Abstract
Lamin A/C (LMNA) is one of the most frequently mutated genes associated with dilated cardiomyopathy (DCM). DCM related to mutations in LMNA is a common inherited cardiomyopathy that is associated with systolic dysfunction and cardiac arrhythmias. Here we modelled the LMNA-related DCM in vitro using patient-specific induced pluripotent stem cell-derived cardiomyocytes (iPSC-CMs). Electrophysiological studies showed that the mutant iPSC-CMs displayed aberrant calcium homeostasis that led to arrhythmias at the single-cell level. Mechanistically, we show that the platelet-derived growth factor (PDGF) signalling pathway is activated in mutant iPSC-CMs compared to isogenic control iPSC-CMs. Conversely, pharmacological and molecular inhibition of the PDGF signalling pathway ameliorated the arrhythmic phenotypes of mutant iPSC-CMs in vitro. Taken together, our findings suggest that the activation of the PDGF pathway contributes to the pathogenesis of LMNA-related DCM and point to PDGF receptor-β (PDGFRB) as a potential therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data are available from the Gene Expression Omnibus (GEO; GSE118885).
References
Carmosino, M. et al. Role of nuclear lamin A/C in cardiomyocyte functions. Biol. Cell 106, 346–358 (2014).
Fatkin, D. et al. Missense mutations in the rod domain of the lamin A/C gene as causes of dilated cardiomyopathy and conduction-system disease. N. Engl. J. Med. 341, 1715–1724 (1999).
Krohne, G. & Benavente, R. The nuclear lamins. Exp. Cell Res. 162, 1–10 (1986).
Hershberger, R. E. & Morales, A. in GeneReviews (eds Pagon, R. A. et al.) (University of Washington, 1993).
Hershberger, R. E., Hedges, D. J. & Morales, A. Dilated cardiomyopathy: the complexity of a diverse genetic architecture. Nat. Rev. Cardiol. 10, 531–547 (2013).
Tesson, F. et al. Lamin A/C mutations in dilated cardiomyopathy. Cardiol. J. 21, 331–342 (2014).
Diecke, S. et al. Novel codon-optimized mini-intronic plasmid for efficient, inexpensive, and xeno-free induction of pluripotency. Sci. Rep. 5, 8081 (2015).
Kodo, K. et al. iPSC-derived cardiomyocytes reveal abnormal TGF-β signalling in left ventricular non-compaction cardiomyopathy. Nat. Cell Biol. 18, 1031–1042 (2016).
Lee, J. et al. SETD7 drives cardiac lineage commitment through stage-specific transcriptional activation. Cell Stem Cell 22, 428–444 (2018).
Burridge, P. W. et al. Chemically defined generation of human cardiomyocytes. Nat. Methods 11, 855–860 (2014).
Karakikes, I. et al. A comprehensive TALEN-based knockout library for generating human induced pluripotent stem cell-based models for cardiovascular diseases. Circ. Res. 120, 1561–1571 (2017).
Termglinchan, V., Seeger, T., Chen, C., Wu, J. C. & Karakikes, I. in Cardiac Gene Therapy (ed. Ishikawa, K.) 55–68 (Springer New York, 2017).
Bers, D. M. Calcium cycling and signaling in cardiac myocytes. Annu. Rev. Physiol. 70, 23–49 (2008).
Lan, F. et al. Abnormal calcium handling properties underlie familial hypertrophic cardiomyopathy pathology in patient-specific induced pluripotent stem cells. Cell Stem Cell 12, 101–113 (2013).
Itzhaki, I. et al. Modeling of catecholaminergic polymorphic ventricular tachycardia with patient-specific human-induced pluripotent stem cells. J. Am. Coll. Cardiol. 60, 990–1000 (2012).
Maizels, L. et al. Patient-specific drug screening using a human induced pluripotent stem cell model of catecholaminergic polymorphic ventricular tachycardia type 2. Circ Arrhythm Electrophysiol 10, e004725 (2017).
