Abstract
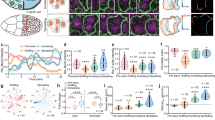
Tissue morphogenesis arises from coordinated changes in cell shape driven by actomyosin contractions. Patterns of gene expression regionalize cell behaviours by controlling actomyosin contractility. Here we report two modes of control over Rho1 and myosin II (MyoII) activation in the Drosophila endoderm. First, Rho1–MyoII are induced in a spatially restricted primordium via localized transcription of the G-protein-coupled receptor ligand Fog. Second, a tissue-scale wave of Rho1–MyoII activation and cell invagination progresses anteriorly away from the primordium. The wave does not require sustained gene transcription, and is not governed by regulated Fog delivery. Instead, MyoII inhibition blocks Rho1 activation and propagation, revealing a mechanical feedback driven by MyoII. We find that MyoII activation and invagination in each row of cells drives adhesion to the vitelline membrane mediated by integrins, apical spreading, MyoII activation and invagination in the next row. Endoderm morphogenesis thus emerges from local transcriptional initiation and a mechanically driven cycle of cell deformation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information files. Raw image data are available upon reasonable request.
Code availability
The custom codes used to process images analyse data and run simulations are available upon request.
Change history
27 August 2019
Owing to a technical error, this Article was not published online on 14 August 2019, as originally stated, and was instead first published online on 15 August 2019. The Article has been corrected online.
References
Lecuit, T. & Lenne, P. F. Cell surface mechanics and the control of cell shape, tissue patterns and morphogenesis. Nat. Rev. Mol. Cell Biol. 8, 633–644 (2007).
Gilmour, D., Rembold, M. & Leptin, M. From morphogen to morphogenesis and back. Nature 541, 311–320 (2017).
Leptin, M. Gastrulation in Drosophila: the logic and the cellular mechanisms. EMBO J. 18, 3187–3192 (1999).
Heisenberg, C. P. & Bellaïche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013).
Martin, A. C., Kaschube, M. & Wieschaus, E. F. Pulsed contractions of an actin–myosin network drive apical constriction. Nature 457, 495–499 (2009).
Solon, J., Kaya-Copur, A., Colombelli, J. & Brunner, D. Pulsed forces timed by a ratchet-like mechanism drive directed tissue movement during dorsal closure. Cell 137, 1331–1342 (2009).
Roh-Johnson, M. et al. Triggering a cell shape change by exploiting preexisting actomyosin contractions. Science 335, 1232–1235 (2012).
Martin, A. C. & Goldstein, B. Apical constriction: themes and variations on a cellular mechanism driving morphogenesis. Development 141, 1987–1998 (2014).
Mason, F. M., Tworoger, M. & Martin, A. C. Apical domain polarization localizes actin–myosin activity to drive ratchet-like apical constriction. Nat. Cell Biol. 15, 926–936 (2013).
Munjal, A., Philippe, J. M., Munro, E. & Lecuit, T. A self-organized biomechanical network drives shape changes during tissue morphogenesis. Nature 524, 351–355 (2015).
Marston, D. J. et al. MRCK-1 drives apical constriction in C. elegans by linking developmental patterning to force generation. Curr. Biol. 26, 2079–2089 (2016).
Michaux, J. B., Robin, F. B., McFadden, W. M. & Munro, E. M. Excitable RhoA dynamics drive pulsed contractions in the early C. elegans embryo. J. Cell Biol. 217, 4230–4252 (2018).
Kerridge, S. et al. Modular activation of Rho1 by GPCR signalling imparts polarized myosin II activation during morphogenesis. Nat. Cell Biol. 18, 261–270 (2016).
Manning, A. J., Peters, K. A., Peifer, M. & Rogers, S. L. Regulation of epithelial morphogenesis by the G protein-coupled receptor mist and its ligand fog. Sci. Signal. 6, ra98 (2013).
Costa, M., Wilson, E. T. & Wieschaus, E. A putative cell signal encoded by the folded gastrulation gene coordinates cell shape changes during Drosophila gastrulation. Cell 76, 1075–1089 (1994).
Dawes-Hoang, R. E. et al. Folded gastrulation, cell shape change and the control of myosin localization. Development 132, 4165–4178 (2005).
Rauzi, M., Lenne, P. F. & Lecuit, T. Planar polarized actomyosin contractile flows control epithelial junction remodelling. Nature 468, 1110–1114 (2010).
Munro, E., Nance, J. & Priess, J. R. Cortical flows powered by asymmetrical contraction transport PAR proteins to establish and maintain anterior–posterior polarity in the early C. elegans embryo. Dev. Cell 7, 413–424 (2004).
Kim, H. Y. & Davidson, L. A. Punctuated actin contractions during convergent extension and their permissive regulation by the non-canonical Wnt-signaling pathway. J. Cell Sci. 124, 635–646 (2011).
