Abstract
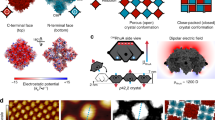
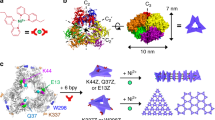
The ability of proteins and other macromolecules to interact with inorganic surfaces is essential to biological function. The proteins involved in these interactions are highly charged and often rich in carboxylic acid side chains1,2,3,4,5, but the structures of most protein–inorganic interfaces are unknown. We explored the possibility of systematically designing structured protein–mineral interfaces, guided by the example of ice-binding proteins, which present arrays of threonine residues (matched to the ice lattice) that order clathrate waters into an ice-like structure6. Here we design proteins displaying arrays of up to 54 carboxylate residues geometrically matched to the potassium ion (K+) sublattice on muscovite mica (001). At low K+ concentration, individual molecules bind independently to mica in the designed orientations, whereas at high K+ concentration, the designs form two-dimensional liquid-crystal phases, which accentuate the inherent structural bias in the muscovite lattice to produce protein arrays ordered over tens of millimetres. Incorporation of designed protein–protein interactions preserving the match between the proteins and the K+ lattice led to extended self-assembled structures on mica: designed end-to-end interactions produced micrometre-long single-protein-diameter wires and a designed trimeric interface yielded extensive honeycomb arrays. The nearest-neighbour distances in these hexagonal arrays could be set digitally between 7.5 and 15.9 nanometres with 2.1-nanometre selectivity by changing the number of repeat units in the monomer. These results demonstrate that protein–inorganic lattice interactions can be systematically programmed and set the stage for designing protein–inorganic hybrid materials.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Design models in PDB format are available on Github (https://github.com/pylesharley/DHR10micaX/tree/master/rosetta_models). Source data for Supplementary Figs. 2–6 are provided with the paper. All other data not included in the manuscript are available upon reasonable request from the corresponding authors.
Code availability
The Rosetta Macromolecular Modelling suite is available for non-commercial use at (https://www.rosettacommons.org). The specific Rosetta applications used were Rosetta Scripts46, Remodel47, and Pyrosetta48. Foldit33, a graphic user interface to Rosetta, was used as well. PyMOL49 was used to view the design models and prepare input files for Rosetta protocols.
A Github repository (https://github.com/pylesharley/DHR10micaX) contains the Rosetta protocols, input files and Python scripts used to model the protein assembles on mica, and corresponding README.txt files with instructions.
References
Sodek, J., Ganss, B. & McKee, M. D. Osteopontin. Crit. Rev. Oral Biol. Med. 11, 279–303 (2000).
Addadi, L., Joester, D., Nudelman, F. & Weiner, S. Mollusk shell formation: a source of new concepts for understanding biomineralization processes. Chemistry 12, 980–987 (2006).
Shaw, W. J. Solid-state NMR studies of proteins immobilized on inorganic surfaces. Solid State Nucl. Magn. Reson. 70, 1–14 (2015).
Staniland, S. S. & Rawlings, A. E. Crystallizing the function of the magnetosome membrane mineralization protein Mms6. Biochem. Soc. Trans. 44, 883–890 (2016).
Fukushima, T. et al. The molecular basis for binding of an electron transfer protein to a metal oxide surface. J. Am. Chem. Soc. 139, 12647–12654 (2017).
Davies, P. L. Ice-binding proteins: a remarkable diversity of structures for stopping and starting ice growth. Trends Biochem. Sci. 39, 548–555 (2014).
DeOliveira, D. B. & Laursen, R. A. Control of calcite crystal morphology by a peptide designed to bind to a specific surface. J. Am. Chem. Soc. 119, 10627–10631 (1997).
Masica, D. L., Schrier, S. B., Specht, E. A. & Gray, J. J. De novo design of peptide-calcite biomineralization systems. J. Am. Chem. Soc. 132, 12252–12262 (2010).
Song, R. & Cölfen, H. Additive controlled crystallization. CrystEngComm 13, 1249–1276 (2011).
Grigoryan, G. et al. Computational design of virus-like protein assemblies on carbon nanotube surfaces. Science 332, 1071–1076 (2011).
Brown, C. L., Aksay, I. A., Saville, D. A. & Hecht, M. H. Template-directed assembly of a de novo designed protein. J. Am. Chem. Soc. 124, 6846–6848 (2002).
Mustata, G.-M. et al. Graphene symmetry amplified by designed peptide self-assembly. Biophys. J. 110, 2507–2516 (2016).
Leow, W. W. & Hwang, W. Epitaxially guided assembly of collagen layers on mica surfaces. Langmuir 27, 10907–10913 (2011).
Tao, J. et al. Energetic basis for the molecular-scale organization of bone. Proc. Natl Acad. Sci. USA 112, 326–331 (2015).
Akutagawa, T. et al. Formation of oriented molecular nanowires on mica surface. Proc. Natl Acad. Sci. USA 99, 5028–5033 (2002).
Loo, R. W. & Goh, M. C. Potassium ion mediated collagen microfibril assembly on mica. Langmuir 24, 13276–13278 (2008).