Bers, D. M. Cardiac sarcoplasmic reticulum calcium leak: basis and roles in cardiac dysfunction. Annu. Rev. Physiol. 76, 107–127 (2014).
Schreiber, K. H. & Kennedy, B. K. When lamins go bad: nuclear structure and disease. Cell 152, 1365–1375 (2013).
Kervestin, S. & Jacobson, A. NMD: a multifaceted response to premature translational termination. Nat. Rev. Mol. Cell Biol. 13, 700–712 (2012).
Seeger, T. et al. A premature termination codon mutation in MYBPC3 causes hypertrophic cardiomyopathy via chronic activation of nonsense-mediated decay. Circulation 139, 799–811 (2019).
Luperchio, T. R., Wong, X. & Reddy, K. L. Genome regulation at the peripheral zone: lamina associated domains in development and disease. Curr. Opin. Genet. Dev. 25, 50–61 (2014).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Perovanovic, J. et al. Laminopathies disrupt epigenomic developmental programs and cell fate. Sci. Transl. Med. 8, 335ra58 (2016).
Kind, J. & van Steensel, B. Genome–nuclear lamina interactions and gene regulation. Curr. Opin. Cell Biol. 22, 320–325 (2010).
Chen, X. et al. ATAC-see reveals the accessible genome by transposase-mediated imaging and sequencing. Nat. Methods 13, 1013–1020 (2016).
Gesson, K. et al. A-type lamins bind both hetero- and euchromatin, the latter being regulated by lamina-associated polypeptide 2 alpha. Genome Res. 26, 462–473 (2016).
Rønningen, T. et al. Prepatterning of differentiation-driven nuclear lamin A/C-associated chromatin domains by GlcNAcylated histone H2B. Genome Res. 25, 1825–1835 (2015).
Poleshko, A. et al. Genome–nuclear lamina interactions regulate cardiac stem cell lineage restriction. Cell 171, 573–587 (2017).
Lachmann, A. et al. Massive mining of publicly available RNA-seq data from human and mouse. Nat. Commun. 9, 1366 (2018).
Andrae, J., Gallini, R. & Betsholtz, C. Role of platelet-derived growth factors in physiology and medicine. Genes Dev. 22, 1276–1312 (2008).
Tompkins, J. D. et al. Mapping human pluripotent-to-cardiomyocyte differentiation: methylomes, transcriptomes, and exon DNA methylation “memories”. EBioMedicine 4, 74–85 (2016).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Chintalgattu, V. et al. Cardiomyocyte PDGFR-β signaling is an essential component of the mouse cardiac response to load-induced stress. J. Clin. Invest. 120, 472–484 (2010).
Mattout, A., Cabianca, D. S. & Gasser, S. M. Chromatin states and nuclear organization in development — a view from the nuclear lamina. Genome Biol. 16, 174 (2015).
Solovei, I. et al. LBR and lamin A/C sequentially tether peripheral heterochromatin and inversely regulate differentiation. Cell 152, 584–598 (2013).
Sharma, A. et al. High-throughput screening of tyrosine kinase inhibitor cardiotoxicity with human induced pluripotent stem cells. Sci. Transl. Med. 9, eaaf2584 (2017).
Lam, C. K. et al. Novel role of HAX-1 in ischemic injury protection involvement of heat shock protein 90. Circ. Res. 112, 79–89 (2013).
Greensmith, D. J. Ca analysis: an Excel based program for the analysis of intracellular calcium transients including multiple, simultaneous regression analysis. Comput. Methods Programs Biomed. 113, 241–250 (2014).
Lund, E., Oldenburg, A. R. & Collas, P. Enriched domain detector: a program for detection of wide genomic enrichment domains robust against local variations. Nucleic Acids Res. 42, e92 (2014).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLOS Comput. Biol. 9, e1003118 (2013).