Maître, J. L., Niwayama, R., Turlier, H., Nédélec, F. & Hiiragi, T. Pulsatile cell-autonomous contractility drives compaction in the mouse embryo. Nat. Cell Biol. 17, 849–855 (2015).
Mayer, M., Depken, M., Bois, J. S., Jülicher, F. & Grill, S. W. Anisotropies in cortical tension reveal the physical basis of polarizing cortical flows. Nature 467, 617–621 (2010).
Behrndt, M. et al. Forces driving epithelial spreading in zebrafish gastrulation. Science 338, 257–260 (2012).
Bement, W. M. et al. Activator–inhibitor coupling between Rho signalling and actin assembly makes the cell cortex an excitable medium. Nat. Cell Biol. 17, 1471–1483 (2015).
Nishikawa, M., Naganathan, S. R., Jülicher, F. & Grill, S. W. Controlling contractile instabilities in the actomyosin cortex. eLife 6, e19595 (2017).
Fernandez-Gonzalez, R., Simoes, Sde. M., Röper, J. C., Eaton, S. & Zallen, J. A. Myosin II dynamics are regulated by tension in intercalating cells. Dev. Cell 17, 736–743 (2009).
Chanet, S. et al. Actomyosin meshwork mechanosensing enables tissue shape to orient cell force. Nat. Commun. 8, 15014 (2017).
Mitrossilis, D. et al. Mechanotransductive cascade of Myo-II-dependent mesoderm and endoderm invaginations in embryo gastrulation. Nat. Commun. 8, 13883 (2017).
Tan, P. Y. & Zaidel-Bar, R. Transient membrane localization of SPV-1 drives cyclical actomyosin contractions in the C. elegans spermatheca. Curr. Biol. 25, 141–151 (2015).
Collinet, C., Rauzi, M., Lenne, P. F. & Lecuit, T. Local and tissue-scale forces drive oriented junction growth during tissue extension. Nat. Cell Biol. 17, 1247–1258 (2015).
Lye, C. M. et al. Mechanical coupling between endoderm invagination and axis extension in Drosophila. PLoS Biol. 13, e1002292 (2015).
Young, P. E., Pesacreta, T. C. & Kiehart, D. P. Dynamic changes in the distribution of cytoplasmic myosin during Drosophila embryogenesis. Development 111, 1–14 (1991).
Sweeton, D., Parks, S., Costa, M. & Wieschaus, E. Gastrulation in Drosophila: the formation of the ventral furrow and posterior midgut invaginations. Development 112, 775–789 (1991).
Casanova, J. & Struhl, G. Localized surface activity of torso, a receptor tyrosine kinase, specifies terminal body pattern in Drosophila. Genes Dev. 3 (12B), 2025–2038 (1989).
Weigel, D., Jürgens, G., Klingler, M. & Jäckle, H. Two gap genes mediate maternal terminal pattern information in Drosophila. Science 248, 495–498 (1990).
Garcia, H. G., Tikhonov, M., Lin, A. & Gregor, T. Quantitative imaging of transcription in living Drosophila embryos links polymerase activity to patterning. Curr. Biol. 23, 2140–2145 (2013).
Edgar, B. A., Lehman, D. A. & O’Farrell, P. H. Transcriptional regulation of string (cdc25): a link between developmental programming and the cell cycle. Development 120, 3131–3143 (1994).
Paré, A. C. et al. A positional Toll receptor code directs convergent extension in Drosophila. Nature 515, 523–527 (2014).
Kerszberg, M. & Wolpert, L. Mechanisms for positional signalling by morphogen transport: a theoretical study. J. Theor. Biol. 191, 103–114 (1998).
Lander, A. D., Nie, Q. & Wan, F. Y. Do morphogen gradients arise by diffusion? Dev. Cell 2, 785–796 (2002).
Seher, T. C., Narasimha, M., Vogelsang, E. & Leptin, M. Analysis and reconstitution of the genetic cascade controlling early mesoderm morphogenesis in the Drosophila embryo. Mech. Dev. 124, 167–179 (2007).
Heissler, S. M. & Sellers, J. R. Kinetic adaptations of myosins for their diverse cellular functions. Traffic 17, 839–859 (2016).
Geiger, B., Spatz, J. P. & Bershadsky, A. D. Environmental sensing through focal adhesions. Nat. Rev. Mol. Cell Biol. 10, 21–33 (2009).
Iskratsch, T., Wolfenson, H. & Sheetz, M. P. Appreciating force and shape—the rise of mechanotransduction in cell biology. Nat. Rev. Mol. Cell Biol. 15, 825–833 (2014).
Rauzi, M. et al. Embryo-scale tissue mechanics during Drosophila gastrulation movements. Nat. Commun. 6, 8677 (2015).
Odell, G. M., Oster, G., Alberch, P. & Burnside, B. The mechanical basis of morphogenesis. I. Epithelial folding and invagination. Dev. Biol. 85, 446–462 (1981).