Shin, S.-H. et al. Direct observation of kinetic traps associated with structural transformations leading to multiple pathways of S-layer assembly. Proc. Natl Acad. Sci. USA 109, 12968–12973 (2012).
Aghebat Rafat, A., Pirzer, T., Scheible, M. B., Kostina, A. & Simmel, F. C. Surface-assisted large-scale ordering of DNA origami tiles. Angew. Chem. Int. Edn Engl. 53, 7665–7668 (2014).
Ma, X. et al. Tuning crystallization pathways through sequence engineering of biomimetic polymers. Nat. Mater. 16, 767–774 (2017).
Brunette, T. J. et al. Exploring the repeat protein universe through computational protein design. Nature 528, 580–584 (2015).
Dyer, K. N. et al. High-throughput SAXS for the characterization of biomolecules in solution: a practical approach. Methods Mol. Biol. 1091, 245–258 (2014).
Schneidman-Duhovny, D., Hammel, M., Tainer, J. & Sali, A. Accurate SAXS profile computation and its assessment by contrast variation experiments. Biophys. J. 105, 962–974 (2013).
Kuwahara, Y. Comparison of the surface structure of the tetrahedral sheets of muscovite and phlogopite by AFM. Phys. Chem. Miner. 28, 1–8 (2001).
Boles, M. A., Engel, M. & Talapin, D. V. Self-assembly of colloidal nanocrystals: from intricate structures to functional materials. Chem. Rev. 116, 11220–11289 (2016).
Ruotolo, B. T. & Robinson, C. V. Aspects of native proteins are retained in vacuum. Curr. Opin. Chem. Biol. 10, 402–408 (2006).
Sahasrabuddhe, A. et al. Confirmation of intersubunit connectivity and topology of designed protein complexes by native MS. Proc. Natl Acad. Sci. USA 115, 1268–1273 (2018).
Whitelam, S. et al. Common physical framework explains phase behavior and dynamics of atomic, molecular, and polymeric network formers. Phys. Rev. X 4, 011044 (2014).
Fallas, J. A. et al. Computational design of self-assembling cyclic protein homo-oligomers. Nat. Chem. 9, 353–360 (2017).
Boyken, S. E. et al. De novo design of protein homo-oligomers with modular hydrogen-bond network-mediated specificity. Science 352, 680–687 (2016).
Wang, J. et al. Differential modulating effect of MoS2 on amyloid peptide assemblies. Chemistry 24, 3397–3402 (2018).
Chan, P., Curtis, R. A. & Warwicker, J. Soluble expression of proteins correlates with a lack of positively-charged surface. Sci. Rep. 3, 3333 (2013).
MacKerell, A. D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Kleffner, R. et al. Foldit Standalone: a video game-derived protein structure manipulation interface using Rosetta. Bioinformatics 33, 2765–2767 (2017).
McDaniel, J. R., Mackay, J. A., Quiroz, F. G. & Chilkoti, A. Recursive directional ligation by plasmid reconstruction allows rapid and seamless cloning of oligomeric genes. Biomacromolecules 11, 944–952 (2010).
Tang, N. C. & Chilkoti, A. Combinatorial codon scrambling enables scalable gene synthesis and amplification of repetitive proteins. Nat. Mater. 15, 419–424 (2016).
Rambo, R. P. ScÅtter: Software for SAXS Analysis. Version 3.0 http://www.bioisis.net/tutorial/9 (Diamond Light Source and SIBYLS beamline (12.3.1) of the Advanced Light Source, 2016).
Rambo, R. P. & Tainer, J. A. Characterizing flexible and intrisically unstructured biological macromoleucles by SAS using the Porod–Debye law. Biopolymers 95, 559–571 (2011).
Bernadó, P. & Svergun, D. I. Structural analysis of intrinsically disordered proteins by small-angle X-ray scattering. Mol. Biosyst. 8, 151–167 (2012).
VanAernum, Z. L. et al. Surface-induced dissociation of noncovalent protein complexes in an extended mass range orbitrap mass spectrometer. Anal. Chem. 91, 3611–3618 (2019).
Waitt, G. M., Xu, R., Wisely, G. B. & Williams, J. D. Automated in-line gel filtration for native state mass spectrometry. J. Am. Soc. Spectrom. 19, 239–245 (2008).
Marty, M. T. et al. Bayesian deconvolution of mass and ion mobility spectra: from binary interactions to polydisperse ensembles. Anal. Chem. 87, 4370–4376 (2015).
Kilpatrick, E. L., Liao, W. L., Camara, J. E., Turko, I. V. & Bunk, D. M. Expression and characterization of 15N-labeled human C-reactive protein in Escherichia coli and Pichia pastoris for use in isotope-dilution mass spectrometry. Protein Expr. Purif. 85, 94–99 (2012).
Frenkel, D. & Eppenga, R. Evidence for algebraic orientational order in a two-dimensional hard-core nematic. Phys. Rev. A 31, 1776–1787 (1985).
Newcomb, C. J., Qafoku, N. P., Grate, J. W., Bailey, V. L. & De Yoreo, J. J. Developing a molecular picture of soil organic matter–mineral interactions by quantifying organo–mineral binding. Nat. Commun. 8, 396 (2017).