Acknowledgements
We thank members of the laboratory of D. M. Bers for providing the pRYR2 antibody, and S. A. Yi, K. H. Nam, G. A. Akgun, C. Chen and S. Zhang for their contribution. This research is supported by AHA 17MERIT33610009, NIH R01 HL128170, R01 HL113006, R01 HL130020, R01 HL132875, R01 HL141851, Leducq Foundation 18CVD05 (J.C.W.); R01 HL139679, R00 HL104002, AHA 17IRG33410532 (I.K.); Prince Mahidol Award Foundation (V.T.); NIH K99 HL133473 (H.W.); the German Research Foundation (T.S.); National Research Foundation of Korea 2012R1A5A2A28671860 and 2019R1C1C1010675 (J.L.); NIH F32 HL139045 (E.L.); CIRM GC1R-06673-A, R24 HL117756 (M.P.S.); and NIH P50 HG007735 and Howard Hughes Medical Institute (H.Y.C.)
Author information
Authors and Affiliations
Contributions
J.C.W., I.K, V.T. and S.D. conceptualized and designed the study; J.C.W., I.K., J.L., V.T. and S.D. wrote the manuscript; V.T. and I.K. designed and performed the gene-editing experiments; J.L. analysed the majority of the data; I.I. and P.G. performed the electrophysiological experiments; C.K.L. and H.W. performed calcium analysis; X.C. and J.Z.Z. performed the ATAC-see experiments; M.A. performed the ATAC-seq experiments; M.G., E.L., J.L. and J.M.C. analysed all of the next-generation sequencing-related data; K.S. and J.-W.R. performed skin biopsies and clinical phenotype assessments; P.J.W. recruited the patients; J.L., R.C., T.S., T.C. and I.P.G. generated iPSC-CMs for the experiments; M.P.S. and H.Y.C. contributed to the experimental design; I.K. and J.C.W. supervised the study and provided funding support.
Corresponding authors
Ethics declarations
Competing interests
H.Y.C. is a advisor to10x Genomics. Stanford University has filed a patent application on ATAC-see technology, in which H.Y.C. is named a co-inventor. J.C.W. is a co-founder of Khloris Biosciences but has no competing interests, as the work presented was performed independently.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 LMNA-mutant iPSC-CMs can recapitulate arrhythmic phenotype of patients with LMNA-related DCM.
a, Schematic pedigree of the family carrying the mutation in LMNA. Patients (III-1, III-3, III-9, III-15 and III-17) and healthy individuals (IV-1 and IV-2) recruited for this study are numbered. Circles represent female family members and squares represent males. The ‘+’ and ‘−’ signs underneath family members indicate the presence or absence of the mutation in LMNA, respectively. b, Schematic view of 348-349insG frameshift mutation in LMNA. c, Genotyping of fibroblasts derived from patients and healthy controls. MT, mutant. d, Clinical features of patients and healthy individuals. e, Electrocardiogram of patients III-1 and III-3 and a healthy individual (IV-1). The electrocardiogram data were measured once per individual. f, g, Electrophysiological measurements of spontaneous action potentials in mutant iPSC-CMs (III-15 and III-17) recorded by patch clamp in current-clamp mode. The experiments were repeated three times independently with similar results.
Extended Data Fig. 2 The mutation in LMNA is a cause of the arrhythmic phenotype in LMNA-mutant iPSC-CMs.
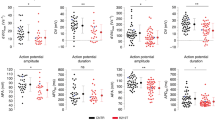
a, Gene-editing strategy using the TALEN method. The piggyBac system was used to generate isogenic lines as previously described11,12. b, Genotyping of gene-edited isogenic lines (III-3 corrected, insertion, deletion; IV-1 insertion). For LMNA del-KO/MUT, we used TALEN pairs that target the start codon of LMNA. Genotyping showed the C insertion in the wild-type allele that leads early stop codon. c, Immunostaining of NANOG (red, left), POU5F1 (red, middle) and SOX2 (red, right) in iPSC lines. Blue, DAPI. Scale bars, 10 μm. The experiments were repeated twice independently with similar results. d–f, Electrophysiological recordings of spontaneous action potentials in control (IV-1) and mutant (III-9, isogenic IV-1; WT/ins-MUT) iPSC-CMs measured by patch clamp in current-clamp mode. Red arrows indicate delayed afterdepolarization-like arrhythmias. The experiments were repeated three times independently with similar results. g, Action potential parameters of ventricular-like iPSC-CMs. MDP, maximal diastolic potential; APA, action potential amplitude; APD, action potential duration at 50%, 70%, 90% of repolarization; bpm, beats per min.