He, B., Doubrovinski, K., Polyakov, O. & Wieschaus, E. Apical constriction drives tissue-scale hydrodynamic flow to mediate cell elongation. Nature 508, 392–396 (2014).
Stark, K. A. et al. A novel α integrin subunit associates with βPS and functions in tissue morphogenesis and movement during Drosophila development. Development 124, 4583–4594 (1997).
Münster, S. et al. Attachment of the blastoderm to the vitelline envelope affects gastrulation of insects. Nature 568, 395–399 (2019).
Gelens, L., Anderson, G. A. & Ferrell, J. E. Jr. Spatial trigger waves: positive feedback gets you a long way. Mol. Biol. Cell 25, 3486–3493 (2014).
Allard, J. & Mogilner, A. Traveling waves in actin dynamics and cell motility. Curr. Opin. Cell Biol. 25, 107–115 (2013).
Diz-Muñoz, A., Fletcher, D. A. & Weiner, O. D. Use the force: membrane tension as an organizer of cell shape and motility. Trends Cell Biol. 23, 47–53 (2013).
Hashimoto, H., Robin, F. B., Sherrard, K. M. & Munro, E. M. Sequential contraction and exchange of apical junctions drives zippering and neural tube closure in a simple chordate. Dev. Cell 32, 241–255 (2015).
Morize, P., Christiansen, A. E., Costa, M., Parks, S. & Wieschaus, E. Hyperactivation of the folded gastrulation pathway induces specific cell shape changes. Development 125, 589–597 (1998).
Huang, J., Zhou, W., Dong, W., Watson, A. M. & Hong, Y. From the cover: directed, efficient, and versatile modifications of the Drosophila genome by genomic engineering. Proc. Natl Acad. Sci. USA 106, 8284–8289 (2009).
Venken, K. J. et al. Versatile P[acman] BAC libraries for transgenesis studies in Drosophila melanogaster. Nat. Methods 6, 431–434 (2009).
Bertet, C., Sulak, L. & Lecuit, T. Myosin-dependent junction remodelling controls planar cell intercalation and axis elongation. Nature 429, 667–671 (2004).
Cavey, M. & Lecuit, T. Imaging cellular and molecular dynamics in live embryos using fluorescent proteins. Methods Mol. Biol. 420, 219–238 (2008).
Müller, H. A. Immunolabeling of embryos. Methods Mol. Biol. 420, 207–218 (2008).
Fuse, N., Yu, F. & Hirose, S. Gprk2 adjusts Fog signaling to organize cell movements in Drosophila gastrulation. Development 140, 4246–4255 (2013).
Kosman, D., Small, S. & Reinitz, J. Rapid preparation of a panel of polyclonal antibodies to Drosophila segmentation proteins. Dev. Genes Evol. 208, 290–294 (1998).
Aigouy, B. et al. Cell flow reorients the axis of planar polarity in the wing epithelium of Drosophila. Cell 142, 773–786 (2010).
Butler, L. C. et al. Cell shape changes indicate a role for extrinsic tensile forces in Drosophila germ-band extension. Nat. Cell Biol. 11, 859–864 (2009).
Acknowledgements
We thank all members of the Lecuit and Lenne group for stimulating and useful discussions during the course of this project, the IBDM imaging facility for assistance with maintenance of the microscopes, FlyBase for maintaining curated databases and Bloomington for providing fly stocks. We are grateful to P. Tomancak for sharing the information regarding the expression pattern of scab published in ref. 48 and its function in Tribolium. This work was supported by the ERC grants Biomecamorph #323027 and SelfControl #788308. C.C. was supported by a Human Frontier Science Program Long-Term Fellowship (LT000733/2011-L) and the CNRS, A.B. was supported by a PhD fellowship from the LabEx INFORM (ANR-11-LABX-0054) and of the A∗MIDEX project (ANR-11-IDEX-0001–02), funded by the Investissements d’Avenir French government program. E.M. was supported by NIGMS 1RO1 GM098441-06. We acknowledge the France-BioImaging infrastructure supported by the French National Research Agency (ANR–10–INBS-04-01, Investments for the future).
Author information
Authors and Affiliations
Contributions
A.B., C.C. and T.L. conceived the project and planned experiments. A.B. and C.C. performed all experiments and quantifications. A.B. did the simulations with the help of E.M. J.-M.P. designed and generated the molecular constructs. A.B., C.C. and T.L. analysed the results and discussed them with E.M. P.-F.L. provided inputs. A.B., C.C. and T.L. wrote the manuscript and all authors made comments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Guy Blanchard, Emmanuel Farge, Kristen Panfilio and Barry Thompson for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Quantification of the morphogenetic wave within the dorsal posterior epithelium.
