Abstract
Living systems are capable of locomotion, reconfiguration and replication. To perform these tasks, cells spatiotemporally coordinate the interactions of force-generating, ‘active’ molecules that create and manipulate non-equilibrium structures and force fields of up to millimetre length scales1,2,3. Experimental active-matter systems of biological or synthetic molecules are capable of spontaneously organizing into structures4,5 and generating global flows6,7,8,9. However, these experimental systems lack the spatiotemporal control found in cells, limiting their utility for studying non-equilibrium phenomena and bioinspired engineering. Here we uncover non-equilibrium phenomena and principles of boundary-mediated control by optically modulating structures and fluid flow in an engineered system of active biomolecules. Our system consists of purified microtubules and light-activatable motor proteins that crosslink and organize the microtubules into distinct structures upon illumination. We develop basic operations—defined as sets of light patterns—to create, move and merge the microtubule structures. By combining these operations, we create microtubule networks that span several hundred micrometres in length and contract at speeds up to an order of magnitude higher than the speed of an individual motor protein. We manipulate these contractile networks to generate and sculpt persistent fluid flows. The principles of boundary-mediated control that we uncover may be used to study emergent cellular structures and forces and to develop programmable active-matter devices.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the Caltech Research Data Repository at https://data.caltech.edu/records/1160. All plasmids used in this study are available at https://www.addgene.org. All of the other reagents and the source code used for this study are available from the corresponding authors upon reasonable request.
References
Marchetti, M. C. et al. Hydrodynamics of soft active matter. Rev. Mod. Phys. 85, 1143–1189 (2013).
Dumont, S. & Prakash, M. Emergent mechanics of biological structures. Mol. Biol. Cell 25, 3461–3465 (2014).
Needleman, D. & Dogic, Z. Active matter at the interface between materials science and cell biology. Nat. Rev. Mater. 2, 17048 (2017).
Nédélec, F. J., Surrey, T., Maggs, A. C. & Leibler, S. Self-organization of microtubules and motors. Nature 389, 305–308 (1997).
Surrey, T., Nédélec, F., Leibler, S. & Karsenti, E. Physical properties determining self-organization of motors and microtubules. Science 292, 1167–1171 (2001).
Sanchez, T., Chen, D. T. N., DeCamp, S. J., Heymann, M. & Dogic, Z. Spontaneous motion in hierarchically assembled active matter. Nature 491, 431–434 (2012).
DeCamp, S. J., Redner, G. S., Baskaran, A., Hagan, M. F. & Dogic, Z. Orientational order of motile defects in active nematics. Nat. Mater. 14, 1110 (2015).
Wu, K.-T. et al. Transition from turbulent to coherent flows in confined three-dimensional active fluids. Science 355, eaal1979 (2017).
Bricard, A., Caussin, J.-B., Desreumaux, N., Dauchot, O. & Bartolo, D. Emergence of macroscopic directed motion in populations of motile colloids. Nature 503, 95–98 (2013).
Nédélec, F., Surrey, T. & Maggs, A. C. Dynamic concentration of motors in microtubule arrays. Phys. Rev. Lett. 86, 3192–3195 (2001).
Lee, H. Y. & Kardar, M. Macroscopic equations for pattern formation in mixtures of microtubules and molecular motors. Phys. Rev. E 64, 056113 (2001).
Keber, F. C. et al. Topology and dynamics of active nematic vesicles. Science 345, 1135–1139 (2014).
Aoyama, S., Shimoike, M. & Hiratsuka, Y. Self-organized optical device driven by motor proteins. Proc. Natl Acad. Sci. USA 110, 16408–16413 (2013).
Guntas, G. et al. Engineering an improved light-induced dimer (iLID) for controlling the localization and activity of signaling proteins. Proc. Natl Acad. Sci. USA 112, 112–117 (2015).
Schuppler, M., Keber, F. C., Kröger, M. & Bausch, A. R. Boundaries steer the contraction of active gels. Nat. Commun. 7, 13120 (2016).