Extended Data Fig. 3 Abnormal calcium handling in LMNA-mutant iPSC-CMs.
a, Confocal imaging of Fluo-4 AM calcium events in control (III-3; WT/cor-WT) and mutant (III-3; WT/MUT) iPSC-CMs while being treated with increasing extracellular Ca2+ concentrations. All representative traces were recorded from three individual cells (presented as red, blue and black). b, Spontaneous calcium events per 100 s of control and mutant iPSC-CMs for each extracellular Ca2+ concentration. c, Summary of the percentage of cells that have spontaneous Ca2+ release events from the sarcoplasmic reticulum in control and mutant iPSC-CMs. d, qPCR analysis of CAMK2D and RYR2 expression in control and mutant iPSC-CMs. Data are mean ± s.e.m. e, f, Immunoblot analysis of pRYR2, RYR2, pCAMK2D and CAMK2D protein levels in control and mutant iPSC-CMs. Data are mean ± s.e.m.; a two-tailed Student’s t-test was used to calculate P values; n = 3; values above the lines indicate significance. g, qPCR analysis of CAMK2D expression in control and mutant iPSC-CMs. Expression level of GAPDH was used as control. Data are mean ± s.e.m.; n = 8. h, Representative Ca2+ transients of mutant iPSC-CMs (III-3; WT/MUT) treated with 1 μM of KN92 or KN93 for 24 h. i, Quantification of the percentage of cells that exhibit arrhythmic waveforms in mutant iPSC-CMs (III-17 WT/MUT) at baseline, as well as after the treatment with 1 μM of KN92 or KN93 for 24 h. j, Immunoblot analysis of pRYR2, RYR2, pCAMK2D and CAMK2D protein levels after treatment of DMSO, KN92 or KN93 for 24 h. The experiments in a were repeated twice independently with similar results. The Ca2+ transient analyses in h were repeated as described in Fig. 2e independently with similar results. The immunoblot analyses in e and j were repeated twice independently with similar results.
Extended Data Fig. 4 Downregulation of mutant mRNA through NMD pathway in LMNA-mutant iPSC-CMs.
a, Quantification of cells showing abnormal nuclear structures in control and mutant iPSC-CMs. The images were recorded from three differentiation batches. n = 215 (WT/cor-WT), n = 286 (WT/MUT), n = 222 (ins-MUT/MUT) and n = 280 (del-KO/MUT). b, Representative confocal images of control and mutant lines. Micro-patterned iPSC-CMs were stained with specific antibodies against TNNT2 (red), LMNA (white) and LMNB1 (green). Blue, DAPI. Scale bar, 20 μm. The experiments were repeated three times independently with similar results. c, Quantification of cells showing abnormal nuclear structures in control and mutant iPSC-CMs. The images were recorded from three differentiation batches. Data are mean ± s.e.m.; a two-tailed Student’s t-test was used to calculate P values; n = 3 (total number of counted cells, 175 (WT/WT) and 203 (WT/ins-MUT)); the value above the line indicates significance. d, Immunoblot analysis of lamin A/C levels in control and mutant iPSC-CMs. e, Quantification of signal intensity of the lamin A/C band in d. Data are mean ± s.e.m.; statistical significance was obtained using one-way ANOVA; values above the line indicate significance; n = 10 (WT/WT), n = 7 (WT/ins-MUT), n = 5 (WT/MUT). f, Immunoblot analysis of lamin A/C levels in two different clones of control and mutant iPSC-CMs. Two different antibodies that recognize the N terminus of lamin A/C were used. GAPDH was used as loading control. g, Relative mRNA expression of total LMNA in control and mutant iPSC-CMs. Data are mean ± s.e.m.; a two-tailed Student’s t-test was used to calculate P values; the value above the line indicates significance; n = 10 (WT/WT), n = 7 (WT/ins-MUT). h, Confirmation of allele-specific primers using plasmid carrying wild-type LMNA or mutant LMNA. Digital PCR using allele-specific primers detected the ratio of wild-type/mutant LMNA, which was consistent with the ratio of wild-type/mutant plasmids. Data are mean ± s.d.; n = 3. i, Immunoblot analysis of cell lysates from mutant iPSC-CMs treated with emetine and wortmannin. Two different batches of antibodies were used. Red asterisks indicate the truncated lamin A/C with a 14-kDa size. j, Immunoblot analysis of cell lysates from mutant iPSC-CMs treated with wortmannin. Three different batches of E-1 antibody detect the N terminus of LMNA and the 131C3 antibody detects the C terminus. k, Immunoblot analysis of cell lysates from control iPSC-CMs treated with emetine and wortmannin. The experiments in f, i–k were repeated twice independently with similar results.