a–d, Procedure to generate kymograph heat maps from time lapses of endoderm morphogenesis. a, Stills of a dual-colour time lapse, E-cad–GFP is in magenta and MyoII is in green. b, A kymograph generated along the yellow horizontal line in a. A single cell, highlighted in yellow, first moves towards the anterior and then increases its projected apical area before recruiting MyoII. Solid yellow lines: time of the stills in a; dashed vertical line: the position of the cell at time t0. c, Kymograph heat map of MyoII integrated intensity measured using cell tracking in a. Data are averaged along the M–L axis within the white dashed line box in a and cell positions are the positions at time t. Note that cell tracks display movements similar to the kymograph in b. d, Kymograph heat map of MyoII integrated intensity registered on the cell positions at time t0 extracted from cell tracking in a. Data are averaged along the M–L axis as in c, but each cell track is plotted according to its position along the A–P axis at time t0. Cell tracks do not display any movement and appear as straight vertical lines. As a result, tissue deformation is not considered and the movement of the wave across the tissue can be visualized. Time t0 is the onset of endoderm morphogenesis. In a–d, n = 1 embryo for demonstration purposes. e, f, Kymograph heat maps of A–P apical cell projected length (e) and cell aspect ratio (f). The primordium and propagation regions are indicated. n = 947 cells from 5 embryos. g, Representation of the bending angle θ in a lateral view from a tracked cell (yellow contour) (Methods). h, Average time traces of the corrected area (Methods) and MyoII integrated intensity. Cells are registered on t0, the time of MyoII activation. The corrected area decreases when MyoII intensity increases indicating that cells are truly constricting. i, Time trace of the relative error in the measurement of cell area (that is, underestimation of the real cell area) due to the bending angle. In g–i, n = 24 cells, 1 embryo, with θ measured automatically (method 1, Methods) on MyoII side views. Mean ± s.d. are shown.
Extended Data Fig. 2 MyoII activation in cells of the primordium and propagation regions occurs differently.
a, Close-up views of cells in the primordium and propagation regions. b, Projected apical cell area and MyoII integrated intensity of the cells in a. Yellow and green boxes, steps of MyoII recruitment in the primordium cell and the cell in the propagation region, respectively. c, MyoII recruitment rate in cells of the indicated regions (n = 288 cells for primordium and 456 cells for propagation region, 6 embryos). Box plots show the median, 25th and 75th percentiles. The grey crosses label outliers. P = 5.1 × 10−58, two-sided Mann–Whitney test.
Extended Data Fig. 3 Rho–GTP and Rok propagate together with MyoII during the wave.
a, c, High-resolution stills of cells in the propagation region labelled with MyoII and with the Rho1–GTP sensor anillin(RBD)–GFP (n = 4 embryos) (a), and RokKD–GFP (n = 4 embryos) (c). Cell contours are in yellow. b, d, Curves of Rho1 sensor (b) and Rok (d) mean intensity over time along with MyoII in one representative cell of the propagation region. Intensity values are normalized to the maximum of each curve for visualization purposes. In the insets, scatter plots of MyoII mean intensity versus Rho1 sensor mean intensity (b) or Rok mean intensity (d) from about 10 cells in the embryo shown in a and about 7 cells in the embryo shown in c (Methods). In the insets in b and d, R2 is the square of the Pearson correlation coefficient. In a–d, t0 is arbitrary.
Extended Data Fig. 4 Terminal patterning controls MyoII activation in the posterior endoderm.
a, Schematics of the terminal patterning-dependent pathway controlling MyoII activation in the posterior endoderm. b, MyoII activation in the posterior endoderm in embryos mutant for torso (mat tor−/−), depleted of maternal and zygotic fog (fog RNAi) and mutant for the Gα12/13 concertina (mat cta−/−). Yellow contours mark cells in the propagation region. n = 5 embryos for wild type, n = 4 for mat tor−/−, n = 5 for fog RNAi and n = 5 for mat cta−/−. c, Sagittal sections of embryos immunostained for Fog and Tll (together with the membrane marker neurotactin in the merge) at different stages of posterior endoderm morphogenesis. The white and green lines indicate the boundaries of the expression domains of fog and tll, respectively. n = 8 embryos for primordium contraction stages and n = 5 embryos for MyoII propagation stages, from one independent experiment in this configuration.
Extended Data Fig. 5 MyoII propagation does not depend on gene transcription.
a, fog expression in the posterior endoderm visualized with the MS2–MCP system35 along with MyoII in living embryos. Top and side views are shown for the indicated time points. White dashed boxes indicate the positions of the close-up views on the right, and the yellow lines the region from which side views were generated. fog is expressed in the primordium but not in the propagation region. n = 4 embryos. b, Stills of a control- (water) and α-amanitin-injected embryo at a late stage of invagination. Asterisks indicate dividing cells (ROI1) and arrowheads indicate elongated cells (ROI2) in the posterior-ventrolateral ectoderm. Elongated cells in the ectoderm are a hallmark of cell stretching typical of conditions under which cell intercalation is affected29,30,62. n = 11 for water- and n = 10 for α-amanitin-injected embryos.