Belmonte, J. M., Leptin, M. & Nédélec, F. A theory that predicts behaviors of disordered cytoskeletal networks. Mol. Syst. Biol. 13, 941 (2017).
Foster, P. J., Fürthauer, S., Shelley, M. J. & Needleman, D. J. Active contraction of microtubule networks. eLife 4, e10837 (2015).
Good, M. C., Vahey, M. D., Skandarajah, A., Fletcher, D. A. & Heald, R. Cytoplasmic volume modulates spindle size during embryogenesis. Science 342, 856–860 (2013).
Weiner, O. D., Marganski, W. A., Wu, L. F., Altschuler, S. J. & Kirschner, M. W. An actin-based wave generator organizes cell motility. PLoS Biol. 5, e221 (2007).
Gardel, M. L., Schneider, I. C., Aratyn-Schaus, Y. & Waterman, C. M. Mechanical integration of actin and adhesion dynamics in cell migration. Annu. Rev. Cell Dev. Biol. 26, 315–333 (2010).
Theurkauf, W. E. Premature microtubule-dependent cytoplasmic streaming in cappuccino and spire mutant oocytes. Science 265, 2093–2096 (1994).
Ganguly, S., Williams, L. S., Palacios, I. M. & Goldstein, R. E. Cytoplasmic streaming in Drosophila oocytes varies with kinesin activity and correlates with the microtubule cytoskeleton architecture. Proc. Natl Acad. Sci. USA 109, 15109–15114 (2012).
Goldstein, R. E., Tuval, I. & van de Meent, J.-W. Microfluidics of cytoplasmic streaming and its implications for intracellular transport. Proc. Natl Acad. Sci. USA 105, 3663–3667 (2008).
Drescher, K., Dunkel, J., Cisneros, L. H., Ganguly, S. & Goldstein, R. E. Fluid dynamics and noise in bacterial cell–cell and cell–surface scattering. Proc. Natl Acad. Sci. USA 108, 10940–10945 (2011).
Drescher, K., Goldstein, R. E., Michel, N., Polin, M. & Tuval, I. Direct measurement of the flow field around swimming microorganisms. Phys. Rev. Lett. 105, 168101 (2010).
He, B., Doubrovinski, K., Polyakov, O. & Wieschaus, E. Apical constriction drives tissue-scale hydrodynamic flow to mediate cell elongation. Nature 508, 392–396 (2014).
Shinar, T., Mana, M., Piano, F. & Shelley, M. J. A model of cytoplasmically driven microtubulebased motion in the single-celled Caenorhabditis elegans embryo. Proc. Natl Acad. Sci. USA 108, 10508–10513 (2011).
Mittasch, M. et al. Non-invasive perturbations of intracellular flow reveal physical principles of cell organization. Nat. Cell Biol. 20, 344–351 (2018).
Acknowledgements
We thank M. Anjur-Dietrich, J. Brady, J. Bruck, V. Galstyan, S. Hirokawa, C. Hueschen, Y. Lazebnik, W. Lim, W. Marshall, D. Mullins, D. Needleman, P. Rothemund and E. Winfree for scientific discussions. We thank L. Bugaj, Z. Dogic, A. Frost, W. Huynh, R. Ismagilov, L. Metcalf, H. Nguyen and R. Vale for advice and assistance during the development of the experimental system; K. van den Dries for assistance with three-dimensional visualization of asters; P. Sternberg for use of a microscopy system for initial light-activation experiments. We are grateful to N. Orme for assistance with figures and illustrations. We acknowledge support from the NIH through grants 1R35 GM118043-01 (R.P.) and NIH DP5 OD012194 (M.T.); the NSF through NSF 1330864 (M.T.); the John Templeton Foundation as part of the Boundaries of Life Initiative through grants 51250 & 60973 (R.P.); the Foundational Questions Institute and Fetzer Franklin Fund through FQXi 1816 (R.P., M.T.); and the UCSF Center for Systems and Synthetic Biology NIGMS P50 GM081879 (M.T.). M.T. acknowledges support from the Heritage Medical Research Institute.