Extended Data Fig. 5 Lamin A/C Haploinsufficiency results in an abnormal distribution of open chromatin in LMNA-mutant iPSC-CMs.
a–f, Representative images and normalized signal intensity of ATAC-see and DAPI of control and mutant iPSC-CMs. Data were obtained from different patient lines, including the uncorrected and isogenic lines of patient III-3 (a, b); the uncorrected and isogenic line of control IV-1 (c, d); and the line of patient III-15 (e, f) for normalized signal intensity of ATAC-see and DAPI. Data are mean ± s.e.m. g, Correlation of signal distribution between ATAC-see and DAPI. n = 42 (WT/WT), n = 28 (WT/cor-WT), n = 33 (del-KO/MUT), n = 32 (ins-WT/WT), n = 25 (WT/MUT) for normalized signal intensity of ATAC-see and DAPI. Data are mean and minimum to maximum; two-tailed Student’s t-test was used to calculate P values. The experiments in a, c, and e were repeated three times independently with similar results.
Extended Data Fig. 6 Genomic and chromatin features of LADs in control and mutant iPSC-CMs.
a, Normalized enrichment of lamin A/C ChIP–seq signals, histone markers (H3K4me3 and H3K27me3) and ATAC-seq signals within ±0.4 Mb of mapped LAD borders. The genomic locations of LADs were obtained from ChIP–seq on lamin A/C using two different antibodies (Abcam 8984, blue line, sc-376248, green line) in control iPSC-CMs (III-3). b, Representative images of ChIP–seq, ATAC-seq and RNA-seq of chromosome 12 (133 Mb). The red box shows LADs explicitly called in mutant iPSC-CMs (gain); the purple box shows LADs called in both control and mutant iPSC-CMs (overlapping); the blue box shows LADs explicitly called in control iPSC-CMs (loss). c–e, Number (c), genomic coverage (d) and mean of length of LADs (e) in control, mutant, gain, overlapping and loss LADs. ChIP–seq on lamin A/C (Abcam 8984) was used for data analysis. f, g, Average peak intensity of H3K4me3 and H3K27me3 of each LAD. n = 184 (loss), n = 370 (overlapping), n = 184 (gain) for H3K4me3; n = 273 (loss), n = 504 (overlapping), n = 273 (gain) for H3K27me3. Data are mean and minimum to maximum; Wilcoxon matched-pairs signed-rank test was used to calculate P values. h, Scatter plot of normalized lamin A/C, ATAC and histone marker (H3K4me3 and H3K27me3) enrichment of each LAD. The y axis shows the log2-transformed relative normalized lamin A/C enrichment of each LAD in mutant iPSC-CMs compared to control iPSC-CMs. The x axis shows the log2-transformed relative normalized ATAC and histone marks enrichment of each LAD in mutant iPSC-CMs compared with control iPSC-CMs. Each data point represents one LAD. The statistical significance was obtained using one-way ANOVA; n = 587 for sc-376248 and n = 585 for Abcam 8984. i, Percentage of differentially expressed genes in mutant iPSC-CMs compared to control iPSC-CMs. j, Number of differentially expressed genes located in mutant iPSC-CMs compared to control iPSC-CMs. (FDR-adjusted P < 0.01; log2-transformed fold change in expression of >1 or <−1). k, Distribution of log2-transformed fold change in FPKM in control and mutant iPSC-CMs. A non-parametric Kruskal–Wallis (testing for two-sided differences) followed by Dunn’s post hoc test was used to adjust for multiple comparisons; n = 266 (gain), n = 8171 (non-LADs), n = 835 (overlapping), n = 206 (loss).