Extended Data Fig. 6 Fog diffusion does not control MyoII wave propagation.
a, Schematic of the dorsal epithelium in a sagittal section, illustrating the hypothesis of secreted Fog diffusion from the primordium controlling MyoII propagation. b, Apical views of embryos immunostained for Fog and MyoII at primordium contraction and late propagation stages during endoderm morphogenesis. n = 7 embryos for primordium contraction from 1 independent experiment and n = 17 embryos for propagation stage from 2 independent experiments. c, Quantifications of Fog mean intensity in the propagation region and in the dorsal epithelium. Individual embryo data points are superimposed on box plots. Box plots: black line, mean; grey box, s.d. n = 17 embryos. P = 7.1 × 10−7 from a two-sided Mann–Whitney test. d, Model of Fog secretion, diffusion, receptor binding and MyoII activation (Supplementary Information). e, Position of MyoII-activation front over time from one simulation of the model in b for the diffusion constant D, a given production rate (blue) and its double (red). f, Time-lapse of MyoII in the indicated conditions. The boxes mark the primordium and the activated cells in the propagation region. n = 4 embryos each. g, h, MyoII integrated intensity in cells of the primordium (g; n = 153 and 191 cells for wild type and hkb-fog, respectively, from 4 embryos each) and in a 10-µm-wide band of cells at about 30 µm distance from the primordium at time 0 (h; n = 69 and 67 cells for wild type and hkb-fog, respectively, from 4 embryos each). Mean ± s.d. between different embryos. i, Maximum of MyoII integrated intensity in cells of the indicated conditions. Box plots show median, 25th and 75th percentiles. Grey crosses label outliers. n is the number of cells from four embryos for each condition. P = 1.2 × 10−4 and P = 0.0017 from a two-sided Mann–Whitney test. j, Speed of MyoII propagation in the reference frame of the microscope. Box plots: black line, mean; grey box, s.d. n = 4 embryos each. N.S. indicates P = 1, two-sided Mann–Whitney test.
Extended Data Fig. 7 Patterned Fog signalling does not control MyoII wave propagation.
a, MyoII pattern in embryos in which Fog (UAS-Fog), a constitutively active Gα12/13 (UAS-Gα12[CA]) and Rho1 (UAS-Rho1[CA]) are expressed uniformly. Yellow and orange lines, the limits of the primordium and propagation regions, respectively. White arrows point to the same cell over time. n = 5, n = 8 and n = 4 embryos for UAS-Fog, UAS-Gα12[CA] and UAS-Rho1[CA], respectively. b, Left, pathway of MyoII activation in endoderm cells. Right, schematic of the Fog expression pattern (orange) in wild type and UAS-Fog, and of the observed MyoII pattern (green). The active and inactive zones used to calculate the MyoII tissue level polarity in c are indicated. c, d, Quantifications of MyoII tissue-level polarity 10 min after primordium activation (c) and of MyoII propagation (the number of cell rows of the propagation region activated within 20 min) (d) for the indicated conditions. For c, n = 10 (P = 4.1 × 10−12), n = 5 (P = 2.9 × 10−4), n = 8 (P = 0.0011) and n = 4 (P = 0.0027) embryos for wild type, UAS-Fog, UAS-Ga12[CA] and UAS-Rho1[CA], respectively, with the P values from a one-sample t-test against the hypothesis that the data are normally distributed with mean equals zero. For d, n is as for c, except n = 16 for wild type, and P = 0.076, P = 1.8 × 10−4 and P = 0.80 from a two-sided Mann–Whitney test for the indicated comparisons. e, Sagittal sections of a wild-type embryo and an embryo ubiquitously expressing Fog (UAS-Fog). Immunolabelling for Fog is shown in white and Tll is shown in green. n = 10 wild type and n = 2 UAS-Fog embryos from 2 independent experiments. f, MyoII pattern with Fog ubiquitous expression in a wild-type and a fog zygotic mutant embryos (imaged with 20× objective). n = 3 embryos each. g, Quantifications of MyoII patterns along the A–P axis in the posterior endoderm of the indicated conditions. n = 3 embryos each, mean ± s.d.
Extended Data Fig. 8 Cells in the propagation region are subjected to mechanical stress.
a, Left, regions in the dorsal epithelium (corresponding to the propagation region) in which tension was probed by line cuts with different orientations (M–L cut in blue and A–P cut in green). Right, examples of M–L and A–P line cuts in the indicated regions. Overlays of the pre-cut (magenta) and an image 10-s post-cut (green) are shown. The yellow line indicates the line cut. n is as in b for wild type. b, Quantifications of the tissue initial recoil velocity in the regions and line orientations illustrated in wild-type and H-1152-injected embryos. N indicates the number of independent ablations extracted from 26 and 7 embryos for wild type and H-1152, respectively. Box plots show the median, 25th and 75th percentiles. The red crosses label outliers. P values are from a two-sided Mann–Whitney test.