Author information
Authors and Affiliations
Contributions
T.D.R., H.J.L., R.P. and M.T. conceived the experiments and interpreted the results. T.D.R., H.J.L., R.A.B., Z.Q. and M.T. wrote the manuscript. T.D.R. designed and cloned iLID motor fusion constructs. T.D.R., H.J.L. and R.A.B. performed protein purification. T.D.R. and H.J.L. designed, performed and analysed the active-matter experiments. Z.Q. analysed and modelled flow data and tracked trajectories of moving asters. R.A.B. performed and analysed gliding assays. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Andreas Bausch, François Nédélec and Suraj Shankar for their contribution to the peer review of this work.
Supplementary information
Supplementary information
This file contains supplementary materials and methods, data acquisition, analysis and a supplementary discussion, including Supplementary Figures 1–25 and supplementary references.
Video 1
3D aster 100-µm excitation disk. 3D projection of aster formed with a 100-μm excitation disk. Scale bars are 30 μm. X, Y and Z axis are represented by the red, green, and blue bars, respectively.
Video 2
3D aster 30-µm excitation disk. 3D projection of aster formed with a 300-μm excitation disk. Scale bars are 30 μm. X, Y and Z axis are represented by the red, green, and blue bars, respectively.
Video 3
Aster formation and decay. Formation and decay of an aster with a 50-μm disk. Yellow disk indicates when and where light pattern is being projected. Time stamp is in min: sec.
Video 4
Aster with moving disk. Aster following an excitation disk moving at 100 nm s−1. Time stamp is in min: sec.
Video 5
Aster merger. Asters spaced 375 μm apart are linked together with light. Light pattern is indicated in yellow. Time stamp is in min: sec.
Video 6
Patterned asters. Asters of different sizes and positions are simultaneously patterned.
Video 7
Simultaneous movement of asters. Two asters simultaneously following disk patterns that are moving at 200 nm s−1. Time stamp is in min: sec.
Video 8
Spiralling aster. Aster following a spiralling disk pattern moving at 200 nm s−1. Time stamp is in min: sec.
Video 9
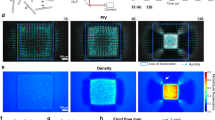
Active flow with labelled microtubules. Fluorescent microscopy time lapse of flow generated with a 750 × 20-μm bar. Time stamp is in min: sec.
Video 10
Active flow with tracer beads. Brightfield microscopy of flow generated with a 700 × 20-μm bars. Sample contains 1-μm beads for measuring advective flow. Time stamp is in min: sec.
Video 11
Plus-shape excitation and flow. Flow generated with a plus-shape pattern. The ‘+’ is composed of two 350 × 20-μm bars. Time stamp is in min: sec.
Video 12
T-shape excitation and flow. Flow generated with a T-shape pattern. The ‘T’ is composed of two 350 × 20-μm bars. Time stamp is in min: sec.
Video 13
L-shape excitation and flow. Flow generated with an L-shape pattern. The ‘L’ is composed of two 350 × 20-μm bars. Time stamp is in min: sec.
Video 14
Active stir bar. Dynamic flow pattern made with a rotating 450 × 30-μm bar. The ends of the bar rotate at 200 nm s−1. Time stamp is in min: sec.
Rights and permissions
About this article
Cite this article
Ross, T.D., Lee, H.J., Qu, Z. et al. Controlling organization and forces in active matter through optically defined boundaries. Nature 572, 224–229 (2019). https://doi.org/10.1038/s41586-019-1447-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1447-1
This article is cited by
-
Controlling liquid–liquid phase behaviour with an active fluid
Nature Materials (2023)
-
Competing instabilities reveal how to rationally design and control active crosslinked gels
Nature Communications (2022)
-
Topological active matter
Nature Reviews Physics (2022)
-
Self-mixing in microtubule-kinesin active fluid from nonuniform to uniform distribution of activity
Nature Communications (2022)
-
Bioinspired Centimeter-scale Sensor Free Obstacle-passing Robots with a Wireless Control System
Journal of Bionic Engineering (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.