Extended Data Fig. 7 Abnormal distribution of H3K9 methylation in mutant iPSC-CMs.
a, b, Representative images of immunofluorescence staining of control mutant iPSC-CMs. iPSC-CMs were stained with specific antibodies against H3K9me2 or H3K9me3 (green). Blue, DAPI. Scale bar, 1,000 nm. The experiments were repeated three times independently with similar results. c–e, Representative images of lamin A/C enrichment and LAD distribution of ChIP–seq data. ChIP–qPCR analysis of H3K9me2 and H3K9me3 enrichment on LAD regions. Data are mean ± s.d.; n = 3.
Extended Data Fig. 8 Transcription factors altered by lamin A/C haploinsufficiency contribute to the activation of genes located outside LADs.
a, Distribution of absolute distances to the nearest LAD (by nucleotide distance) from the transcription start site of genes that are differentially expressed (top) or that show no significant difference in expression between mutant and control iPSC-CMs (bottom). b, Distribution of median absolute log2-transformed change in expression of genes with relatively long (>7.5 × 106 bp) distances to the nearest LAD (top) and genes with relatively short (<2.5 × 106 bp) distances to the nearest LAD (bottom). In each category, 500 genes were sampled with replacement over 10,000 times. c, d, Co-occurrence analyses of transcription factors and genes and coexpression analyses of ARCHS4 transcription factors of differentially expressed genes located in non-LADs. Genes located in non-LADs are shown in blue; genes with no significant difference in gene expression between control and mutant iPSC-CMs are shown in black; genes located in LADs and highly expressed in mutant iPSC-CMs compared with control iPSC-CMs are shown in red. Top 200 differentially expressed genes located in non-LADs were used for the analysis. e, Representative images of ChIP–seq, ATAC-seq and RNA-seq of the genomic region of PRRX1. f, Relative mRNA expression of PRRX1, PDGFRB, GREM1, LUM and DCN in mutant iPSC-CMs treated with scramble or PRRX1 siRNA. Data are mean ± s.e.m.; a two-tailed Student’s t-test was used to calculate P values; n = 3 (PDGFRB and GREM1), n = 4 (DCN, LUM and PRRX1); values above the lines show significance.
Extended Data Fig. 9 PDGFRB is upregulated in LMNA-mutant iPSC-CMs.