Extended Data Fig. 9 Mechanical perturbations affect propagation speed and MyoII recruitment.
a, Time-lapse of a wild-type embryo (left), an embryo with an anterior M–L fence (AF, middle) and a dorsalized embryo (dl−/−, right). Representative tracked cells in the propagation region are marked in yellow. Bottom, schematics of the normal and perturbed conditions. n = 9 wild-type embryos, 4 AF embryos and 5 dl−/− embryos. b, Speed of MyoII propagation in the reference frame of the tissue in the indicated conditions. n = 9 wild-type embryos, 4 AF embryos and 5 dl−/− embryos. P = 0.0028 and P = 1.0 × 10−3 from a two-sided Mann–Whitney test. c, d, Maximum MyoII integrated intensity (c) and minimum rate of apical constriction (d) in cells of the propagation region for the indicated conditions. n = 663 cells from 9 wild-type embryos, 93 cells from 4 AF embryos and 129 cells from 5 dl−/− embryos. P values are from a two-sided Mann–Whitney test. Box plots show median, 25th and 75th percentiles. The red crosses label outliers.
Extended Data Fig. 10 Rho1–GTP propagation is affected after Rok inhibitor injection.
a, High-resolution time lapse of Rho1–GTP (anillin(RBD)–GFP) in a control (water) and an embryo injected with 40 mM H-1152 (Rok inhibitor) during propagation. Larger views and later times after injection are shown (compared to Fig. 3f). The numbers in yellow indicate cells belonging to cell rows at different distances from the invaginating furrow. n = 6 embryos for H-1152 and n = 3 for water injections, respectively. b, c, Measurement of Rho1–GTP (anillin(RBD)–GFP) mean intensity over time in cells (or regions of cells) at different distances from the invaginating furrow in embryos injected with water (b) and H-1152 (c). Representative of n = 47 cells for water- and 47 cells for H-1152-injected embryos.
Extended Data Fig. 11 MyoII activation in cells of the propagation region.
a, Top and side views of MyoII activation in cells of the propagation region. The cell contour is labelled in yellow. White arrows indicate accumulations of MyoII in regions of the cell in contact with the vitelline membrane. n = 50 cells from 3 embryos. b, Time trace of MyoII integrated intensity of the cell in a. The selected stills in a are indicated along with the time of the cell posterior and anterior edge detachment. c, Histogram of the time of cell posterior and anterior edge detachment from the vitelline membrane relatively to the time of MyoII activation (defined as time 0). n = 50 cells from 3 embryos. d, Average velocity (relative to the vitelline membrane) of the posterior and anterior edges of cells in the propagation region. Pink and green boxes: phases of rapid cell displacement and immobilization of the posterior edge respectively. Time t0 is defined for each cell as the time of maximum projected area increase rate. Mean ± s.d.; n = 456 cells, 6 embryos.
Extended Data Fig. 12 The 3D cycle of cell deformations during wave propagation depends on sustained MyoII activity.
a, Time-lapse imaging of dextran injection in the perivitelline space. Left, dextran alone; right, merged images showing E-cad and MyoII. The arrows indicate the same two cells over time as the invaginating furrow approaches them. b, Space-intensity plot of MyoII and dextran intensity in the propagation region. n = 3 embryos; mean ± s.d. c–e, Scatter plots of the average speed of the 3D cycle during wave propagation calculated from the MyoII activation front versus the speed calculated from the projected apical cell area or cell position along the apico-basal axis of the tissue for the indicated conditions (n = 18 embryos). R2 values for a fit y = x are shown. f, A control and an embryo injected with H-1152 (Rok inhibitor) during MyoII propagation. Reconstructed side views from confocal stacks are shown. White asterisks and dashed lines mark a single cell over time. Time t0 is the time of injection. n = 2 embryos for both water- (control) and H-1152-injected embryos.
Supplementary information
Supplementary Information
This file contains Supplementary Methods with details of the simulations of the Myosin-II wave.
Supplementary Table 1
List of genotypes employed in the experiments in the indicated figure panels.
Supplementary Table 2
The sequences of the forward and reverse primers used to synthesize dsRNAs against fog and scab transcripts are listed. In bold is the annealing sequence. The remaining sequence is the T7 promoter.
Video 1
Tissue-level MyoII dynamics during endoderm morphogenesis. Time-lapse of posterior endoderm morphogenesis in a WT embryo. E-cad-GFP is in magenta and MRLC-mCherry in green. The yellow contours mark tracked cells in the propagation region. Scalebar 15 μm. N=13 embryos.
Video 2
Cell-level MyoII dynamics in the primordium and propagation regions. Time-lapse of MyoII recruitment in a cell in the primordium region (left) and in a cell in the propagation region, corresponding to cells in Extended Data Fig. 2a and relative measurements in Extended Data Fig. 2b. E-cad-GFP is in magenta and MRLC-mCherry in green. The cell contours are marked in yellow. Scalebar 3 μm. N=13 embryos.
