725 cells. h, The distribution of XIST and XACT expression levels for different lineages. Both, cells that expressed XIST and XACT; XIST, cells that expressed only XIST; XACT, cells that expressed only XACT. EPI, n = 282 cells; PE, n = 138 cells; TE, n = 2,725 cells.
Extended Data Fig. 8 DNA methylation patterns during implantation.
a, Total number of embryos and cells collected at each stage (days 6–12) for single-cell Trio-seq analysis. b, t-SNE plot of cells based on the expression matrix. Each dot represents one cell and colours represent lineage types. For each lineage, we selected several individual cells for bisulfite sequencing. In total, 2,544 cells were included: EPI, n = 79 cells; PE, n = 136 cells; TE, n = 2,329 cells. c, Principal component analysis of cells based on the expression matrix. Only cells that were also used for bisulfite sequencing were used for the analysis. In total, 130 cells were used: EPI, n = 31 cells; PE, n = 45 cells; TE, n = 54 cells. d–f, The number of CpG sites detected with at least one-, three- and fivefold coverage across the single-cell samples. In total 2.7 Tb of sequencing data was generated, and on average 10 million CpG sites per cell were covered. Black line, median value. EPI, n = 31 cells; PE, n = 45 cells; TE, n = 54 cells. g, h, t-SNE analysis based on promoter methylation levels of genes. Each dot represents one cell and cells were coloured by culture day and lineage identity. EPI, n = 31 cells; PE, n = 45 cells; TE, n = 54 cells. i, Single-cell DNA methylation levels across gene bodies (from transcription start site (TSS) to transcription end site (TES)) and the 20-kb flanking regions of the transcription start and end sites. We found shared distribution patterns: genes were hypomethylated around the transcription start sites, evenly hypermethylated in the gene body regions and significant decrease in methylation after the transcription end sites.
Extended Data Fig. 9 The CNV landscapes of human early embryos by the DNA methylome dataset.
a, The CNVs for each individual cell at different developmental stages. Red represents duplication, blue represents deletion and white represents diploid. b, Representative examples of euploid and aneuploid single-cell samples. Abnormal copy numbers are highlighted in red (duplication) or blue (deletion). c, The number of cells with CNVs for each chromosome. d, The global DNA methylation levels for euploid and aneuploid cells. Black lines indicate median values, the boxes range from the 25th to 75th percentiles and the whiskers correspond to 1.5× the IQR. EPI, n = 61 cells (euploid, n = 31; aneuploidy, n = 30); PE, n = 84 cells (euploid, n = 45; aneuploidy, n = 39); TE, n = 141 cells (euploid, n = 54; aneuploidy, n = 87) (see details in Supplementary Table 1).
Extended Data Fig. 10 The re-methylation levels of different genomic elements vary between the three lineages at the different developmental stages.
a, Global methylation levels on different genomic elements for different cell lineages. EPI, n = 31 cells; PE, n = 45 cells; TE, n = 54 cells. b, The average DNA methylation levels of promoter regions (250 bp upstream to 250 bp downstream of the transcription start site) for each lineage at different developmental stages. Only promoters for which methylation levels were less than 0.1 at day 6 and more than 0.35 at days 10, 12 and 14 are shown in the heat map. Colours from blue to red represent methylation levels from low to high. EPI, n = 31 cells, PE, n = 45 cells; TE, n = 54 cells. c, DNA methylation levels of represent loci for lineage-specific genes at promoter regions. Each column represents one read. Red represents a methylated CpG site, blue represents an unmethylated CpG site and white represents an undetected site. Only reads that covered at least five CpG sites are shown in the heat map.
Supplementary information
Supplementary Information
This file contains the Ethics statement, Methods and Data availability.
Supplementary Tables
This file contains Supplementary Tables 1-10 and a Guide.
Video 1
Three-dimentional reconstruction image of a day6 human embryo (n=2). Nuclei (DAPI) are shown in blue, OCT4 in red.
Video 2
Three-dimentional reconstruction image of a day10 human embryo (n=3). Nuclei (DAPI) are shown in blue, OCT4 in red and GATA6 in green.
Video 3
Confocal Z-sections image of a day10 human embryo (n=3). Nuclei (DAPI) are shown in blue, OCT4 in red and GATA6 in green. Related to Supplementary Video 2.
Video 4
Three-dimentional reconstruction image of a day12 human embryo (n=3). Nuclei (DAPI) are shown in blue, OCT4 in red and GATA6 in green.
Video 5
Confocal Z-sections image of a day12 human embryo (n=3). Nuclei (DAPI) are shown in blue, OCT4 in red and GATA6 in green. Related to Supplementary Video 4.
Rights and permissions
About this article
Cite this article
Zhou, F., Wang, R., Yuan, P. et al. Reconstituting the transcriptome and DNA methylome landscapes of human implantation. Nature 572, 660–664 (2019). https://doi.org/10.1038/s41586-019-1500-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1500-0
This article is cited by
-
Shaoxia: a web-based interactive analysis platform for single cell RNA sequencing data
BMC Genomics (2024)
-
Disruption of maternal vascular remodeling by a fetal endoretrovirus-derived gene in preeclampsia
Genome Biology (2024)
-
Advances in single-cell omics and multiomics for high-resolution molecular profiling
Experimental & Molecular Medicine (2024)
-
PRAME induces genomic instability in uveal melanoma
Oncogene (2024)
-
Hypoblast from human pluripotent stem cells regulates epiblast development
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.