a, Expression levels of PDGFRA and PDGFRB during the human iPSC-CM differentiation process. The data were adapted from previously published data (GSE76523). b, c, Protein and RNA levels of PDGFRB in human tissues. The data were adapted from the Human Protein Atlas Database32 v.18.1 (data available from http://www.proteinatlas.org/). d, qPCR analysis of PDGFRB expression in LMNA-mutant and control iPSC-CMs. Data are mean ± s.e.m.; a two-tailed Student’s t-test was used to calculate P values; n = 13 (WT/WT), n = 5 (WT/ins-MUT); the value above the line shows significance. e, Immunoblot analysis of PDGFRB protein levels in control versus mutant iPSC-CMs. GAPDH was used as loading control. The experiments were repeated twice independently with similar results. f, Flow cytometry analysis of TNNT2+PDGFRB+ cells in control and mutant iPSC-CMs. n = 4. g, Kinase array of control and mutant iPSC-CMs. Fifty different protein kinases were presented in each chip. Top, raw images of the blotting membrane. Two dots carried the same antibody in technical duplicates. Bottom, quantification of the signal intensity of each spot. h, Representative images of ChIP–seq, ATAC-seq and RNA-seq on the genomic regions of PDGFRB. The promoter region of PDGFRB is highlighted by a blue box. i, ChIP–qPCR of H3K4me3 and H3K27me3 enrichment at the promoter region of PDGFRB in control and mutant iPSC-CMs. n = 3. j, k, qPCR analysis of LMNA and PDGFRB expression levels in left ventricular heart tissue from health controls (n = 3) and patients with LMNA-related DCM (n = 2). Data are mean ± s.e.m. The kinase data in g were repeated twice independently with similar results. f, i, Data are mean ± s.e.m.; statistical significance was obtained using one-way ANOVA; values above the lines show significance.
Extended Data Fig. 10 Arrhythmic phenotype in mutant iPSC-CMs is dependent on the activation of the PDGFRB pathway.
a, qPCR analysis of PDGFRB expression levels in mutant iPSC-CMs (WT/MUT) treated with scramble or PDGFRB siRNAs. The cells were treated with siRNAs for 48 h. Data are mean ± s.e.m.; a two-tailed Student’s t-test was used to calculate P values; n = 3; the value above the line indicates significance. b, Representative Ca2+ transients of mutant iPSC-CMs (III-17 WT/MUT) treated with scramble siRNA or PDGFRB siRNA. c, Quantification of the number of cells that exhibited arrhythmic waveforms in b. d, Representative Ca2+ transients of mutant iPSC-CMs treated with PDGRB inhibitors, crenolanib (100 nM) and sunitinib (500 nM), for 24 h. All traces were recorded for 20 s. e, Quantification of mutant iPSC-CMs (III-17, III-15 and III-3) that exhibited arrhythmic waveforms with or without the treatment of PDGRB inhibitors, crenolanib (100 nM) and sunitinib (500 nM), for 24 h. f, Representative Ca2+ transients of mutant iPSC-CMs (III-17 WT/MUT) treated with PDGFRB inhibitors. g, Immunoblot analysis of pRYR2 and RYR2 protein levels with treatment of DMSO, crenolanib or sunitinib. The data were repeated twice independently with similar results. h, Immunoblot analysis of PDGFRB, tubulin, pCAMK2D and CAMK2D protein levels in control iPSC-CMs expressing empty and PDGFRB constructs. The signal intensity of the PDGFRB (left) and p-CAMK2D (right) is shown. The experiments were repeated twice independently with similar results. i, Representative Ca2+ transients of iPSC-CMs expressing empty and PDGFRB constructs. j, Quantification of arrhythmic waveforms of iPSC-CMs in i. The Ca2+ transients in b, d, f and i were repeated as described in c, e and j independently with similar results.
Extended Data Fig. 11 Gene-expression profile of PDGFRB inhibition in LMNA-mutant iPSC-CMs.