Video 3
MyoII and Rho1 activation in cells of the propagation region. High resolution time-lapse of cells in the propagation region labelled with the Rho1-GTP sensor anillin(RBD)-GFP (left) and MyoII (right). The white arrows point to Rho1-GTP and MyoII accumulations during wave propagation. Scalebar 5 μm. N=4 embryos.
Video 4
MyoII and Rok activation in cells of the propagation region. High resolution time-lapse of cells in the propagation region labelled with Rok(KD)-GFP (left) and MyoII (right). The white arrows point to Rok and MyoII accumulations during wave propagation. Scalebar 5 μm. N=4 embryos.
Video 5
Tissue-level MyoII dynamics in the endoderm of torso, fog and concertina mutant embryos. Time-lapse of MyoII activation during posterior endoderm morphogenesis in a torso mutant embryo (laid by mothers tor -/-, top), in a strong knock-down of fog (fog RNAi, middle) and in a concertina (Gα12/13) mutant embryo (laid by mothers cta -/-, bottom). MyoII is labelled with MRLC-mCherry. Scalebar is 20 μm. N=4 embryos for mat tor-/-, 5 for fog RNAi and 5 for mat cta-/-.
Video 6
fog expression and MyoII propagation in a living embryo. Top and side views of fog expression (fog-MS2 revealed by MCP-GFP) on the left and MyoII (MRLC-mCherry) activation on the right during posterior endoderm morphogenesis in a WT embryo. Scalebar 15 μm. N=4 embryos.
Video 7
fog expression and MyoII propagation following α-amanitin injection. Top and side views of fog expression (fog-MS2 revealed by MCP-GFP) on the left and MyoII (MRLC-mCherry) activation on the right during posterior endoderm morphogenesis in a water (top) and α-amanitin (bottom) injected embryos. Scalebar 15 μm. N=2 and N=3 for water and α-amanitin injected embryos respectively.
Video 8
MyoII propagation following α-amanitin injection. Time-lapse of MyoII activation and propagation during posterior endoderm morphogenesis in a water (top) and α-amanitin (bottom) injected embryos. E-cad-GFP is magenta and MRLC-mCherry in green. The yellow contours mark tracked cells in the propagation region and the white arrows indicate cells at the moment of MyoII activation. Scalebar 20 μm. N=11 for water and N=10 for α-amanitin injected embryos.
Video 9
Tissue-level MyoII dynamics in the endoderm in a WT and a hkb-Fog embryos. Time-lapse of MyoII activation and propagation during posterior endoderm morphogenesis in a WT (top) and an embryo over-expressing fog in the primordium (hkb-Fog, bottom). E-cad-GFP is magenta and MRLC-mCherry in green. Time registration has been performed as described in the methods. Scalebar 20 μm. N=4 embryos each.
Video 10
Dorsal tissue-level MyoII dynamics in embryos where the Fog/GPCR/Rho1 pathway is constitutively active. Time-lapse of MyoII activation during posterior endoderm morphogenesis in embryos expressing ubiquitously Fog (UAS-Fog, top), a constitutively active version of Gα12/13, (UAS-Cta[CA], middle) or a constitutively active version of Rho1 (UAS-Rho1[CA], bottom). MyoII is labelled with MRLC-mCherry. Scalebar 20 μm. N=5, N=8 and N=4 embryos for UAS-Fog, UAS-Cta[CA] and UAS-Rho1[CA] respectively.
Video 11
Rho1-GTP and MyoII tissue-level dynamics in the endoderm in a WT embryo. Rho1 and MyoII activation and propagation during posterior endoderm morphogenesis in a water injected embryo at the end of cellularization. Rho1-GFP is detected with the Rho1 sensor anillin(RBD)-GFP (top) and MyoII is labelled with MRLC-mCherry (bottom). The white arrows indicate events of Rho1 and MyoII activation in cells of the propagation region. Scalebar 15 μm. N=10 embryos.
Video 12
Rho1-GTP and MyoII tissue-level dynamics in the endoderm following Rok inhibition. Rho1 and MyoII activation and propagation during posterior endoderm morphogenesis in an embryo injected with the Rok inhibitor H-1152 at the end of cellularization. Rho1-GFP is detected with the Rho1 sensor anillin(RBD)-GFP (top) and MyoII is labelled with MRLC-mCherry (bottom). The white arrows indicate events of Rho1 and MyoII activation in cells of the propagation region. Scalebar 15 μm. N=12 embryos.
Video 13
MyoII propagation in embryos with mechanical perturbations. Posterior endoderm morphogenesis in a WT embryo (top), an embryo with an anterior dorsal fence (middle) and a dorsalized embryo laid by dorsal -/- mothers (bottom). E-cad-GFP is in magenta and MRLC-mCherry in green. The yellow contours mark tracked cells in the propagation region. Scalebar 15 μm. N=9 WT embryos, 4 AF embryos and 5 dl-/- embryos.
Video 14
Gastrulation in a WT and a dorsalized embryo. Gastrulation in a WT (top) and a dorsalized embryo laid by dorsal -/- mothers (bottom). Scalebar 50 μm. N=14 embryos.