Martin-Jimenez, D., Chacon, E., Tarazona, P. & Garcia, R. Atomically resolved three-dimensional structures of electrolyte aqueous solutions near a solid surface. Nat. Commun. 7, 12164 (2016).
Fleishman, S. J. et al. RosettaScripts: a scripting language interface to the Rosetta macromolecular modeling suite. PLoS One 6, e20161 (2011).
Huang, P. S. et al. RosettaRemodel: a generalized framework for flexible backbone protein design. PLoS One 6, e24109 (2011).
Chaudhury, S., Lyskov, S. & Gray, J. J. PyRosetta: a script-based interface for implementing molecular modeling algorithms using Rosetta. Bioinformatics 26, 689–691 (2010).
The PyMOL Molecular Graphics System. Version 2.1.1 https://pymol.org/2/ (Schrödinger, 2018).
Acknowledgements
We thank T. Brunette, P. Huang, F. Parmeggiani, Y. Hsia, W. Sheffler, T. Craven, S. Boyken and Z. Chen for suggestions; D. Alonso, L. Goldschmidt and P. Vecchiato for supporting computational resources; B. Legg and J. Tao for AFM support; B. Legg for providing the model of mica (001); L. Carter for size-exclusion chromatography with multi-angle light scattering (SEC-MALS) support. F. Busch and V. H. Wysocki provided native mass spectrometry support (NIH National Institute of General Medical Sciences award number P41GM128577). Development of the imaging protocols was supported by the Laboratory Directed Research and Development Office through the Materials Synthesis and Simulations Across Scales Initiative at Pacific Northwest National Laboratory (PNNL). AFM experiments on the DHR10-micaX and DHR10-micaX-NC were supported by the US Department of Energy (DOE), the Office of Basic Energy Sciences (BES), Biomolecular Materials Program (BMP) at PNNL. AFM experiments on DHR10-micaX-H formation of hexagonal lattices and analysis of DHR10-micaX binding kinetics were performed at PNNL and supported by the US DOE BES Energy Frontier Research Center CSSAS (The Center for the Science of Synthesis Across Scales) located at the University of Washington (award number DE-SC0019288). Design and synthesis of DHR10-micaX-H and its variants were performed at the University of Washington and supported by the US DOE BES BMP (award number DE-SC0018940). H.P. was supported by the Institute for Protein Design Materials Science Research Gift Fund and Michelson Medical Research Foundation, Protein Design Initiative Fund, and DOE Biomolecular Materials Program (award number DE-SC0018940). D.B. is funded by the Howard Hughes Medical Institute and Bruce and Jeannie Nordstrom / Patty and Jimmy Barrier Gift for the Institute for Protein Design Directors Fund. We thank the staff at the Advanced Light Source SIBYLS beamline at Lawrence Berkeley National Laboratory, including K. Burnett, G. Hura and J. Tainer for the services provided through the mail-in SAXS programme, which is supported by the DOE Office of Biological and Environmental Research Integrated Diffraction Analysis program DOE BER IDAT grant (award number DE-AC02-05CH11231) and NIGMS supported ALS-ENABLE (award number GM124169-01). PNNL is a multi-programme national laboratory operated for the Department of Energy by Battelle under contract number DE-AC05-76RL01830.
Reviewer information
Nature thanks Roberto A. Chica, Rajesh R. Naik and Sarah S. Staniland for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
H.P. designed, expressed, and purified the proteins. S.Z. developed the protocol to assemble protein on mica and performed the AFM experiments. S.Z. and J.J.D.Y. analysed the AFM data. D.B. and J.J.D.Y. supervised the project. All authors wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1-29 and Supplementary Tables 1-2 are in the Supplementary Information pdf, arranged so that each Supplementary Figure is on the page corresponding to its number. Table S1 is on page 10 with related Figure S10. Table S2 contains amino-acid sequences and is on pages 30-33.
Supplementary Data
This file contains source data for supplementary figure 2.
Supplementary Data
This file contains source data for supplementary figure 3.
Supplementary Data
This file contains source data for supplementary figure 4.
Supplementary Data
This file contains source data for supplementary figure 5.
Supplementary Data
This file contains source data for supplementary figure 6.
Rights and permissions
About this article
Cite this article
Pyles, H., Zhang, S., De Yoreo, J.J. et al. Controlling protein assembly on inorganic crystals through designed protein interfaces. Nature 571, 251–256 (2019). https://doi.org/10.1038/s41586-019-1361-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1361-6
This article is cited by
-
Blueprinting extendable nanomaterials with standardized protein blocks
Nature (2024)
-
Directing polymorph specific calcium carbonate formation with de novo protein templates
Nature Communications (2023)
-
Crystal engineering of quercetin via combined adsorption of polyvinylpyrrolidone and tannin
Korean Journal of Chemical Engineering (2023)
-
Suppressing high-dimensional crystallographic defects for ultra-scaled DNA arrays
Nature Communications (2022)
-
Bio-inspired synthesis of transition-metal oxide hybrid ultrathin nanosheets for enhancing the cycling stability in lithium-ion batteries
Nano Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.