a, GO analysis of downregulated genes (n = 352) in LMNA-mutant iPSC-CMs treated with PDGFRB inhibitors, crenolanib (100 nM) and sunitinib (500 nM), for 24 h. b, Heat map of the expression profile of the gene set related to the GO function of ion transport. The FDR-adjusted P values were obtained using the GO enrichment analysis tool. c, Hierarchical clustering of AmpliSeq RNA-seq data using one-way ANOVA (P = 0.05; n = 230). Two different siRNAs against PDGFRB and a scramble siRNA were used in LMNA-mutant iPSC-CMs (III-15 WT/MUT). d, e, Heat map of expression profile of gene (n = 25) sets related with the GO function of cardiac muscle contraction (d) and actin-mediated cell contraction (e). The FDR-adjusted P values were obtained using the GO enrichment analysis tool. f, No significant changes in abnormal nuclear structures of mutant iPSC-CMs by inhibition of PDGFRB were found. Representative images of mutant iPSC-CMs treated with PDGFRB inhibitors, crenolanib (100 nM) and sunitinib (500 nM), for 24 h. iPSC-CMs were stained with specific antibodies against LMNB1 (green). Blue, DAPI. Scale bars, 10 μm. The experiments were repeated three times independently with similar results. g, Quantification of cells showing abnormal nuclear structures in mutant iPSC-CMs treated with PDGFRB inhibitors. The images were recorded from three differentiation batches. n = 90 (DMSO), n = 69 (crenolanib), n = 79 (sunitinib). Data are mean ± s.e.m.; statistical significance was analysed using one-way ANOVA; values above the lines indicate significance. h, Immunoblot analysis of lamin A/C and GAPDH protein levels in mutant iPSC-CMs treated with PDGFRB inhibitors. CB, crenolanib; SB, sunitinib. The experiments were repeated twice independently with similar results.
Extended Data Fig. 12 Proposed disease model of LMNA-related DCM.
We recruited a large family cohort with DCM and generated patient-specific iPSCs from several patients (n = 5) and healthy individuals (n = 2). We next used gene-edited isogenic iPSC lines (n = 4) and patient heart tissues to address the question why patients with LMNA-related DCM have increased manifestation of cardiac arrhythmias. The electrophysiological studies of mutant iPSC-CMs demonstrated that a mutation in LMNA was the cause of the increased arrhythmogenicity in LMNA-mutant iPSC-CMs. We also found that the LMNA mutation caused lamin A/C haploinsufficiency, which led to abnormal calcium homeostasis in mutant iPSC-CMs through upregulation of calcium-handling genes. Whole-transcriptome profiling (RNA-seq) further demonstrated an abnormal activation of the PDGF pathway in mutant iPSC-CMs. The inhibition of the PDGF signalling pathway by treatment with siRNA or FDA-approved inhibitors, such as sunitinib and crenolanib, could reverse the arrhythmic phenotype of LMNA-mutant iPSC-CMs. Cross-analysis of ChIP–seq, ATAC-seq and RNA-seq data revealed a possible underlying mechanism that lamin A/C haploinsufficiency could disrupt global chromatin conformation, resulting in abnormal gene expression in mutant iPSC-CMs. These findings were further corroborated by studies in cardiac tissues from healthy individuals and patients with LMNA-related DCM, thus validating a novel mechanism of LMNA-related DCM pathogenesis both in vitro and in vivo.
Supplementary information
Supplementary Figures
This file contains Source Data Gels
Rights and permissions
About this article
Cite this article
Lee, J., Termglinchan, V., Diecke, S. et al. Activation of PDGF pathway links LMNA mutation to dilated cardiomyopathy. Nature 572, 335–340 (2019). https://doi.org/10.1038/s41586-019-1406-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1406-x
This article is cited by
-
PDGF-BB accelerates TSCC via fibroblast lactates limiting miR-26a-5p and boosting mitophagy
Cancer Cell International (2024)
-
Cellular reprogramming as a tool to model human aging in a dish
Nature Communications (2024)
-
Systematic in vivo candidate evaluation uncovers therapeutic targets for LMNA dilated cardiomyopathy and risk of Lamin A toxicity
Journal of Translational Medicine (2023)
-
An atlas of lamina-associated chromatin across twelve human cell types reveals an intermediate chromatin subtype
Genome Biology (2023)
-
Spatiotemporally resolved transcriptomics reveals the subcellular RNA kinetic landscape
Nature Methods (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.