Video 15
Rho1-GTP and MyoII tissue-level dynamics in the endoderm in WT following Rok inhibition during propagation. Time-lapse of a water (left) or Rok inhibitor (H-1152) injection during Rho1/MyoII wave propagation. Rho1-GFP is detected with the Rho1 sensor anillin(RBD)-GFP (top) and MyoII is labelled with MRLC-mCherry (bottom). Large views and long time lapse after injection. Note the slow and low-level activation of Rho1 in cells anterior to the invaginating furrow following Rok inhibition (top right). The labels ‘Water’ and ‘H-1152’ mark the time of injection. Scalebar 5 μm. N=3 embryos for WT and N=6 embryos for H-1152 injection.
Video 16
Rho1-GTP and MyoII cell-level dynamics in the endoderm in WT following Rok inhibition during propagation. Time-lapse of a water (left) or Rok inhibitor (H-1152) injection during Rho1/MyoII wave propagation. Rho1-GFP is detected with the Rho1 sensor anillin(RBD)-GFP (top) and MyoII is labelled with MRLC-mCherry (bottom). Close-up on cells in the process of activating Rho1/MyoII at the time of injection. The white arrows point to Rho1-GTP and MyoII accumulations just before injection. The labels ‘Water’ and ‘H-1152’ mark the time of injection. Scalebar 5 μm. N=3 embryos each.
Video 17
Tissue-level side view of endoderm morphogenesis. Side view of MyoII recruitment and cell deformations during posterior endoderm morphogenesis. MyoII is labelled MRLC-GFP and the white arrows indicate events of MyoII activation during wave propagation. Scalebar 20 μm. N=2 embryos.
Video 18
Cell-level 3D views of MyoII activation during wave propagation. High-resolution time-lapse of MyoII (MRLC-mCherry) recruitment in a cell in the propagation region. Top and side views are shown. The cell contours are marked in yellow. At the bottom, a graph of MyoII integrated intensity with a moving vertical bar indicating the time in the time lapse. The white arrows point to bright accumulations of MyoII and the yellow arrows indicate the time of cell detachment at the posterior. Scalebar 5 μm. N=3 embryos.
Video 19
Cell-level 3D views of apical cell spreading during wave propagation. High-resolution time-lapse of E-cad (E-cad-GFP) during apical cortex spreading onto the vitelline membrane in a cell in the propagation region. Top and side views are shown. Scalebar 5 μm.
Video 20
Perivitelline dextran during wave propagation. Time lapse of a perivitelline injection of fluorescent Dextran labelling the extracellular space between cells and the vitelline membrane during MyoII wave propagation. Dextran is in magenta, MRLC-mCherry is labelled in green and E-cad-GFP is in white. Scalebar 5 μm. N=3 embryos.
Video 21
3D views of cells and MyoII following Rok inhibition during wave propagation. Time-lapse of a Rok inhibitor (H-1152) injection during MyoII propagation. Top and side views are shown. F-actin, labelling cell contours, is detected with Utrophin(ADB)-GFP (top) and MyoII is labelled with MRLC-mCherry (bottom). The label 'H-1152' marks the time of injection. Scalebar 15 μm. N=2 embryos each.
Video 22
Cell attachments to the vitelline membrane during wave propagation. Time-lapse of cells in the propagation region illustrating events of transient cell adherence to the vitelline membrane (seen as cells lagging behind the moving invaginating furrow and then suddenly detaching in the invagination). MyoII, labelled by MRLC-GFP, is in white and cell plasma membrane, labelled by Gap-43-mCherry, is in green. Scalebar 5 μm. N=4 embryos.
Video 23
Comparison of tissue and MyoII dynamics in a WT and a scab RNAi embryo. Top and side views of posterior endoderm morphogenesis in a control (water injection) embryo (top), and an embryo injected with dsRNAs against scab (bottom). E-cad-GFP is in magenta and MRLC-mCherry in green. The yellow contours mark tracked cells in the propagation region. Scalebar 15 μm. N=4 embryos each.
Rights and permissions
About this article
Cite this article
Bailles, A., Collinet, C., Philippe, JM. et al. Genetic induction and mechanochemical propagation of a morphogenetic wave. Nature 572, 467–473 (2019). https://doi.org/10.1038/s41586-019-1492-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1492-9
This article is cited by
-
Friction forces determine cytoplasmic reorganization and shape changes of ascidian oocytes upon fertilization
Nature Physics (2024)
-
In silico analysis shows that dynamic changes in curvature guide cell migration over long distances
Biomechanics and Modeling in Mechanobiology (2024)
-
Growth anisotropy of the extracellular matrix shapes a developing organ
Nature Communications (2023)
-
Hexanematic crossover in epithelial monolayers depends on cell adhesion and cell density
Nature Communications (2023)
-
The first embryo, the origin of cancer and animal phylogeny. II. The neoplastic process as an evolutionary engine
Journal of Biosciences (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.